- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Modified Single-Cell Transient Gene Expression Assay in Barley Epidermal Cells
Published: Vol 3, Iss 9, May 5, 2013 DOI: 10.21769/BioProtoc.690 Views: 10690

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
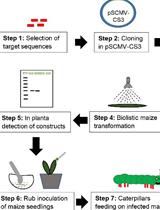
Inoculation of Maize with Sugarcane Mosaic Virus Constructs and Application for RNA Interference in Fall Armyworms
Iram Gull and Georg Jander
Jul 20, 2023 2252 Views
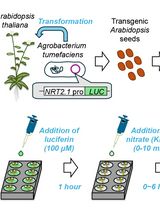
A Microplate-Based Expression Monitoring System for Arabidopsis NITRATE TRANSPORTER2.1 Using the Luciferase Reporter
Yoshiaki Ueda and Shuichi Yanagisawa
Dec 5, 2024 1820 Views
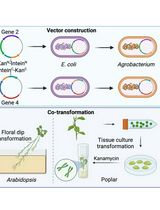
A Novel Gene Stacking Method in Plant Transformation Utilizing Split Selectable Markers
Guoliang Yuan [...] Xiaohan Yang
Feb 20, 2025 2021 Views
Abstract
Transient gene expression via biolistic particle delivery is a widely used technique for gene functional analysis in plants. In this protocol we describe a modified single-cell transient expression assay through transformation with a particle inflow gun of the model PDS-1000/He system (Bio-Rad). This assay was originally optimized for analyzing cell death activity and disease resistance function of the barley MLA (mildew locus A) disease resistance proteins against the powdery mildew fungus, which can be further adopted for other purposes for other types of plant proteins and in some other plant species, including Arabidopsis thaliana.
Keywords: Transient gene expressionMaterials and Reagents
- Barley (Hordeum vulgare L.) plants, 1 week old seedlings
- Powdery mildew strain(s), fresh conidiospores as inoculum
- Benzimidazol (Genview, catalog number: 51-17-2 )
- Agar (Japan, plant cell culture tested)
- CaCl2 (Sigma-Aldrich, catalog number: C7902 )
- X-gluc (Inalco, catalog number: 1758-0600 )
- Coomassie Brilliant Blue R-250 (Amresco, catalog number: 0472 )
- Ethanol (Beijing Chemical Works)
- Glycerol (Beijing Chemical Works)
- Methanol (Beijing Chemical Works)
- Lactic acid (Beijing Chemical Works)
- Plasmid
- Reporter DNA
- K3Fe [CN6] (Sinopharm Chemical Reagent, catalog number: 10016718 )
- Triton X-100 (AMRESCO, catalog number: 0694 )
- Spermidine (Sigma-Aldrich, catalog number: S-4139 ) (see Recipes)
- GUS staining solution (see Recipes)
- Destaining solution (see Recipes)
- Benzimidazol plates (see Recipes)
- Coomassie blue solution (see Recipes)
Equipment
- PDS-1000/He delivery system (Bio-Rad Laboratones)
- Macrocarrier (Bio-Rad Laboratones, catalog number: 1652335 )
- Rupture disc (900 psi) (Bio-Rad Laboratones, catalog number: 1652328 )
- Centrifuges (Eppendorf, catalog number: 5424 )
- Fluorescence microscope (Carl Zeiss, Axio Scope. A1 )
- pH meter (Mettler Toledo, FE20K)
- Gold microcarrier: 1.0 μm in diameter (Bio-Rad Laboratones,catalog number: 165-2263 )
Procedure
- Preparations
- Inoculate plants with powdery mildew spores to prepare inoculum and sow barley seeds to grow plants for bombardment one week in advance. The plants were grown in a growth chamber under a 16 h/8 h, 20 °C/18 °C day/night cycle with 70% relative humidity.
- Prepare Benzimidazol plates one day in advance.
- Inoculate plants with powdery mildew spores to prepare inoculum and sow barley seeds to grow plants for bombardment one week in advance. The plants were grown in a growth chamber under a 16 h/8 h, 20 °C/18 °C day/night cycle with 70% relative humidity.
- Bombardment
- Cut primary leaves and put them on Benzimidazol plates with adaxial side up (Figure 1), 3-5 leaves per petridish (90 mm) per shot, incubate at least 4 h before shooting.

Figure 1. Picture showing preparation of Benzimidazol agar plate with barley leaves for bombardment. 5-6 barley primary leaves were detached from 1 week old barley seedlings and put side by side with adaxial side up on prepared Benzimidazol agar plate. - Prepare gold particles (20 shots):
- Weigh 9 mg gold particles in a 1.5 ml tube.
- Add 1 ml 70% ethanol, vortex 5 min, sediment particles for 15 min on bench.
- Spin 2 sec. (about 2,000 rpm), discard supernatant.
- Repeat 3 times: add 1 ml sterile H2O, vortex 2 min, sediment 1 min, spin 2 sec. (about 2,000 rpm), discard supernatant.
- Add 1 ml of 50% glycerol (in water), vortex (gold particles can be stored at -20 °C for 2-3 weeks).
- Weigh 9 mg gold particles in a 1.5 ml tube.
- Coat the gold particles (use 50 μl gold particle solution for one shot):
- Vortex gold particle for at least 5 min.
- Mix equal molar plasmid and reporter DNA (e.g. GUS or GFP reporter), do not use more than 2 μg DNA in total, add ddH2O when volume is less than 5 μl.
- Aliquot 50 μl gold particles into each empty tube, then add DNA solution.
- While vortexing, add: 50 μl 2.5 M CaCl2 drop-by-drop, then 20 μl 0.1 M spermidine, vortex for 3 min in total.
- Sediment particles for 1 min, spin 2 sec. (2,000 rpm), discard supernatant.
- Add 140 μl 70% ethanol, vortex, spin 2 sec. (2,000 rpm), discard supernatant.
- Add140 μl 100% ethanol, vortex, spin 2 sec. (2,000 rpm), discard supernatant.
- Add 15 μl 100% ethanol, vortex, store on ice until used.
- Vortex gold particle for at least 5 min.
- Bombard, for each shot repeat the following steps:
- Fix the macrocarriers in macrocarier holder, suspend particles by pipetting, and apply the particles onto the macrocarrier. Dry on the bench.
- Dip rupture disc (900 psi) in 100% (v/v) 2-propanol and subsequentlyplace it into rupture disk retaining cap, add few more drops of 2-propanol.
- Insert macrocarrier holder with stop-screen in stop screen holder at position 1 (from top) (Figure 2).
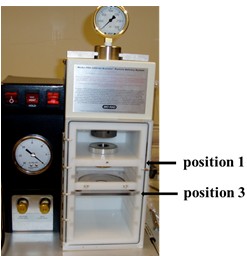

Figure 2. Picture showing the PDS-1000/He delivery system. Indicated are position 1 and 3 that reserved for macrocarrier holder and patridish holder, respectively. - Insert petridish with leaves at position 3 (Figure 2).
- Apply vacuum up to 27 inches of mercury, trigger the shot.
- Arrange leaves on the petridish, put in incubator.
Note: We put the leaves on the dish side by side with adaxial side up (Figure 1).
- Fix the macrocarriers in macrocarier holder, suspend particles by pipetting, and apply the particles onto the macrocarrier. Dry on the bench.
- (Omit this step if fungal inoculation is not necessary)
Inoculate with powdery mildew condiospores at least 4 h after bombardment.
- Cut primary leaves and put them on Benzimidazol plates with adaxial side up (Figure 1), 3-5 leaves per petridish (90 mm) per shot, incubate at least 4 h before shooting.
- For GFP index scoring
36-48 h after bombardment count GFP expressing cell numbers using fluorescence microscope.
Note: The total number of cells here is the sum of compatible (haustorium, seccondary hyphae) and incompatible (only appressorium) on GUS expressing cells. We score all of the five leaves and at least 60 cells were scored.
- For Fungal Haustorium index scoring
- 48 h after fungal spores inoculation
Stain leaves for GUS expression: put leaves into 15 ml falcon tube containing about 8 ml X-gluc staining solution, vacuum infiltrate 5 min for 3 times, and incubate overnight to 24 h at 37 °C. - 1 day after GUS staining
Remove GUS staining solution, add about 10 ml destaining solution, store at RT at least 2 days. - When time available:
Stain for the fungus:- Transfer leaves to large volume of ddH2O for 1 h.
- Stain in coomassie solution for few seconds.
- Wash twice in water.
- Mount on microscope slide in 50% glycerol. Once on the slide the samples should be scored within few days.
- Transfer leaves to large volume of ddH2O for 1 h.
- Score compatible (visible intracellular haustorium, and sometimes secondary hyphae on leaf surface) and incompatible (only fungal appressorium) interaction cell/site for GUS expressing cells.
- 48 h after fungal spores inoculation
Recipes
- Benzimidazol plates
1% agar in water with 85 μM Benzimidazol (from 8.5 mM stock solution in water, 100 x)
Note: The pH value for these plates is about 6.5. It is not necessary to adjust the pH value. - Spermidine
0.1 M solution, 1 g solution mix with 67.8 ddH2O, filter sterilized.
Aliquot and stored at -20 °C (note: it's very hygroscopic and air sensitive, close and put back to freezer immediately when done.)
2.5 M CaCl2 in water, sterile filtrate store in room temperature - GUS staining solution
0.1 M Na2HPO4/NaH2PO4 (pH 7.0)
10 mM Na-EDTA
5 mM K4Fe[CN6]
5 mM K3Fe[CN6]
0.1% Triton X-100 (v/v)
20% methanol (v/v)
1 g/L X-gluc
adjust to pH 7.0 - Destaining solution: stock solution
50% glycerol
25% lactic acid
25% H2O
Dissolve 1 volume stock solution in 2 volumes ethanol. - Coomassie blue solution
0.6% coomassie blue (w/v) in 100% (v/v) methanol/or ethanol
Acknowledgments
This protocol is adapted from Shirasu et al. (1999); Shen et al. (2012) and Bai et al. (2012).
References
- Bai, S., Liu, J., Chang, C., Zhang, L., Maekawa, T., Wang, Q., Xiao, W., Liu, Y., Chai, J., Takken, F. L., Schulze-Lefert, P. and Shen, Q. H. (2012). Structure-function analysis of barley NLR immune receptor MLA10 reveals its cell compartment specific activity in cell death and disease resistance. PLoS Pathog 8(6): e1002752.
- Shen, Q. H., Zhou, F., Bieri, S., Haizel, T., Shirasu, K. and Schulze-Lefert, P. (2003). Recognition specificity and RAR1/SGT1 dependence in barley Mla disease resistance genes to the powdery mildew fungus. Plant Cell 15(3): 732-744.
- Shirasu, K., Nielsen, K., Piffanelli, P., Oliver, R., and Schulze-Lefert, P. (1999). Cell-autonomous complementation of mlo resistance using a biolistic transient expression system. Plant J 17: 293-299.
Article Information
Copyright
© 2013 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Bai, S., Chang, C., Han, X. and Shen, Q. (2013). Modified Single-Cell Transient Gene Expression Assay in Barley Epidermal Cells. Bio-protocol 3(9): e690. DOI: 10.21769/BioProtoc.690.
- Bai, S., Liu, J., Chang, C., Zhang, L., Maekawa, T., Wang, Q., Xiao, W., Liu, Y., Chai, J., Takken, F. L., Schulze-Lefert, P. and Shen, Q. H. (2012). Structure-function analysis of barley NLR immune receptor MLA10 reveals its cell compartment specific activity in cell death and disease resistance. PLoS Pathog 8(6): e1002752.
Category
Plant Science > Plant molecular biology > DNA > Gene expression
Cell Biology > Single cell analysis > Cell carrier
Molecular Biology > DNA > Gene expression
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










