- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
MAPK Phosphorylation Assay with Leaf Disks of Arabidopsis
Published: Vol 3, Iss 19, Oct 5, 2013 DOI: 10.21769/BioProtoc.929 Views: 13915
Reviewed by: Anonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Western Blot Analysis of Chloroplast HSP70B in Chlorella Species
Stephka Chankova [...] Nadezhda Yurina
Aug 5, 2013 10947 Views
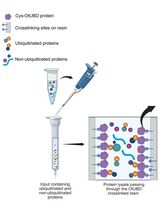
Use of a High-Affinity Ubiquitin-Binding Domain to Detect and Purify Ubiquitinated Substrates and Their Interacting Proteins
Nitu Saha [...] Mark Hochstrasser
Sep 5, 2025 3825 Views
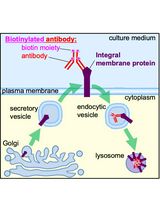
Monitoring Endocytosis of Integral Membrane Proteins Using Western Blot-Based Detection of Biotinylated Antibody Uptake
Alexandra Graninger and Prasanna Satpute-Krishnan
Nov 20, 2025 2244 Views
Abstract
Activation of mitogen activated protein kinases (MAPKs) is involved in many abiotic and biotic stress responses including plant defense. MAPK acitvation is based on the dual phosphorylation of threonine (T) and tyrosin (Y) residues (T-x-Y motif) in the activation loop of the MAPK protein. By determination of the phosphorylation status of a specific MAPK one can detect if the MAPK has been activated or not.
This protocol describes how to analyze the phosphorylation status of Arabidopsis MAPKs MPK3 and MPK6 by using leaf disks, western blotting and a specific antibody (Figure 1). It can also be used for the analysis of MAPKs in other plant systems although some alterations regarding protein extraction might be necessary.
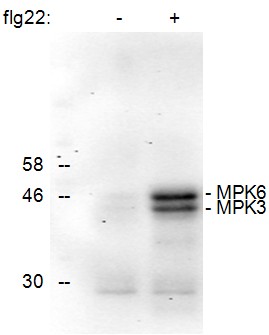
Figure 1. Detection of the phosphorylation of Arabidopsis thaliana MAPKs MPK6 and MPK3 upon treatment of seedlings for 15 min with the active epitope (flg22, 1 μM) of the bacterial elicitor flagellin (+). No phosphorylated MAPKs were detected in the control treated sample (-).
Materials and Reagents
- Arabidopsis thaliana adult plants
- Liquid nitrogen
- Phospho-p44/42 MAPK (Erk1/2) (Thr202/Tyr204) Rabbit mAb #4370 (Cell Signaling Technology) (http://www.cellsignal.com/products/4370.html)
- Anti-Rabbit secondary antibody
- Tris-HCl
- Glycerol
- SDS
- Dithiothreitol (DTT)
- Bromphenol blue
- Reagents for SDS-PAGE and Western Blotting
- 6x Protein extraction/loading buffer (see Recipes)
Equipment
- 1.5 ml Microcentrifuge tubes
- Small petri dishes (~ 5 cm diameter) or 6-well plates
- Cork borer (~ 10 mm diameter)
- Glass beads (~ 1-2 mm diameter)
- Silamat S6 (http://www.ivoclarvivadent.com/en/products/equipment/mixer/silamat-s6)
- Alternatively for 4. and 5. Mortar (~ 5 cm diameter) and pestle
- Vortexer
- Heating block
- Centrifuge
- Equipment for SDS-PAGE, Western Blotting and detection
Procedure
- Leaf disks are cut from adult Arabidopsis (Arabidopsis thaliana) leaves using the cork borer. Try to take leaves of similar age and fitness but from different plants.
- Weigh one exemplary leaf disk and note the weight (usually ~ 20 mg).
- Float disks on distilled water in small petri dishes or 6-well plates (at least 3 per dish or well in approximately 5 ml of water).
- Wait overnight (usually 16 h, dishes closed) and keep the dishes in an undisturbed and controlled place (e.g. growth chamber of origin). This will lead to dephosphorylation of MAPKs which have been phosphorylated upon harvesting.
- Next morning perform the treatment, for example add an elicitor peptide of which you would like to know if it triggers MAPK phosphorylation. We add it to a final concentration of 1 μM, mix carefully to not mechanically stress the leaf tissue and in most cases wait for 15 min. As a proper control use a mock treatment (e.g. solvent only) and also incubate for 15 min. This is critical, since MAPK phosphorylation is triggered very easily by many different stresses. Thus you need to show that your control treatment does not lead to phosphorylation of MAPKs. Other treatments could be addition of chemicals, wounding stress, UV-treatment etc.
Note: Phosphorylation and activation of MAPKs happens very quickly and is transient. It starts roughly 2-5 min after treatment and lasts at least for 1 h or longer.
- Harvest all disks from one well after the desired incubation time quickly with a forceps and dry them briefly on a paper towel (residual water will dilute your sample and interfere with the grinding procedure). Place them into a microcentrifuge tube (preferably safe lock) that contains 5 glass beads and shock-freeze them in liquid nitrogen. Repeat this procedure for each well.
Note: Here you have to be quick! Harvesting the disks (mechanical stress) will induce MAPK phosphorylation, thus you should not take longer than 1 min for each sample to avoid false positive phosphorylation results!
- Grind tissue to fine powder (the tissue should not thaw before the next step!). You can do this by using a mortar and pestle (cooled in liquid nitrogen) but we do not recommend this. Especially if you have many samples grind the tissue by using a Silamat S6 (or similar machine) that grinds the tissue in the tube due to rapid movement and the previously added glass beads.
- Add extraction/loading buffer. For example for 60 mg of tissue, use 60 μl and use the same ratio depending on sample weight.
- Vortex vigorously until the sample has completely thawed and potential buffer precipitates have been dissolved again. Keep at room temperature and continue with further samples.
- Spin all samples 15 sec at 11,000 x g to get down the sample from the lid.
- Boil (95 °C) for 5-10 min.
- Let cool for 3 min.
- Centrifuge at 11,000 x g for 5 min to precipitate glass beads and debris.
- Take supernatant from the top (do not disturb the pellet) and load 15 μl on an SDS-PAGE gel (10% or 12%, 1 mm thickness).
- Perform your standard Western Blotting method (for us wet and semi-dry worked, and similarly both Alkaline Phosphatase and Horseradish peroxidase worked). We use the primary antibody at 1:2,000 dilutions.
Notes
- The “extraction buffer” used in this protocol is a 6x loading buffer. Usually a buffer called “Lacus” (which is very complex) is used for the extraction of phosphorylated MAPKs but we observed, that it works well with just adding the undiluted 6x loading buffer. Since it contains high amounts of SDS do not keep the samples on ice after adding the buffer. Since the samples are very cold SDS might precipitate anyway (white precipitate). In that vortex the samples at room temperature until the SDS dissolved again.
- This MAPK antibody detects in principle all phosphorylated MAPKs so it might produce more bands than just the two of MPK3 and MPK6 (especially when using other plant material). If possible include mpk3 and mpk6 mutants in your analysis to clarify the origin of the bands you see.
- According to the manufacturer the used MAPK antibody detects single as well as dual phosphorylated MAPKs. To our knowledge you cannot discriminate between the two states. Keep that in mind when interpreting the data.
Recipes
- 6x Protein extraction/loading buffer
0.35 M Tris-HCl pH 6.8
30% (v/v) glycerol
10% (v/v) SDS
0.6 M DTT
0.012% (w/v) bromphenol blue
Acknowledgments
This work was supported by the Swiss National Science Foundation (grant 31003A_127563; to TB) and by stipends to SB from the European Molecular Biology Organisation (EMBO: ALTF 61-2010) and the Leopoldina Fellowship Programme of the National Academy of Science Leopoldina (LPDS 2009-35).
References
- Flury, P., Klauser, D., Schulze, B., Boller, T. and Bartels, S. (2013). The anticipation of danger: microbe-associated molecular pattern perception enhances AtPep-triggered oxidative burst. Plant Physiol 161(4): 2023-2035.
Article Information
Copyright
© 2013 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Flury, P., Klauser, D., Boller, T. and Bartels, S. (2013). MAPK Phosphorylation Assay with Leaf Disks of Arabidopsis. Bio-protocol 3(19): e929. DOI: 10.21769/BioProtoc.929.
- Flury, P., Klauser, D., Schulze, B., Boller, T. and Bartels, S. (2013). The anticipation of danger: microbe-associated molecular pattern perception enhances AtPep-triggered oxidative burst. Plant Physiol 161(4): 2023-2035.
Category
Plant Science > Plant immunity > Perception and signaling
Cell Biology > Cell signaling > Phosphorylation
Biochemistry > Protein > Immunodetection > Western blot
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










