- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Hypochlorite Stress Assay for Phenotypic Analysis of the Halophilic Archaeon Haloferax volcanii Using an Improved Incubation Method and Growth Monitoring
Published: Vol 12, Iss 22, Nov 20, 2022 DOI: 10.21769/BioProtoc.4557 Views: 1553
Reviewed by: Gal HaimovichYufang LuWolf Dieter Röther

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
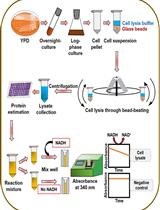
An Assay to Determine NAD(P)H: Quinone Oxidoreductase Activity in Cell Extracts from Candida glabrata
Anamika Battu [...] Rupinder Kaur
Nov 5, 2021 3056 Views
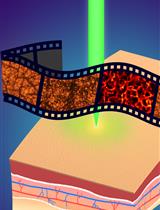

Quantitative and Anatomical Imaging of Human Skin by Noninvasive Photoacoustic Dermoscopy
Zhiyang Wang [...] Sihua Yang
Apr 5, 2022 2296 Views
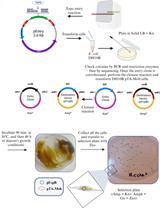
Mobilization of Plasmids from Bacteria into Diatoms by Conjugation Technique
Federico Berdun [...] Eduardo Zabaleta
Mar 5, 2024 1919 Views
Abstract
The study of haloarchaea provides an opportunity to expand understanding of the mechanisms used by extremophiles to thrive in and respond to harsh environments, including hypersaline and oxidative stress conditions. A common strategy used to investigate molecular mechanisms of stress response involves the deletion and/or site-directed mutagenesis of genes identified through omics studies followed by a comparison of the mutant and wild-type strains for phenotypic differences. The experimental methods used to monitor these differences must be controlled and reproducible. Current methods to examine recovery of halophilic archaea from extreme stress are complicated by extended incubation times, nutrients not typically encountered in the environment, and other related limitations. Here we describe a method for assessing the function of genes during hypochlorite stress in the halophilic archaeon Haloferax volcanii that overcomes these types of limitations. The method was found reproducible and informative in identifying genes needed for H. volcanii to recover from hypochlorite stress.
Keywords: ArchaeaBackground
Accumulation of reactive species that are redox-active compounds usually leads to cytotoxic activity. Hypochlorite (HOCl) is a reactive species that is particularly cytotoxic as it reacts in vivo with low molecular weight inorganic molecules and organic molecules including the functional groups of lipids, proteins, carbohydrates, and nucleic acids (Panasenko et al., 2013). HOCl levels usually become more abundant during oxidative stress when there is an increase in molecular oxygen (O2) leading to incomplete reactions stopping at reactive oxygen species such as superoxide anions (O2•−), hydrogen peroxide (H2O2), and the highly reactive hydroxyl radical (OH•) (Loi et al., 2015). The rise in H2O2 levels leads to chlorination where H2O2 reacts with Cl– anions leading to the formation of HOCl (Winterbourn and Kettle, 2013).
H2O2 + Cl– + H+ → HOCl + H2O
HOCl ↔ OCl– + H+
While haloarchaea are of interest to understand how organisms thrive in harsh environments, mechanisms of HOCl stress response are best understood in pathogenesis, as neutrophils of mammalian innate immunity kill exogenous pathogens through an oxidative burst that includes HOCl production (Imlay, 2003, 2008, 2013; Ulfig and Leichert, 2021). Haloarchaea are microorganisms that often dominate hypersaline ecosystems where the concentration of NaCl is greater than in seawater (3.5% w/v NaCl) (Jones and Baxter, 2017). Of note in HOCl stress is the exceedingly high concentrations of chloride ions that are constantly present as exemplified by the hypersaline Lake Tyrrell that fluctuates from 4 to 5 M Cl– (Podell et al., 2014). These conditions lead to an increase in HOCl and consistent exposure of cells to cytotoxic agents. The study of haloarchaea is of interest to the scientific community, as these microorganisms can thrive in such harsh conditions. This inquiry has led to examining the mechanisms used by the haloarchaea to survive stress, including co-expression networks of coordinately regulated genes used to combat or control oxidative stress (Martinez-Pastor et al., 2017).
A common method used in biology to determine if a protein plays an important role in the cell is to delete the gene that encodes the protein of interest and observe the mutant strain phenotype. For example, here we are interested in comparing the growth rate and recovery of mutant and wild-type strains of the halophilic archaeon Haloferax volcanii before and after exposure to HOCl stress. For HOCl, supplementation of cultures with sodium hypochlorite (NaOCl) in aqueous solution leads to the spontaneous conversion of NaOCl into sodium hydroxide (NaOH) and HOCl, which can further dissociate into hydroxide (OH–) and hypochlorite (OCl–).
NaOCl + H2O ↔ NaOH + HOCl ↔ Na+ + OH– + H+ + OCl–
However, cultivating haloarchaea in a controlled environment can be difficult due to their demand for high concentrations of salts (e.g., > 2.5 M NaCl) and thermophilic temperatures (42–55 °C) (Robinson et al., 2005). Both environmental factors lead to evaporation, which exacerbates stress and can extend the time of cultivating cells for HOCl stress assays.
Here we describe a newly designed and improved method to examine the response of the haloarchaeon Haloferax volcanii to HOCl stress. Our initial analysis was conducted in a circular rotary shaker (Figure 1A) that yielded variable results. The inconstancy in results was likely due to the extensive incubation times required to detect cell recovery after HOCl exposure in minimal medium, as the conditions were microaerobic and dehydrating. These findings led to a demand for better growth conditions where the recovery of H. volcanii from HOCl stress could be monitored in a reproducible manner. Thus, a new protocol was developed that reduces the lag time of cellular recovery from HOCl stress and allows for experimental reproducibility by cultivating the cells using a mini rotator to improve aeration and redesigning the incubator to promote humidity (Figure 1B). The method is performed in minimal medium to avoid the complexity of antioxidants otherwise present in yeast extract and other common additions to undefined medium.
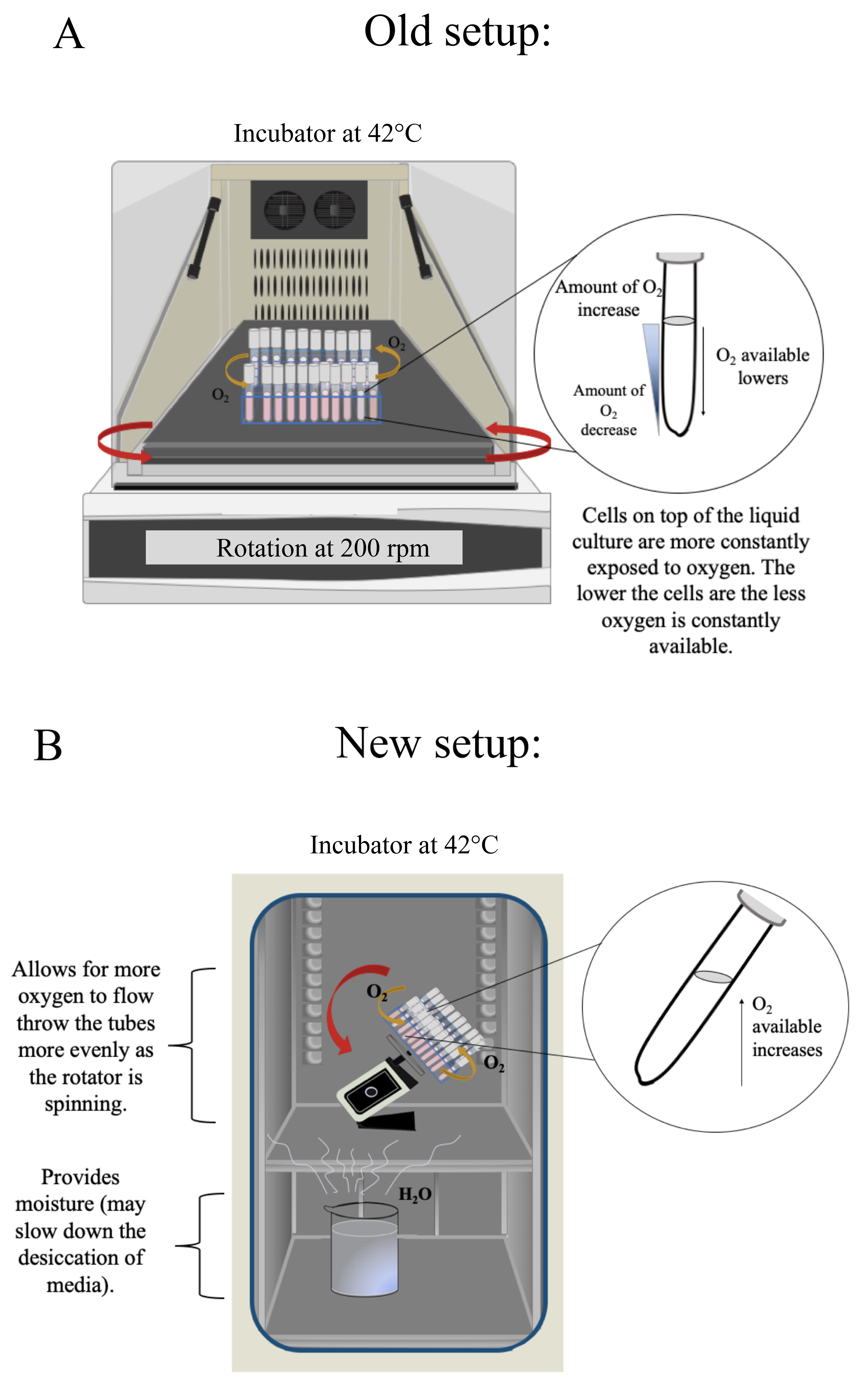
Figure 1. Diagram comparing two different setups for cell growth in culture tubes. A) Old setup with culture tubes oriented upright and agitated by rotary shaking, which causes an uneven distribution of oxygen and irregular growth patterns. B) New setup with culture tubes rotated at an angle to allow for more even aeration. A beaker is filled with water and included at the bottom of the incubator to enhance moisture.
Materials and Reagents
"Zipper" seal sample bags; thickness, 2 mil, and size, 6 × 9 in.
10 mL Sterile polystyrene disposable serological pipets (Genesee Scientific, catalog number: 12-104)
15 mL Conical centrifuge tubes, racked (polypropylene, Olympus Plastics, catalog number: 28-101)
2.5” Toothpicks, needle point, autoclavable (sterile) (LevGo, catalog number: 18250-NP)
200 µL XTIP4 barrier tips, low binding, racked, pre-sterilized (RNase and DNase free) (Genesee Scientific, catalog number: 24-712)
250 mL Erlenmeyer flasks (Pyrex, manufacturer number: 4980250/EMD) (Fisher Scientific, catalog number: S63271)
500 mL Erlenmeyer flasks (Pyrex, manufacturer number: 4980500/EMD) (Fisher Scientific, catalog number: S63273)
Disposable culture tubes, borosilicate glass 13 × 100 mm (Fisher Scientific, catalog number: 14-961-27)
Disposable plastic cuvettes semi-micro, 1.5 mL (Fisher Scientific, catalog number: 14955127)
Disposable petri dishes, polystyrene, sterile, semi-stackable (100 × 15 mm) (VWR, catalog number: 25384-088)
Plain PTFE stir bars, length: 70 mm; diameter: 10 mm (Fisher Scientific, catalog number: 16255801)
Plastic caps (to cap culture tubes) (DWK Life Sciences, manufacturer number: 7366013) (Fisher Scientific, catalog number: 14-957-91)
Poxygrid 96-place test tube rack; for 13–16 mm tubes (Bel-Art, catalog number: F18765-0001)
PYREX griffin low form 1 L beaker, double scale, graduated (Pyrex, manufacturer number: 10001L/EMD) (Fisher Scientific, catalog number: S14276)
Agar ash 2.0%–4.5% (Sigma-Aldrich, catalog number: A7002-250G)
Aluminum foil roll (used to cover opening for the Erlenmeyer flasks for sterilization and growing strains) (Fisher Scientific, catalog number: 01-213-105)
Ammonium chloride (NH4Cl) (Fisher Scientific, catalog number: A661-500)
Calcium chloride dihydrate (CaCl2·2H2O) (Fisher Scientific, catalog number: C79-500)
Copper sulfate pentahydrate (CuSO4·5H2O) (Sigma-Aldrich, catalog number: C3036)
D-Biotin (Fisher Bioreagents, catalog number: 58-85-5)
Glycerol for molecular biology (Fisher Bioreagents, catalog number: BP229-4)
Haloferax volcanii strains used including the H26 parent and SH125 mutant (ΔoxsR) that are previously described (Mondragon et al., 2022)
Iron sulfate heptahydrate (FeSO4·7H2O) (Alfa Aesar, catalog number: 7782-63-0)
L–Shaped cell spreader (Fisher Scientific, catalog number: 14-665-230)
Magnesium chloride hexahydrate (MgCl2·6H2O) (Sigma-Aldrich, catalog number: M0250-KG)
Magnesium sulfate heptahydrate (MgSO4·7H2O) (Fisher Science Education, catalog number: S25414A)
Manganese chloride tetrahydrate (MnCl2·4H2O) (Fisher Chemical, catalog number: M87-100)
Nanopure water (purified from Barnstead/Sybron Nanopure II 4-Module Water Purification System)
Novobiocin sodium salt (≥93% HPLC) (Sigma-Aldrich, catalog number: 74675-1G)
Potassium chloride (KCl) (Fisher Chemical, catalog number: P217-3)
Potassium phosphate dibasic anhydrous (K2HPO4) (Fisher Chemical, catalog number: P288-500)
Potassium phosphate monobasic (KH2PO4) (Fisher Chemical, catalog number: P285-500)
Potassium sulfate (K2SO4) (Fisher Chemical, catalog number: P304-500)
Sodium chloride (NaCl) certified ACS crystalline (Fisher Scientific, catalog number: S271-10)
Sodium hypochlorite (NaOCl) reagent grade, available chlorine 10%–15% (Sigma-Aldrich, catalog number: 425044-250mL)
Thiamine (Sigma cell culture, catalog number: T3902)
Tris-base (Fisher Bioreagents, catalog number: BP152-500)
Tryptone (Fisher Bioreagents, catalog number: BP1421-500)
Uracil, 99+% (Acros organics, catalog number: 66-22-8)
Yeast extract molecular genetics powder (Fisher Bioreagents, catalog number: BP1422-500)
Zinc sulfate heptahydrate (ZnSO4·7H2O) (Fisher Scientific, catalog number: 7446-20-0)
Concentrated salt water (SW) stock solution at 30% (w/v) (see Recipes)
GMM base (see Recipes)
Supplements (see Recipes)
0.5 M Potassium phosphate buffer (KPB), pH 7.5 for 100 mL total
1.5 mg/mL Uracil
Thiamine and biotin solution for 10.8 mL total
Hv-minimal salts for 12 mL total
Trace elements for 100 mL total
Volume of supplements to add to GMM base for total of 1 L of GMM + uracil (see Recipes)
For GMM + uracil agar plates a total of 500 mL (see Recipes)
For ATCC 974 medium plates a total of 500 mL (see Recipes)
Equipment
Cimarec basic stirring hotplate (ThermoFisher Scientific, catalog number: SP194715)
Genesys 40 visible spectrophotometer (ThermoFisher Scientific, catalog number: SP194715)
Genie 2 vortex mixer (Fisher Scientific, catalog number: 12-812)
Heratherm incubator (Thermo Scientific)
I24 Incubator shaker series (New Brunswick Scientific)
Mini rotator (20° angle, 2–80 rpm with disk and clamps, 120 V, Glas-Col Terre Haute, USA)
Pipette (10–100 µL, Rainin, pipet-lite)
Spectronic 20+ spectrophotometer (ThermoSpectronic, Filter: 600–950 nm)
Software
Microsoft Excel (version 16.16.27)
Procedure
Preparation of the strains
Using a sterile toothpick, pick a small aliquot of the frozen H. volcanii strain from a -80 °C glycerol stock and streak onto a glycerol minimal medium (GMM) plate. Repeat this for each strain including the parent, mutant, and complement strains. The stocks consist of stationary phase cells frozen in GMM with 20% (v/v) glycerol.
Incubate the plates for 5 days in a closed plastic zippered bag at 42 °C, until colonies appear.
Using a sterilized toothpick, collect five isolated colonies and put them into 20 mL of GMM in a 250 mL Erlenmeyer flask. This approach avoids dilution stress.
Grow the cells at 42 °C (200 rpm, rotary shaking) using the I24 Incubator shaker.
After 48 h of incubation, prepare a 1:5 dilution of each strain by sterile transfer of 0.2 mL culture and 0.8 mL fresh GMM in disposable plastic cuvettes. Measure the optical density at 600 nm (OD600) using a Genesys 40 visible spectrophotometer using uninoculated GMM to blank the instrument. Multiply the value obtained on the instrument by 5 to determine the original OD600. The cells should be at an OD600 of 0.8 to 1.0, which is equivalent to late log phase.
Dilute the cells with fresh GMM to an OD600 of 0.1 in 65 mL of final volume in a sterile 125 mL Erlenmeyer flask. For example, for cells at an OD600 of 0.89, mix 7.3 mL of cell culture with 57.7 mL of fresh GMM. Gently swirl the diluted cells to mix.
Using 10 mL sterile pipets, transfer aliquots (5 mL) of this cell suspension into 12 loosely capped sterile 13 × 100 mm glass culture tubes per strain type. Include tubes with 5 mL of fresh uninoculated GMM as negative controls.
Incubate the cell cultures for 12 h at 42 °C with aeration using a mini rotator (Glas-Col from Terre Haute in the USA) fitted with a culture rack (see details below). Use the maximum percent speed setting of 50 for the rotation. Perform the incubation in a Heratherm incubator.
Measure the OD600 of the culture by directly inserting the culture tube with the plastic cap into the Spectronic 20+ spectrophotometer (refer to Figure 2 to use the Spectronic 20+ spectrophotometer before measuring samples). Use the uninoculated controls to determine the background signal. The cells should reach log phase, which is estimated at an OD600 of 0.4–0.6. Make sure to close the lid of the instrument prior to taking the OD600 measurements.

Figure 2. Steps to set up the Spectronic 20+ spectrophotometer before measuring samples. 1) Turn on the spectrophotometer by turning the Power switch/Zero control to the right. 2) Make sure the filter lever is set to 600–900 nm and the red light is turned on. 3) The wavelength should be adjusted to 600 nm. 4) Allow for the spectrophotometer to warm up for 15 minutes. 5) Adjust the Power switch/Zero control to percent transmittance to 0. 6) Place into the sample compartment the 13 × 100 mm glass culture tube with only the medium (GMM) that will serve as a blank to 7) adjust the Transmittance/ Absorbance control (100%T/0A) to 100/0.Note: Do not remove plastic cap; the cap and the tube should both fit in the compartment.
Setup for incubation
Incubate the culture tubes in a Heratherm incubator with the following setup (Figure 3):
Place the culture tubes on a 96-place test tube rack attached to a mini rotator.
Place the mini rotator with the culture tubes on the top shelf of the incubator.
On the bottom shelf of the incubator, fill a 1 L beaker with distilled water up to the 1,000 mL mark to maintain moisture in the incubator.
Turn on rotator and slowly increase the maximum percent speed to 50.
Note: The same incubation setup is used after supplementation of NaOCl.
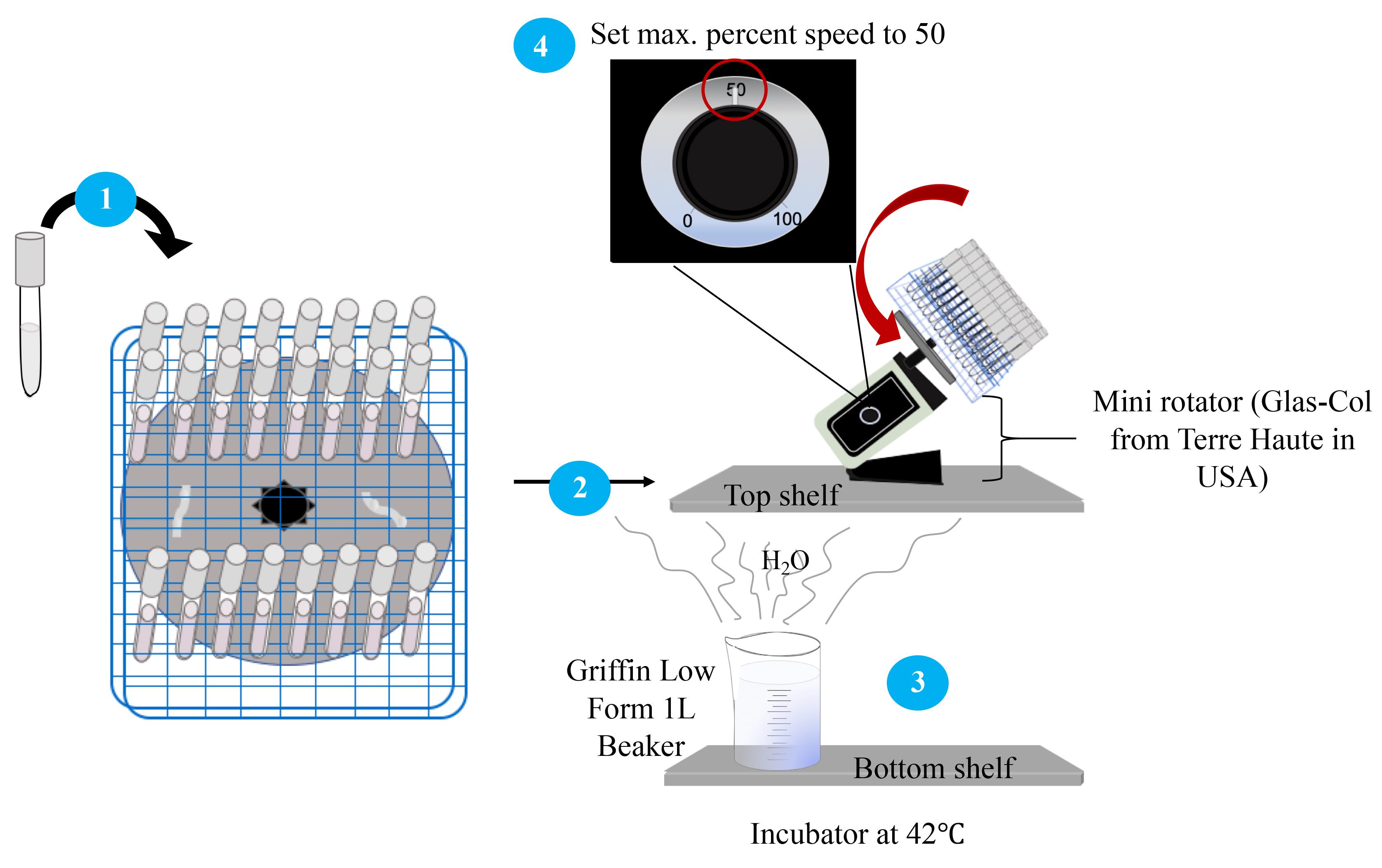
Figure 3. Schematic of the workflow for setting up for incubation. Circled numbers refer to the steps in the text.Preparation of NaOCl
Upon arrival of shipment, dispense the NaOCl reagent in 10 mL aliquots in 15 mL conical tubes and store at -80 °C (time of NaOCl storage was not monitored).
For each experiment, thaw 10 mL of frozen NaOCl stock to room temperature. Briefly vortex the solution once it is thawed.
Then, transfer 5 mL of the thawed NaOCl solution to a fresh 15 mL conical tube, which serves as the 16.2 M NaOCl stock.
Perform a serial dilution of the 16.2 M NaOCl stock to a final concentration of 2.025 M as follows: dispense 2.5 mL of nanopure water to three 15 mL conical tubes, perform a 1:2 dilution by transferring 2.5 mL of the 16.2 M stock to the first of the three tubes and mixing the solution to generate an 8.1 M stock, repeat the 1:2 dilutions two more times to a final concentration of 2.025 M (Figure 4A).

Figure 4. Schematic of the workflow for preparation and supplementation of NaOCl. A) Preparation of NaOCl; B) Supplementation of NaOCl. Circled numbers refer to the steps in the text.Supplementation of NaOCl (Figure 4B)
Randomly select half of the 12 culture tubes for each strain for supplementation with or without 5 mM NaOCl.
Supplement the first set of 6 culture tubes with 12.3 µL of nanopure water.
Attention: Vortex each individual cell culture tube immediately after addition of the water.
Supplement the second set of 6 cell cultures with 12.3 µL of 2.025 M NaOCl for a final concentration of 5 mM.
Attention: Vortex each individual cell culture tube immediately after addition of the NaOCl.
Monitoring of cell growth after supplementation with NaOCl
Remove all cell culture tubes from the mini rotator every 24 h of incubation at 42 °C after supplementing with NaOCl.
Examine the culture tubes to see if there are marks visible on the outside of the tube. If marks are present on the outside of the tubes, use a Kimwipe with ethanol to remove the marks. This action will allow the OD600 to be more accurately measured using the Spectronic 20+ spectrophotometer.
Place each culture tube directly into the spectrophotometer individually to determine the OD600 value.
If using the Spectronic 20+ spectrophotometer, make sure to close the lid before recording the OD600.
Repeat these OD600 measurements for 10 days.
Further analysis of strains that may exhibit two distinct responses in hypochlorite stress
Exposing cells to second round of hypochlorite stress (Figure 5).
For further analysis of the biological significance of a mutation, pool tubes with a similar growth pattern and treat as independent groups. For example, the mutant strain of this example is treated as three independent groups after the first stress assay. The first group consists of culture tubes pooled for the mock control. The second group is the pool of cell cultures that did not grow under 5 mM NaOCl. The third group is the pool of cell cultures that grew in the presence of 5 mM NaOCl.
Measure the OD600 for the two groups, by using the same measuring method from step A5.
Repeat steps A6 and A7.
Incubate the cell cultures for 10 h as in Section B.
Repeat all steps from Sections C–E.
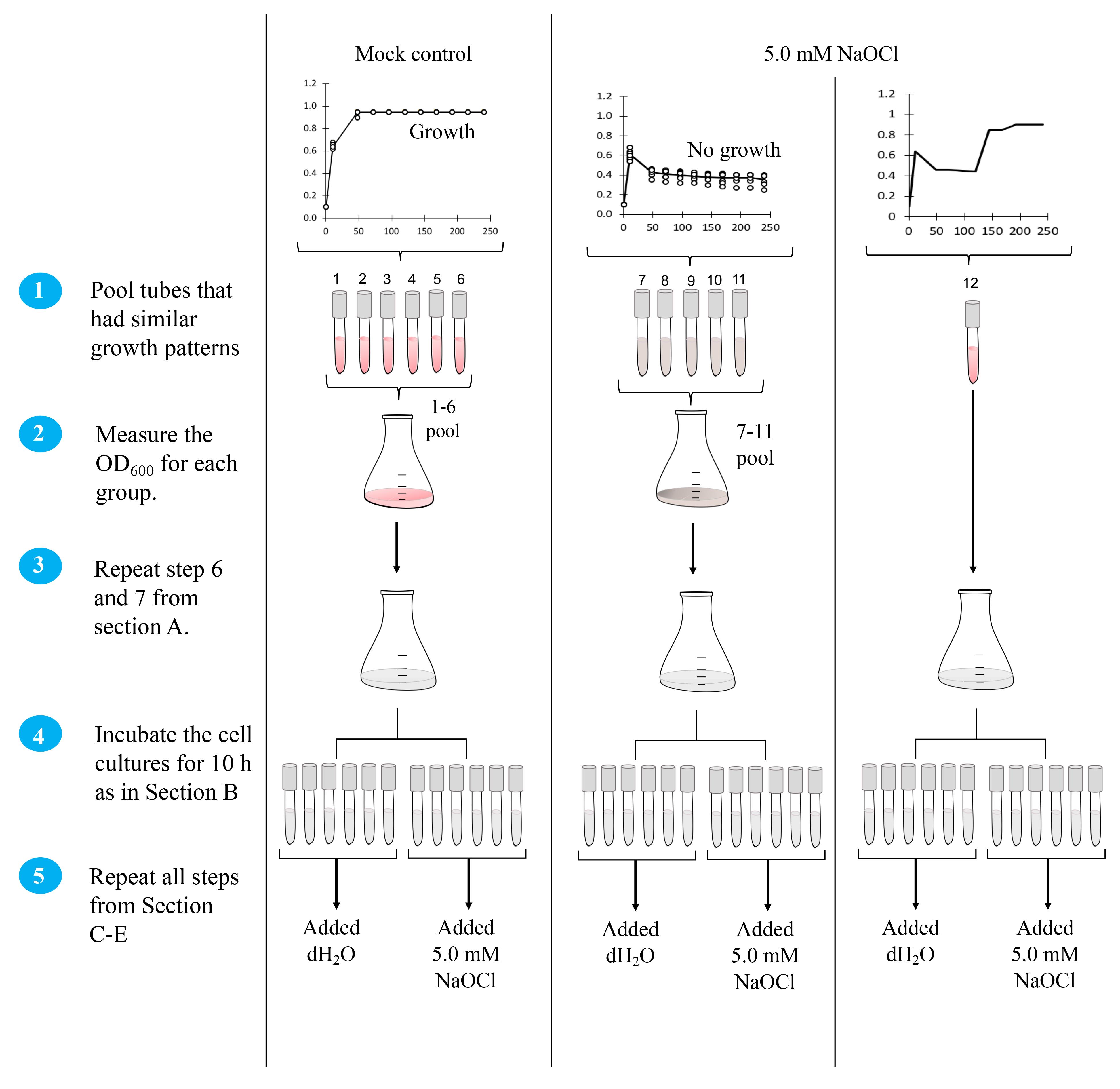
Figure 5. Schematic of the workflow for preparing strains for second incubation with hypochlorite. Circled numbers refer to the steps in the text.Plate count assay
After 10 days of monitoring the growth of each cell culture tube, randomly choose 2 or 3 technical replicates of each sample type to perform a serial dilution in 18% concentrated salt water (SW) (Figure 6).
First transfer 1 mL of cell culture into a sterilized 1.5 mL microcentrifuge tube.
Then, transfer 50 µL of this cell culture to a 1.5 mL microcentrifuge tube with 950 µL of 18% SW, and mix the sample by pipetting up and down.
Repeat step b three more times to generate a serial dilution series of 1:20, 1:400, 1:8,000, and 1:16,000.
Plate 100 µL of the diluted samples on ATCC 974 agar medium plates using an L-shaped cell spreader.
Allow the plates to air dry for ~15 min. Place the air-dried plates in a plastic bag and incubate at 42 °C for 5 days.
When the colonies are visible, choose a plate that appears to have between 30 and 300 colonies. Count the total colonies.
Calculate the CFUs per mL (CFUs/mL) of the sample by multiplying the number of colonies times the dilution factor of the counted plate. Note that because only 100 µL of cells were plated, a multiple of 10 must be included in the calculations of CFU per mL in addition to the original dilution factors (*DF) that range from 400 to 16,000.
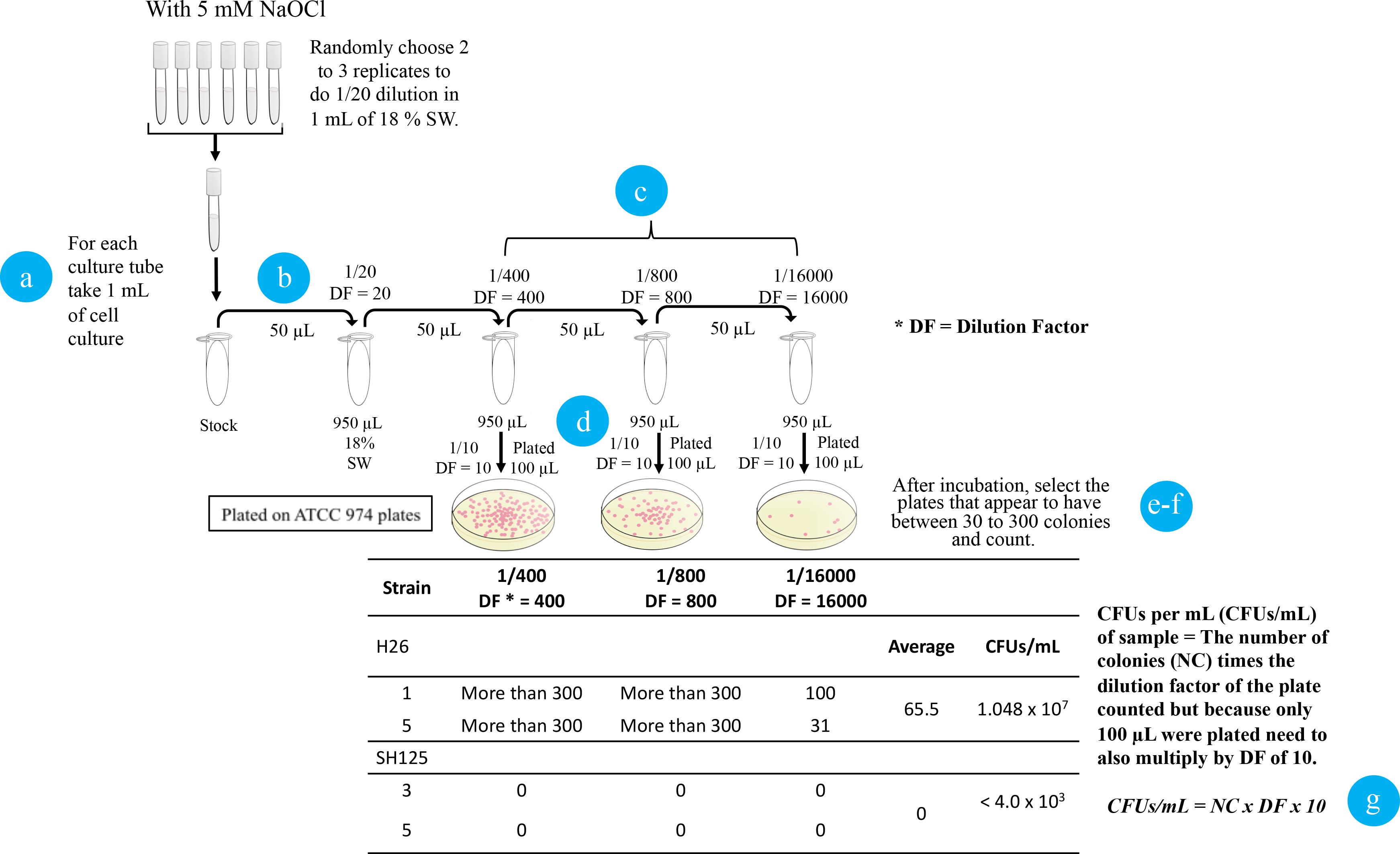
Figure 6. Schematic workflow for plate count assay. Circled numbers refer to the steps in the text.Data analysis
Record and plot the OD600 units over time (hours) in Microsoft Excel. Use the scatter plot to compare growth patterns of the different strains and culture conditions including the mock (H2O) and experimental sets (exposed to 5 mM NaOCl). An example is shown in Figure 7.
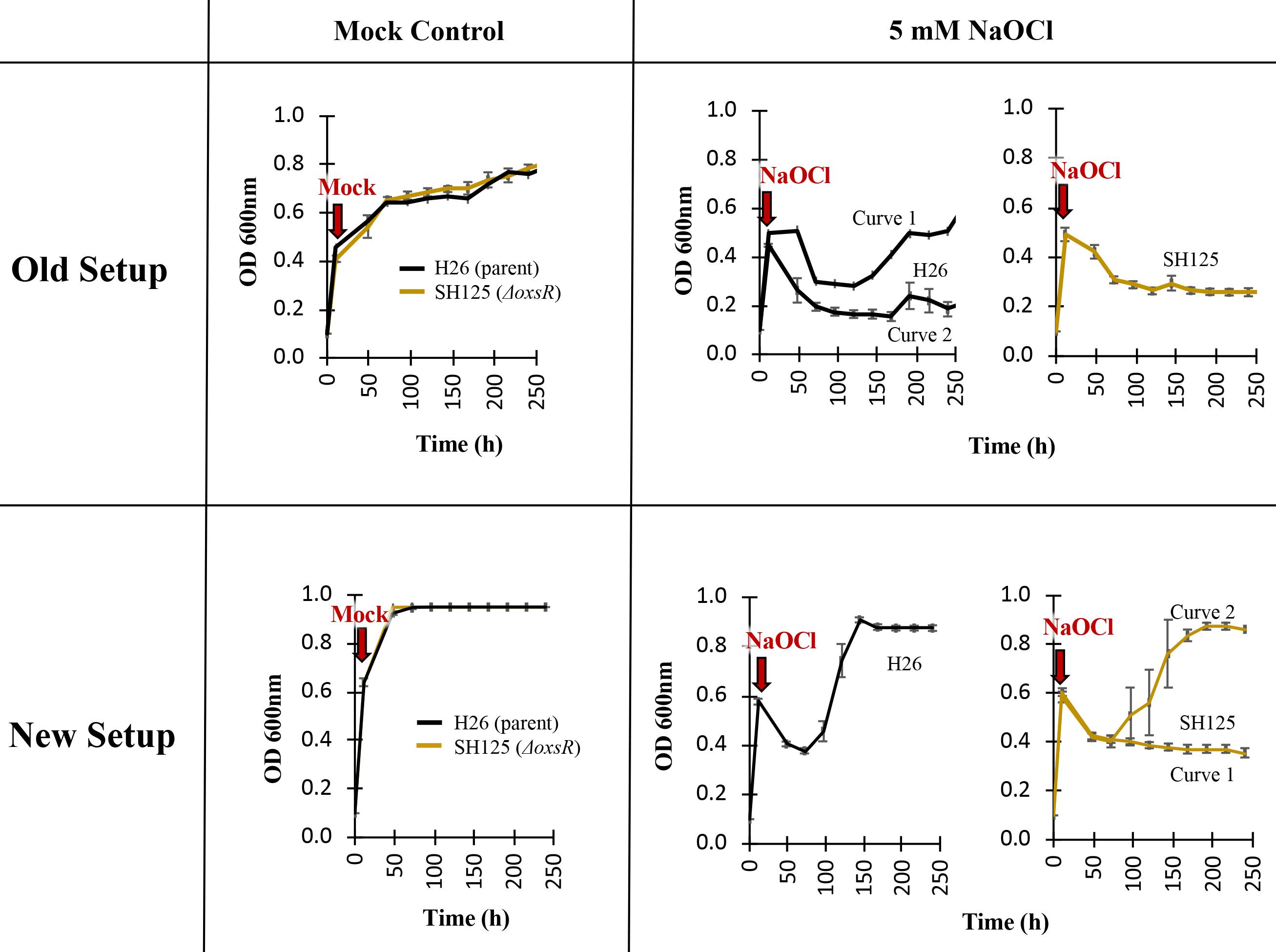
Figure 7. Scatter plots comparing the growth curve from “New setup” of Figure 1Calculate the growth rate and doubling time of each strain.
Plot time (h) against LN(OD of 600) to better identify the exponential phase (straight line). Choose the best two (or three) points that give the best straight line (Figure 8).
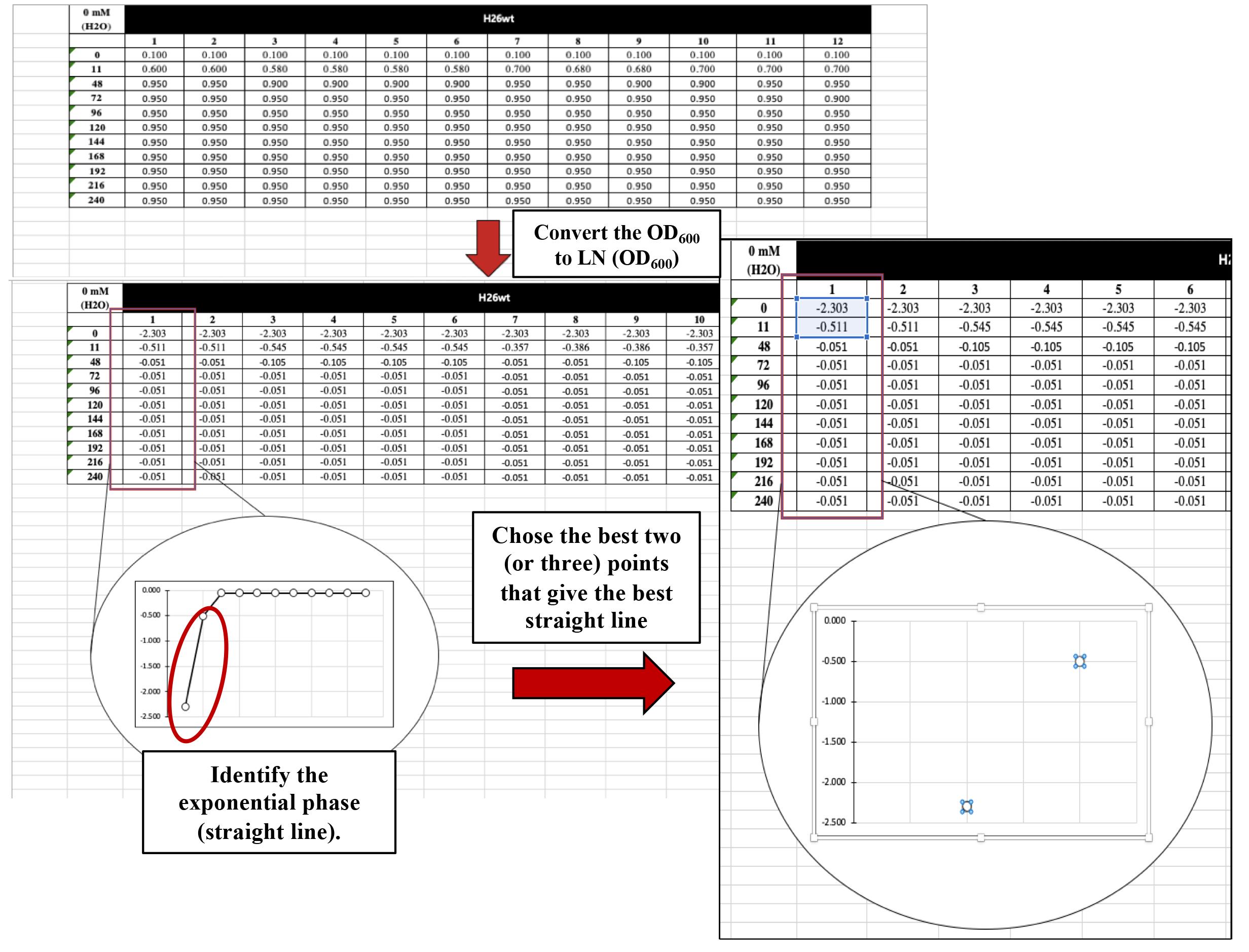
Figure 8. Plot time (h) against LN(OD of 600) to better identify the exponential phase (straight line)Calculate the growth rate by adding a linear trendline by going to Chart Design < Press Add Chart Element < Go to Trendline < Choose Linear < Go to More Trendline Options < Select Display Equation on chart (Figure 9). This will give you an equation for the trendline that includes the slope of the line that is the growth rate in generation per hour (gen/h).
Note: The growth rate for each 12 technical replicates is calculated separately. Then, the average growth rate (µ) is calculated based on all growth rates from each replicate.
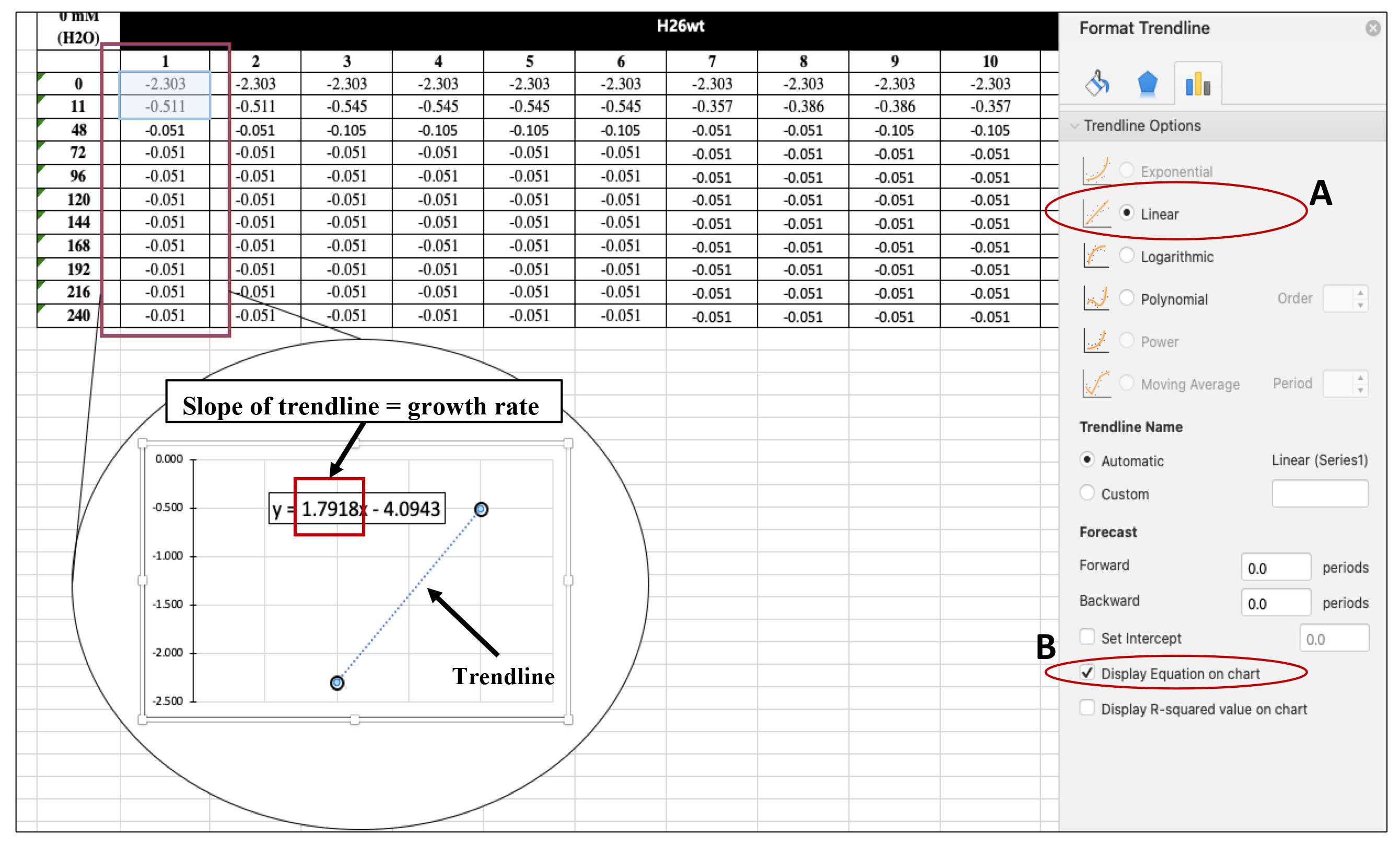
Figure 9. Finding the growth rate. After selecting More Trendline Options make sure the option is set to Linear (A) and select Display Equation on chart (B). The slope of the trendline is the growth rate in generation per hour (gen/h).To calculate the doubling time (k), use the average growth rate in the following formula:
k = 1/µ
Calculate the standard deviation (±) for each average growth rate and doubling time.
For the strains supplemented with 5 mM NaOCl, the exponential phase will be seen much later than the mock control (H2O). As shown in Figure 10, the two points that made a straight line were selected after adding the 5 mM of NaOCl.
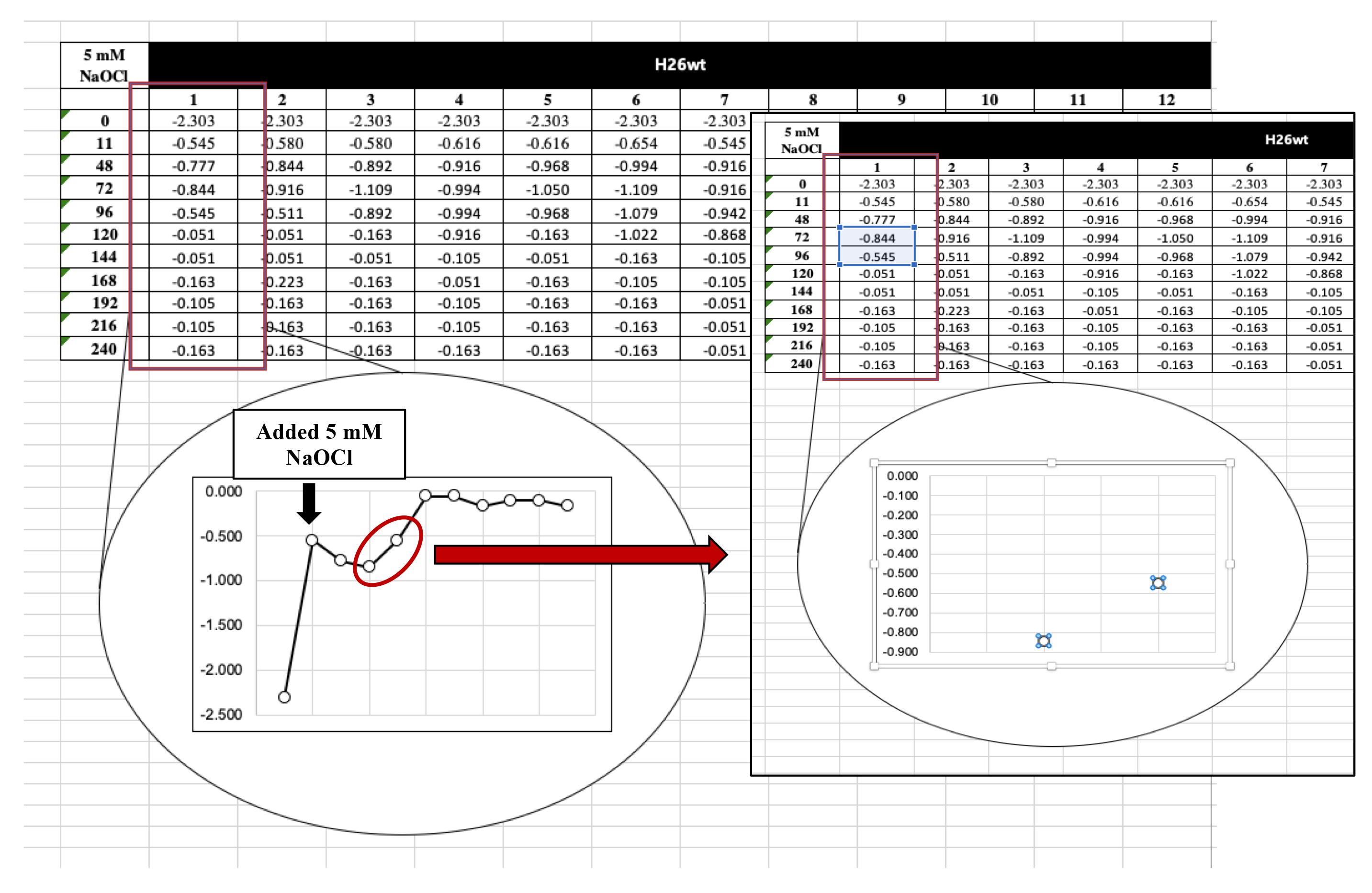
Figure 10. Finding the points to make trendline for strains after adding 5 mM of NaOCl. Make sure to select points that make a straight line.Calculating Area Under the Curve (A.U.C.) by using Trapezoidal Rule
For each technical replicate find the area under the curve by finding the area of each trapezoid. For example, first trapezoid is between x = 1 and x = 2 by using the following formula (Figure 11A):
area = (y1 + y2)/2*(x2 - x1)
Calculate the area for the other trapezoids (Figure 11B).
Then, add the areas of all trapezoids to find the sum that will be the A.U.C. for the growth curve (Figure 11C).
Repeat the above steps for each technical replicate for each strain, then find the average and standard error.
For helpful reference to guide A.U.C. calculations see Github link https://rdrr.io/github/leonpheng/lhtool/man/AUC.html.
Plot the average of A.U.C. of each strain in the bar graph and add the technical replicates individually and convert them to a scatter plot (Figure 11D). Use the average for each experiment to calculate if it has equal variance by F-test. If so, use the two-tailed Student’s t-test analysis to find if there are significant differences between strains.
Note: If you find that the variances are unequal, you can do the two-tailed, unpaired Welch's t-test instead of the Student's t-test.
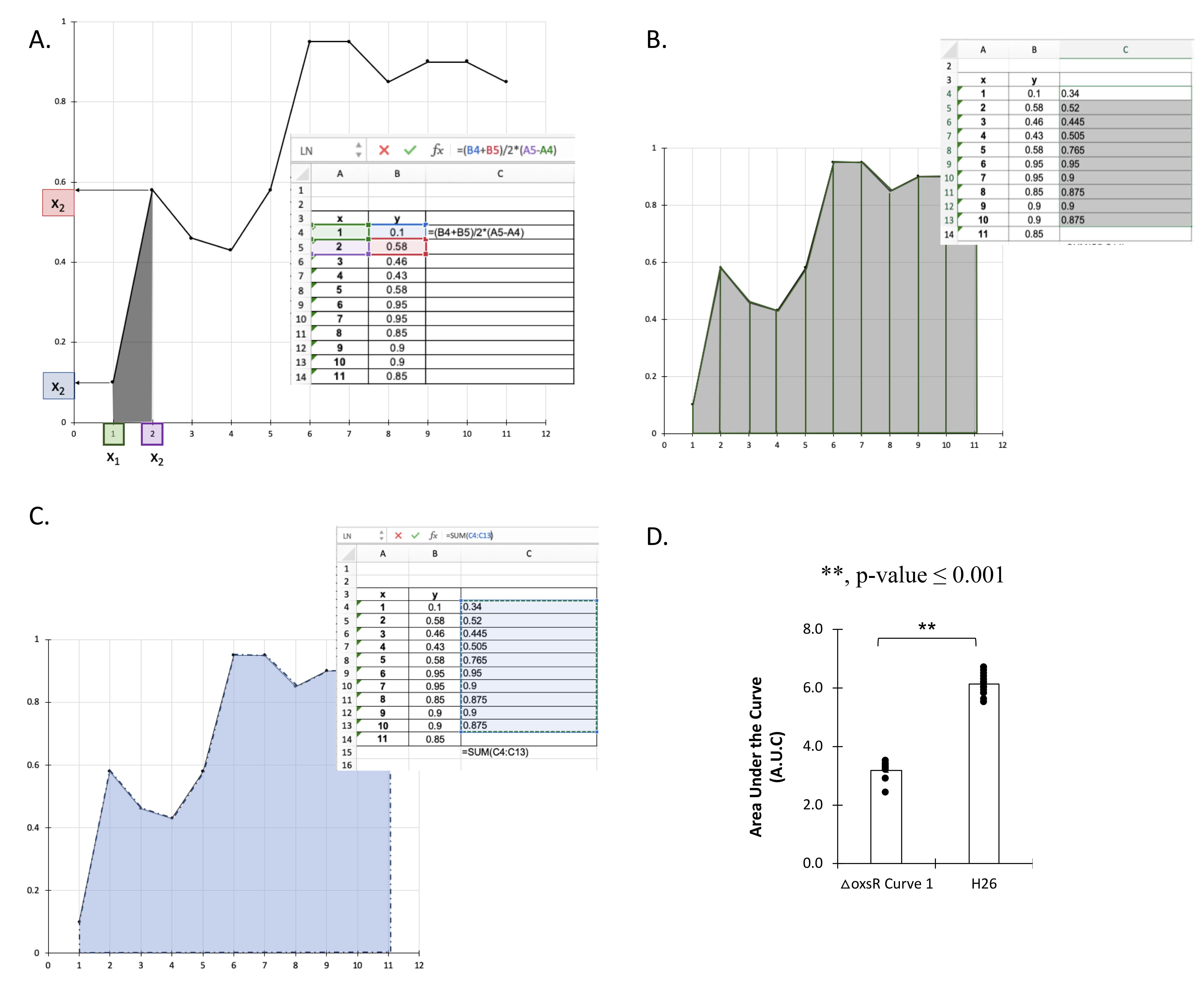
Figure 11. Calculating the Area Under the Curve (A.U.C.) for a Haloferax volcanii strain (H26). A) Find the area of the trapezoid by using area = (y1 + y2)/2*(x2 - x1). B) Calculate the area for the other trapezoids. C) Find the sum of all areas to find the area under the curve. D) Example of bar graph showing the A.U.C. and the significant differences between strains by using the Student’s t-test analysis.
Recipes
The first set of recipes are for glycerol minimal media plus uracil (GMM + uracil) and were modified from the Halohandbook (Dyall-Smith, 2009).
Concentrated salt water (SW) stock solution at 30% (w/v) (1 L)
Use nanopure water as approximately 70% of the total volume of the solution (~700 mL), and warm it on a hot plate.
Note: A microwave could also be used to warm up the water.
When the water gets warm, add the following salts one by one allowing each to completely dissolve:
NaCl 240 g
MgCl2·6H2O 30 g
MgSO4·7H2O 35 g
KCl 7 g
1 M Tris base (with pH of 7.5, the pH is adjusted with HCl) 20 mL
GMM base (1 L)
Nanopure water 20.4 mL
30% SW 647.2 mL
1 M Tris base (with pH of 7.5, the pH is adjusted with HCl) 32.4 mL
Autoclave solution before use.
Supplements
0.5 M KPB, pH 7.5 for 100 mL total
1 M K2HPO4 83.4 mL
1 M KH2PO4 16.6 mL
pH should be approximately 7.5.
Note: When mixing compounds, add in small volumes of the 1 M KH2PO4 to the 1 M K2HPO4 while monitoring the pH. When all 1 M KH2PO4 has been added, the pH should be approximately 7.5. Do not add HCl or NaOH to adjust pH to 7.5.
Filter the solution before use and store at room temperature.
1.5 mg/mL Uracil
The Halohandbook suggests making a 50 mg/mL solution of uracil by dissolving in DMSO. However, DMSO is a reactive compound that may expose the cells to oxidative stress. To better control the hypochlorite stress conditions, uracil is dissolved at 1.5 mg/mL in nanopure water overnight at 42 °C with rotary shaking at 200 rpm using an I24 Incubator shaker. After ~16 h the dissolved uracil is filtered and stored at room temperature.
1 mg/mL of Novobiocin
Novobiocin is used to select for cells transformed with plasmids that harbor the novobiocin resistance marker. Plasmids are often used to carry the wild-type gene of interest and demonstrate that the mutation can be complemented. Dissolve the novobiocin at 10 mg/mL in nanopure water and then dilute 10-fold to a working stock of 1 mg/mL. Store excess reagent in aliquots at -20 °C for future use.
Thiamine and biotin solution for 10.8 mL total
1 mg/mL thiamine 9.6 mL
1 mg/mL D-biotin 1.2 mL
Filter solution before use and store at 4 °C.
Hv-minimal salts for 12 mL total
1 M NH4Cl 5.0 mL
0.5 M CaCl2 6.0 mL
Trace elements 1.0 mL
Fill up to 12 mL with nanopure water.
Filter the solution. Make Hv-minimal salts fresh before use.
Trace elements for 100 mL total
Dissolve the following elements in ~70 mL of nanopure water:
MnCl2·4H2O 36 mg
ZnSO4·4H2O 44 mg
FeSO4·7H2O 230 mg
CuSO4·5H2O 5 mg
Fill up to 100 mL with nanopure water and filter before storing at 4 °C.
Volume of supplements to add to GMM base for total of 1 L of GMM + uracil
Add the calculated amount of GMM base to a sterile flask (total GMM minus total volume of all supplements). Then add the following volumes of each supplement:
1 M glycerol 20 mL
0.5 M KPB, pH 7.5 1.9 mL
1.5 mg/mL uracil 33.4 mL
Thiamine and biotin solution 0.882 mL
Hv-minimal salts 11.8 mL
Note: For the complement and empty vector strains, GMM + uracil is supplemented with 0.3 µg/mL of novobiocin by using a stock solution of 1 mg/mL of novobiocin.
For GMM + uracil agar plates a total of 500 mL
Autoclave the volume of GMM base (total GMM minus total volume of all supplements) with 7.5 g of agar with a stirring rod. Cool the solution to 60°C for 30 min. After cooling the medium, add the same supplements while the GMM base with agar are stirring on the plate:
1 M glycerol 10 mL
0.5 M KPB, pH 7.5 0.950 mL
1.5 mg/mL uracil 16.7 mL
Thiamine and biotin solution 0.441 mL
Hv-minimal salts 5.9 mL
Pour the medium in petri dishes (~20 mL for each plate). Allow the medium to solidify and cool before use.
Note: For the complement and empty vector strains, GMM + uracil plates are supplemented with 0.3 µg/mL of novobiocin by spreading the volume of 1 mg/mL of novobiocin onto the dry GMM + uracil plate. The novobiocin solution is allowed to dry on the plates before inoculating the strain from the -80 °C glycerol stock.
The following recipe is for 1 L of ATCC 974 medium plates that was modified:
Nanopore water ~700 mL
NaCl 125 g
MgCl2·6H2O 50 g
K2SO4 5 g
CaCl2·2H2O 0.134 g
Tryptone 5 g
Yeast extract 5 g
Adjust to pH 6.8 with 1 M KOH
Agar 15 g
Fill up to 1,000 mL with nanopore water and autoclave.
Pour the medium in petri dishes (~20 mL for each plate). Allow the medium to solidify and cool before use.
Note: For strains that contain plasmids, novobiocin with the same concentration as for GMM + uracil was used.
Acknowledgments
Funds awarded to JMF and AS to develop systems biology tools were through the Bilateral NSF/BIO-BBSRC program (NSF 1642283). Funds awarded to JMF to determine archaeal redox regulation were through the U.S. Department of Energy, Office of Basic Energy Sciences, Division of Chemical Sciences, Geosciences and Biosciences, Physical Biosciences Program (DOE DE-FG02-05ER15650) and to provide evolutionary insight in biological systems were through the National Institutes of Health (NIH R01 GM57498).
Competing interests
The Authors declare that there are no conflicts of interest.
References
- Dyall-Smith, M. (2009). The Halohandbook: Protocols for Halobacterial Genetics v.7.2.
- Imlay, J. A. (2003). Pathways of oxidative damage. Annu Rev Microbiol 57: 395-418.
- Imlay, J. A. (2008). Cellular defenses against superoxide and hydrogen peroxide. Annu Rev Biochem 77: 755-776.
- Imlay, J. A. (2013). The molecular mechanisms and physiological consequences of oxidative stress: lessons from a model bacterium.Nat Rev Microbiol 11(7): 443-454.
- Jones, D. L. and Baxter, B. K. (2017). DNA repair and photoprotection: Mechanisms of overcoming environmental ultraviolet radiation exposure in halophilic archaea. Front Microbiol 8: 1882.
- Loi, V. V., Rossius, M. and Antelmann, H. (2015). Redox regulation by reversible protein S-thiolation in bacteria. Front Microbiol 6: 187.
- Martinez-Pastor, M., Tonner, P. D., Darnell, C. L. and Schmid, A. K. (2017). Transcriptional regulation in archaea: From individual genes to global regulatory networks. Annu Rev Genet 51: 143-170.
- Mondragon, P., Hwang, S., Kasirajan, L., Oyetoro, R., Nasthas, A., Winters, E., Couto-Rodriguez, R. L., Schmid, A. and Maupin-Furlow, J. A. (2022). TrmB family transcription factor as a thiol-based regulator of oxidative stress response. mBio: e0063322.
- Panasenko, O. M., Gorudko, I. V. and Sokolov, A. V. (2013). Hypochlorous acid as a precursor of free radicals in living systems.Biochemistry (Mosc) 78(13): 1466-1489.
- Podell, S., Emerson, J. B., Jones, C. M., Ugalde, J. A., Welch, S., Heidelberg, K. B., Banfield, J. F. and Allen, E. E. (2014). Seasonal fluctuations in ionic concentrations drive microbial succession in a hypersaline lake community. ISME J 8(5): 979-990.
- Robinson, J. L., Pyzyna, B., Atrasz, R. G., Henderson, C. A., Morrill, K. L., Burd, A. M., Desoucy, E., Fogleman, R. E., Naylor, J. B., Steele, S. M., et al. (2005). Growth kinetics of extremely halophilic Archaea (family Halobacteriaceae) as revealed by Arrhenius plots. J Bacteriol 187(3): 923-929.
- Ulfig, A. and Leichert, L. I. (2021). The effects of neutrophil-generated hypochlorous acid and other hypohalous acids on host and pathogens. Cell Mol Life Sci 78(2): 385-414.
- Winterbourn, C. C. and Kettle, A. J. (2013). Redox reactions and microbial killing in the neutrophil phagosome.Antioxid Redox Signal 18(6): 642-660.
Article Information
Copyright
© 2022 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Mondragon, P., Hwang, S., Schmid, A. and Maupin-Furlow, J. A. (2022). Hypochlorite Stress Assay for Phenotypic Analysis of the Halophilic Archaeon Haloferax volcanii Using an Improved Incubation Method and Growth Monitoring. Bio-protocol 12(22): e4557. DOI: 10.21769/BioProtoc.4557.
Category
Microbiology > Microbial physiology > Stress response
Environmental science
Biological Sciences > Microbiology
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










