- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
3H-Penciclovir (3H-PCV) Uptake Assay
Published: Vol 3, Iss 17, Sep 5, 2013 DOI: 10.21769/BioProtoc.887 Views: 8952
Reviewed by: Lin FangFanglian He

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
NanoPDLIM2-Based Combination Therapy for Lung Cancer Treatment in Mouse Preclinical Studies
Thi Hoa Le [...] Zhaoxia Qu
Sep 5, 2025 2274 Views
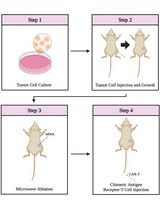
Combining Microwave Ablation With CAR-T-Cell Therapy in Tumor-Bearing Mouse Models
Bihui Cao [...] Jia Shen
Oct 20, 2025 2372 Views
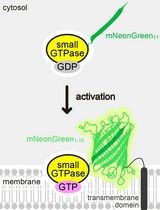
Detecting the Activation of Endogenous Small GTPases via Fluorescent Signals Utilizing a Split mNeonGreen: Small GTPase ActIvitY ANalyzing (SAIYAN) System
Miharu Maeda and Kota Saito
Jan 5, 2026 561 Views
Abstract
Thymidine Kinase from human Herpes simplex virus type 1 (HSV1-TK) in combination with specific substrate prodrug nucleotide analogue ganciclovir (GCV) has been widely used as suicidal therapeutic gene for cancer gene therapy. HSV1, and its mutant (HSV1-sr39TK) with improved substrate specificity, were used as reporter genes for PET-imaging of various biological functions in small animals, by combining with radiolabeled substrates such as 18F-FHBG and 124I-FIAU. 3H-Penciclovir (PCV) uptake assay is a method of choice used to determine the expression level of HSV1-TK in mammalian cells and tissues. HSV1-TK phosphorylate PCV and result in the formation of penciclovir monophosphate, and its subsequent phopsphorylation by cellular TK lead to the formation of penciclovir triphosphate, which is trapped selectively in cells express HSV-TK. 3H-Penciclovir enables the detection of penciclovir uptake of mammalian cells and tissues by radioactive procedures such as scintillation counting. Here we describe the protocol to carry out 3H-Penciclovir uptakes in mammalian cells.
Materials and Reagents
- Control and experimental cells (HEK293T and HEK293T-sr39TK)
- Dulbecco's Modified Eagle Medium (DMEM) (Life technologies, catalog number: 11965-084 )
- 3H-Penciclovir (specific activity 14.9 ci mmol-1) (Moravek Biochemicals, La Brea, CA, USA)
- Sodium hydroxide (NaOH) (0.1 N)
- Phosphate buffered saline (PBS) (pH 7.4) (Life technologies, catalog number: 10010-056 )
- Glass Scintillation vials
- Cytoscint Scintillation fluid (Acros Organics, Geel, Belgium)
- Cell culture plates (Greiner Bio-One, USA)
- Bradford protein assay reagent (Bio-Rad Laboratories, Hercules, CA)
- BSA (Sigma-Aldrich, catalog number: A9418 )
- Wash solution (Ice cold PBS) (pH 7.4)
Equipment
- 37 °C 5% CO2 Cell culture incubator (Themo Fisher Scientific, model: Napco 8000)
- Scintillation counter (Beckman Coulter, Brea, CA)
- Shaker (Boekel scientific, Model: 260350 )
- Radioactive Chemical hood (Hamilton, Two Rivers, WI)
- Pipettes (Gilson, Middleton, WI)
- Disposal (As per Environment and Health safety regulation)
- Spectrophotometer (Varian Inc, model: Varian Cary 50 )
Procedure
- Mammalian HEK293T and HEK293T cells stably expressing sr39-HSV1-TK (HEK293T-sr39TK) should be plated at a confluency of 60-80% in 12 well culture plates (150,000 cells/well in 1 ml DMEM). Incubate at 37 °C with 5% CO2 for 24 h.
- Twenty-four hours after initial plating remove the medium and add 0.5 μl of 3H-Penciclovir/well (0.5 mCi/ml of 8-3H-Penciclovir; specific activity 14.9 ci mmol-1) in 0.5 ml of respective culture medium.
- Incubate at 37 °C for 3 h. Maintain the incubation time consistent between samples.
- Keep the same cells (HEK293T) plated in equal number express no HSV1-TK as control.
- After 3 hours of incubation remove the medium carefully without disturbing cells, and wash the plates twice with 0.5 ml each of ice cold PBS. Pool the medium and wash solutions from respective well to measure the remaining total activities.
- Add 0.5 ml of 0.1 N NaOH to each well and leave it for 10 min at room temperature in a shaker to lyse the cells.
- Transfer 20 μl of homogenous cell lysate to measure total protein concentration by using Bradford protein assay reagent. In brief, to each 20 μl of cell lysate add 1 ml of 1x Bradford protein assay reagent and read the absorbance at 595 nm (A595) 5 min after incubation at room temperature in a spectrophotometer. Use standard graph prepared with the standards (BSA) diluted in 0.1 N NaOH to calculate the total protein present in 0.5 ml of cell lysate.
- Transfer the remaining cell lysate (480 μl) to scintillation vial and 5 ml of biodegradable scintillation fluid (Cytoscint fluid). Cells lacking HSV1-TK expression exposed to 3H-Penciclovir serve as negative control.
- The total remaining activity from each sample should be measured by using 480 μl of solution taken from the pooled medium and the wash solution, and also measured from 0.5 μl of 3H-Penciclovir diluted in 480 μl of medium.
- Measure the radioactivity from different samples in a Scintillation counter by taking an average reading/min by reading for 5 min. Normalize the results with protein concentration and relate with total activity.
- Results should be expressed as the percentage conversion of 3H-PCV per milligram of protein/total count, or as the conversion of 3H -PCV in moles per milligram of protein/total activity (Radioactivity within cells/[Radioactivity remained in medium + Radioactivity within cells]).
Acknowledgments
We thank Dr Sanjiv Sam Gambhir, Chairman, Department of Radiology, Stanford University for providing helpful support and facility for conducting this research. We thank the Department of Radiology, Stanford University for funding support (R. Paulmurguan), and thank the Canary Center at Stanford for providing the facilities. We also thank NIH-NCI RO1CA161091 (R. Paulmurugan) for partial funding support.
References
- Sekar, T. V., Foygel, K., Willmann, J. K. and Paulmurugan, R. (2013). Dual-therapeutic reporter genes fusion for enhanced cancer gene therapy and imaging. Gene Ther 20(5): 529-537.
Article Information
Copyright
© 2013 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Sekar, T. V. and Paulmurugan, R. (2013). 3H-Penciclovir (3H-PCV) Uptake Assay. Bio-protocol 3(17): e887. DOI: 10.21769/BioProtoc.887.
Category
Cancer Biology > General technique > Cancer therapy > Gene expression
Cell Biology > Cell signaling > Stress response
Biochemistry > Protein > Activity
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










