- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Plant Endo-β-mannanase Activity Assay
Published: Vol 3, Iss 17, Sep 5, 2013 DOI: 10.21769/BioProtoc.883 Views: 9981
Reviewed by: Tie Liu

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Immunofluorescence for Detection of TOR Kinase Activity In Situ in Photosynthetic Organisms
Ana P. Lando [...] Giselle M. A. Martínez-Noël
Dec 20, 2024 1866 Views

An Activity-Based Proteomics with Two-Dimensional Polyacrylamide Gel Electrophoresis (2D-PAGE) for Identifying Target Proteases in Arabidopsis Apoplastic Fluid
Sayaka Matsui and Yoshikatsu Matsubayashi
Mar 5, 2025 2037 Views
Advancing 2-DE Techniques: High-Efficiency Protein Extraction From Lupine Roots
Sebastian Burchardt [...] Emilia Wilmowicz
Oct 5, 2025 1804 Views
Abstract
Endo-β–mannanases in plant require post-translational modification, such as N-glycosylation and disulfide-linked dimerization, for their catalytic activity. Determination of the plant endo-β–mannanase activity needs to modify the assay conditions for optimizing their enzymatic reaction. Here, we describe a modified method for plant endo-β–mannanase assay. A high-salt buffer without thiol reductants is required for effective extraction of the enzyme. The enzyme is able to digest water-insoluble AZCL galactomannan to release water soluble dyed fragments, which is detected through measurement of absorbance at 590 nm wavelength. Increase in absorbance at 590 nm is correlated directly with enzyme activity.
Materials and Reagents
- Liquid nitrogen
- BCA Reagent (Tiangen Biotech, catalog number: PA115-01 )
- Bovine serum albumin
- AZC L-galactomannan (Megazyme, catalog number: I-AZGMA )
- Commercial Aspergillus niger endo-β-mannanase (Megazyme, catalog number: E-BMANN )
- 100 mM phenylmethanesulfonyl fluoride (PMSF) (see Recipes)
- 0.5 M ethylene diaminete traacetic acid (EDTA) (see Recipes)
- Extraction buffer (see Recipes)
- 0.1 M sodium acetate buffer (pH 5.0) (see Recipes)
Equipment
- Mortar and pestle
- 10,000 Mr cut-off filter (EMD Millipore, catalog number: UFC801096 )
- Centrifuge
- Incubator shaker
- Water bath
- Microplate reader or Spectrophotometer
Procedure
- Samples (~ 10 g developing xylem or leaves from one-year-old poplar) are ground in liquid nitrogen to a fine powder and homogenized at 4 °C in 1.5-volume fold of extraction buffer for 1 h (strong enzymatic activity can be detected in developing xylem).
- The homogenate is centrifuged at 10,000 x g for 30 min at 4 °C.
- The supernatant is then passed through a 10,000 Mr cut-off filter and dehydrated to < 500 μl, then the protein is diluted to ~1 μg/μl in 0.1 M sodium acetate buffer (pH 5.0) at 4 °C.
- The protein extraction is measured by BCA Reagent using bovine serum albumin as a standard.
- 200 μl of reaction mixture containing 100 μl of 1% AZC L-galactomannan (w/v, in 0.1 M sodium acetate buffer, pH 5.0) and 20 μg of extracted proteins or heated inactive proteins (100 °C 10 min, as control) is incubated at 40 °C for 2 h with continuous shake.
- The reaction mixture is boiled at 100 °C for 5 min and centrifuged at 12,000 x g for 5 min.
- The absorbance (A) of the supernatant at 590 nm is determined. The background values (A0) obtained using heated inactive proteins are subtracted from values (A1) obtained using active extract (A = A1 - A0).
- Standardization
- Enzyme activity of a serial dilutions of a commercial Aspergillus niger endo-β-mannanase (E-BMANN, Megazyme) is determined under the conditions: 200 μl of reaction mixture containing 100 μl of 1% AZC L-galactomannan and E-BMANN is incubated at 40 °C for 2 h.
- A standard curve correlated with E-BMANN activity is shown in Figure 1. For absorbance values in a range of 0.05~0.9, these values can be calculated by reference to the equation: Y = SX + C. Where:
Y = endo-β-Mannanase activity (in micro-Units/assay, i.e. per 200 μl)
S = Slope of the calibration graph
X = Absorbance of the reaction at 590 nm (A)
C = Intercept on the Y-axis - According to the manufacturer’s instruction, one Unit of activity is defined as the amount of enzyme required to release one micromole of mannose reducing-sugar equivalents per minute under the defined assay conditions (1 micro-Unit = 1 pmol/min).
- Enzyme activity of a serial dilutions of a commercial Aspergillus niger endo-β-mannanase (E-BMANN, Megazyme) is determined under the conditions: 200 μl of reaction mixture containing 100 μl of 1% AZC L-galactomannan and E-BMANN is incubated at 40 °C for 2 h.
- Calculation of enzyme activity: endo-β-Mannanase activity in the sample is determined by reference to the standard curve to convert absorbance values to micro-Units per assay (Y), then further to micro-Units per μg protein (Y/[20 μg protein], pmol/min/[μg protein]).
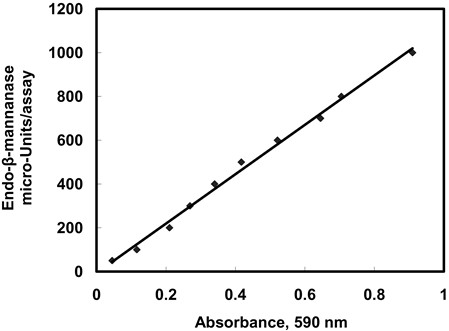
Figure 1. Endo-β-mannanase standard curve on the commercial endo-β-mannanase.
Recipes
- 100 mM Phenylmethanesulfonyl fluoride (PMSF) 10 ml
Mix 0.174 g of PMSF with 10 ml isopropanol
Store in small aliquots at -20 °C - 0.5 M Ethylene diamine tetraacetic acid (EDTA) (pH 8.0) (1 L)
Dissolve 186.1 g EDTA-Na.2H2O in 800 ml dH2O
Adjust pH to 8.0 with NaOH (~20 g NaOH particles)
Add dH2O to 1 L
Autoclave at 121 °C for 20 min
Store at RT - Extraction buffer (1 L)
1 M sodium acetate buffer (pH 5.0)
10 mM EDTA
10 mM sodium azide
3 mM PMSF
Mix 57 ml glacial acetic acid (1.05 g/ml) and 20 ml 0.5 M EDTA with 800 ml dH2O
Adjust pH to 5.0 with NaOH
Add 0.65 g sodium azide
Add dH2O to 1 L
Add 30 μl of 100 mM PMSF per ml extraction buffer immediately before use
Note: Do not add the sodium azide until pH is adjusted. Acidification of sodium azide will release a poisonous gas. - 0.1 M sodium acetate buffer (pH 5.0)
Mix 5.7 ml glacial acetic acid (1.05 g/ml) with 800 ml dH2O
Adjust pH to 5.0 with NaOH
Add 0.65 g sodium azide
Add dH2O to 1 L
Acknowledgments
This protocol was adapted from Zhao et al. (2013).
References
- Zhao, Y., Song, D., Sun, J. and Li, L. (2013). Populus endo-beta-mannanase PtrMAN6 plays a role in coordinating cell wall remodeling with suppression of secondary wall thickening through generation of oligosaccharide signals. Plant J 74(3): 473-485.
Article Information
Copyright
© 2013 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Zhao, Y. and Li, L. (2013). Plant Endo-β-mannanase Activity Assay. Bio-protocol 3(17): e883. DOI: 10.21769/BioProtoc.883.
Category
Plant Science > Plant biochemistry > Protein > Activity
Biochemistry > Protein > Activity
Plant Science > Plant biochemistry > Protein > Isolation and purification
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link












