- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Femtoliter Injection of ESCRT-III Proteins into Adhered Giant Unilamellar Vesicles
(*contributed equally to this work) Published: Vol 12, Iss 4, Feb 20, 2022 DOI: 10.21769/BioProtoc.4328 Views: 3741
Reviewed by: Anonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Fluorescence-based Heme Quantitation in Toxoplasma Gondii
Amy Bergmann and Zhicheng Dou
Jun 20, 2021 4031 Views

Artificial Metalloenzymes in Artificial Sanctuaries Through Liquid–Liquid Phase Separation
Kaixin Wang [...] Tong Wu
Oct 5, 2025 1594 Views
Dynamic Mapping of RNA-Binding Proteins During Bacillus subtilis Sporulation Using Orthogonal Organic Phase Separation
Thomas Kaboré and Clémentine Delan-Forino
Mar 5, 2026 174 Views
Abstract
The endosomal sorting complex required for transport (ESCRT) machinery mediates membrane fission reactions that exhibit a different topology from that observed in clathrin-coated vesicles. In all of the ESCRT-mediated events, the nascent vesicle buds away from the cytosol. However, ESCRT proteins are able to act upon membranes with different geometries. For instance, the formation of multivesicular bodies (MVBs) and the biogenesis of extracellular vesicles both require the participation of the ESCRT-III sub-complex, and they differ in their initial membrane geometry before budding starts: the protein complex acts either from outside the membrane organelle (causing inward budding) or from within (causing outward budding). Several studies have reconstituted the action of the ESCRT-III subunits in supported bilayers and cell-sized vesicles mimicking the geometry occurring during MVBs formation (in-bud), but extracellular vesicle budding (out-bud) mechanisms remain less explored, because of the outstanding difficulties encountered in encapsulation of functional ESCRT-III in vesicles. Here, we provide a different approach that allows the recreation of the out-bud formation, by combining giant unilamellar vesicles as a membrane model and a microinjection system. The vesicles are immobilized prior to injection via weak adhesion to the chamber coverslip, which also ensures preserving the membrane excess area required for budding. After protein injection, vesicles exhibit outward budding. The approach presented in this work can be used in the future to disentangle the mechanisms underlying ESCRT-III-mediated fission, recreating the geometry of extracellular bud production, which remains a challenge. Moreover, the microinjection methodology can be also adapted to interrogate the action of other cytosolic components on the encapsulating membranous organelle.
Graphic abstract:
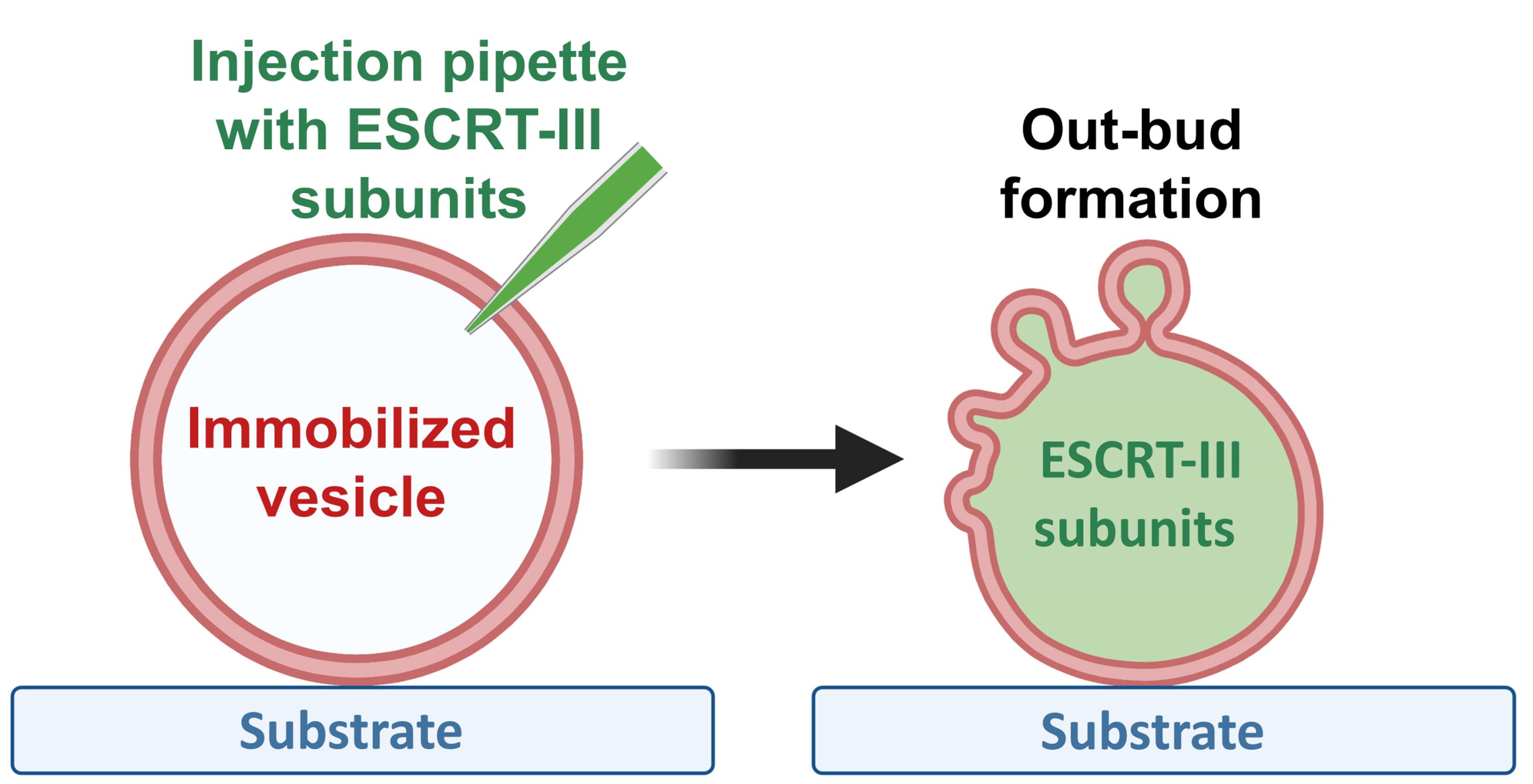
Out-bud formation after ESCRT-III protein injection into GUVs
Background
Extracellular vesicles (EVs) are defined as cell-derived vesicles confined by a lipid bilayer. They participate in cellular disposal and in intercellular communication by delivering antigens, genetic material, and lipids to recipient cells (Raposo and Stahl, 2019). EV secretion seems to be an ubiquitous process present throughout all domains of life and, in most of the cases, it is related to normal physiological conditions (Herrmann et al., 2021). However, a considerable increase in EV number is observed during pathogenic processes including cancer (Wolfers et al., 2001; Tao et al., 2019; Clos-Garcia et al., 2018; Xu et al., 2018), hereditary α-tryptasemia (Glover et al., 2019), multiple sclerosis (Moyano et al., 2016), cytotoxic drug intoxication (Keklikoglou et al., 2019), and parasitic diseases, among which cerebral malaria has been widely characterized (Combes et al., 2004; Schofield and Grau, 2005; Campos et al., 2010; Mfonkeu et al., 2010; Nantakomol et al., 2011; Martin-Jaular et al., 2011). EVs can be classified into two major classes, depending on their size and origin: exosomes, with a typical diameter of 30–150 nm, and microvesicles, that have a diameter of 100–1,000 nm (Raposo and Stahl, 2019). Whereas microvesicles are shed by outward budding of the plasma membrane, exosomes are generated by the fusion of multivesicular bodies (MVBs) with the plasma membrane, followed by the release of intraluminal vesicles (ILVs) (van Niel et al., 2018). EV populations are considered highly heterogeneous both in content and in size, probably due to the different pathways along which they originate from (Raposo and Stahl, 2019), making it difficult to resolve the mechanisms associated with cargo-packing and specific activity during cell signaling. Understanding the molecular processes that govern their biogenesis, via employing simple mimetic systems, could provide a clue to solve their mechanism of action, and therefore, help to understand the pathophysiology of certain diseases.
One of the pathways for EV biogenesis is orchestrated by the endosomal sorting complex required for transport (ESCRT) machinery, which comprises several protein subunits organized into four different complexes (ESCRT-0, -I, -II and -III), and the accessory Vps4 complex (reviewed in Vietri et al., 2020). Typically, ESCRT members are recruited to the endosomal membrane in a stepwise manner, with subsequent activation of the different ESCRT complexes. Two non-canonical pathways have been also identified: 1) activation of ESCRT-I by Bro1, which functions as an ubiquitin acceptor (Tang et al., 2016), and 2) ESCRT-III activation by Alix, which mediates the ubiquitin-independent, but ESCRT-III-dependent endocytosis (Dores et al., 2012). On the other hand, the ESCRT machinery has also been implicated in the production of nano-sized vesicles that are enriched in cell surface proteins, reflecting its participation during microvesicle formation (Nabhan et al., 2012; Wang and Lu, 2017).
ESCRT-III recruitment in Plasmodium falciparum operates through a non-canonical pathway in which PfBro1 activates PfVps32 and PfVps60, both ESCRT-III members, triggering EV biogenesis (Avalos-Padilla et al., 2021b). Involving an intracellular parasite, the study of this process is problematic. Moreover, the knockdown or deletion of ESCRT genes in other organisms results in the formation of aberrant structures that lack ILVs (Doyotte et al., 2005; Nickerson et al., 2006). To address these difficulties, membrane models have been widely used to analyze in vitro ESCRT-III-mediated events. In this direction, giant unilamellar vesicles (GUVs) (Dimova and Marques, 2019; Dimova, 2019) combined with recombinant ESCRT proteins have become an established platform to examine the formation of MVBs (Im et al., 2009; Avalos-Padilla et al., 2018 and 2021a; Booth et al., 2019;Alqabandi et al., 2021). To mimic the geometry occurring in this process, ESCRT components are introduced in the vesicle surroundings. The proteins induce membrane invaginations towards the vesicle interior, which can lead to the formation of ILVs, connected to the mother membrane through a thin neck, and the final cleavage of the neck results in the formation of MVB-like GUVs.
The out-budding processes (as in the formation of microvesicles shed by the plasma membrane) exhibit a reverse budding topology, compared to that of MVB formation. Thus, to explore such processes, the ESCRT units have to act from within the vesicle model. In other words, the proteins have to be introduced into the GUVs’ lumen. One approach to accomplish this consists of encapsulating ESCRT proteins inside GUVs, by forming the vesicles in the presence of the proteins; using this strategy, nanotubes with the correct topology for scission were pulled and subsequently cleaved (Schöneberg et al., 2018). However, under these conditions, one cannot observe the vesicle response upon immediate interaction with the proteins, as it relies on ATP photo-uncaging. To evade these drawbacks, we have designed an approach in which pre-formed GUVs encapsulating the buffer necessary for protein activity are injected with the ESCRT-III proteins. With this technique, we are able to observe in real time the dynamics of out-bud formation (mimicking the process driven during EV biogenesis), and to evaluate the effects specific to a particular protein.
Injection approaches in GUVs have been applied previously (Wick et al., 1996; Hurtig and Orwar, 2008; Lefrançois et al., 2018). In isolated GUVs, it is important to ensure control over the vesicle volume and area. In particular, the injection of isotonic solutions can pull out the excess membrane area needed for deformation, which would then prohibit outward budding. Thus, we adapted this protocol, by employing osmolarities of the injected solutions which lead to vesicle deflation, to create excess vesicle area for deformation and budding. Furthermore, isolated vesicles need to be immobilized to facilitate puncturing the membrane without displacing and losing them. In previous work, the immobilization was ensured by working with GUVs which were directly formed on the substrate for GUV swelling. However, such vesicles are typically connected to other GUVs and structures, thus not ensuring area/volume conservation. Here, we employed biotin-avidin-based adhesion, using biotinylated lipids in the vesicle and an avidin-coated substrate to which the GUVs were fixed as proposed earlier (Maan et al., 2018). Successful injection of ESCRTs and further registration of the functionality of the proteins, namely formation of out-buds, requires fine adjustment of the adhesion level. On one hand, strong adhesion stabilizes the vesicles during the puncturing procedure, while on the other hand, it increases the membrane tension while consuming the area available for deformation. The latter effect limits the ability of the membrane to bend and thus hinders budding. We thus optimized the avidin surface concentration to ensure mild adhesion. Another important aspect to consider is the buffer in which proteins remain active: in the case of ESCRT-III proteins, the buffer used was 150 mM NaCl and 25 mM Tris-HCl, pH 7.4 (~325 mOsmol/kg). This high salt concentration hamperedGUVs growing through the standard electroformation protocol (Angelova and Dimitrov, 1986). Despite some recent developments of this protocol aiming at application to solutions of high ionic strength (Montes et al., 2007; Pott et al., 2008; Li et al., 2016; Lefrançois et al., 2018), it is clear that the incorporation of negatively charged lipids imposes strong limitations, resulting in poor vesicle quality and small size (as confirmed by our own tests, data not shown). To overcome these drawbacks, we used the gel-assisted method (Weinberger et al., 2013), in which we were able to grow vesicles encapsulating the protein buffer. It must be noted that this method may lead to the incorporation of polymers into the formed vesicles, thus altering their mechanical properties (Dao et al., 2017). Indeed, occasionally we observed vesicles with denser content (presumably resulting from encapsulated hydrogel). For the work here, we always selected “clean” GUVs with no visible alterations in their membranes and, as detailed later, substantiate the results with control experiments where vesicles were injected with protein-free buffer. As we are working with P. falciparum ESCRT-III subunits, the GUV lipid composition was selected to mimic the inner leaflet of the red blood cell plasma membrane (Virtanen et al., 1998). However, this can be modified depending on the system. We also demonstrate the injection and outward budding process for GUVs injected with another ESCRT-III system, namely the protozoan parasite responsible for amoebiasis, Entamoeba histolytica, whose characterization in GUVs has been previously reported (Avalos-Padilla et al., 2018), using a suitable membrane composition. Finally, as mentioned above, to deflate the GUVs, and thus to ensure that the vesicles exhibit excess membrane area needed for the formation of the out-buds, the injected proteins were kept in a 0.8× buffer (prepared by a direct dilution in Milli-Q water of the 1× protein buffer, see Recipe 4). As a control, upon injection of GUVs with the same volume and osmolarity of protein-free solution as in the experimental set-up, no out-buds appeared, demonstrating the validity of our approach.
Materials and Reagents
Lipids
1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine [POPC] (Avanti Polar Lipids, catalog number: 850457)
1-palmitoyl-2-oleoyl-sn-glycero-3-phospho-L-serine [POPS] (Avanti Polar Lipids, catalog number: 840034)
Cholesterol (ovine) [chol] (Avanti Polar Lipids, catalog number: 700000)
1,2-dioleoyl-sn-glycero-3-phospho-(1’-myo-inositol-3’-phosphate) [PI(3)P] (Avanti Polar Lipids, catalog number: 850150)
1,2-distearoyl-sn-glycero-3-phosphoethanolamine-N-[biotinyl(polyethyleneglycol)-2000] [DSPE-PEG 2000 Biotin] (Avanti Polar Lipids, catalog number: 880129)
1,2-dipalmitoyl-sn-glycero-3-phosphoethanolamine-N-(lissamine rhodamine B sulfonyl) [DPPE-Rhodamine] (Avanti Polar Lipids, catalog number: 810158)
Note: All lipids are dissolved in chloroform at a final concentration of 10 mg·mL-1 and can be stored at -20°C for up to one month.
Reagents
Avidin from egg white (Sigma-Aldrich, catalog number: A9275)
Bovine serum albumin [BSA] (Sigma-Aldrich, catalog number: A8806)
Biotinylated BSA (Thermo Fisher Scientific, catalog number: 29130)
Chloroform (Merck, Supelco, catalog number: 288306)
Ethanol absolute (Merck, catalog number: 1009831011)
Methanol (Merck, catalog number: 34860)
Milli-Q® water (Millipore system)
Polyvinyl alcohol [PVA] (Approximate MW 145,000 g·mol-1, Merck, catalog number: 814894)
Polyethylene glycol fluorescein isothiocyanate [PEG-FITC] (Nanocs, catalog number: PG1-FC-40k)
NaCl (Sigma-Aldrich, catalog number: S7653)
Trizma® hydrochloride (Sigma-Aldrich, catalog number: T6666)
Purified recombinant PfBro1 and PfVps32 from P. falciparum (Avalos-Padilla et al., 2021b)
Purified recombinant EhVps20t and EhVps32 from E. histolytica (Avalos-Padilla et al., 2018)
PVA solution (5% w/v) (see Recipe 1)
Lipid mix (1 mg·mL-1) for the injection of recombinant P. falciparum PfBro1 and PfVps32 in vesicles made of POPC:POPS:DSPE-PEG 2000 Biotin:DPPE-Rhodamine (see Recipe 2)
Lipid mix (1 mg·mL-1) for the injection of recombinant E. histolytica EhVps20t and EhVps32 in vesicles made of POPC:POPS:chol:PI(3)P:DSPE-PEG 2000 Biotin:DPPE-Rhodamine (see Recipe 3)
Protein buffer 1×, pH 7.4 (see Recipe 4)
Note: The purification of the P. falciparum proteins (PfBro1 and PfVps32) and E. histolytica proteins (EhVps20t and EhVps32) followed a conventional protocol of affinity purification using the GST-tag present in the recombinant proteins and GSH-Sepharose 4B for its capture. The GST-tag was later removed, and recombinant proteins were purified by size exclusion chromatography as detailed in Avalos-Padilla et al. (2021b) and Avalos-Padilla et al. (2018), respectively.
Consumables
Thin wall borosilicate capillaries with filament (internal radius: 0.78 mm, external radius: 1 mm), to be used in the fabrication of the injection micropipettes (Harvard Apparatus, catalog number: 30-0038)
Cover glass (Menzel Gläser, 26 mm × 56 mm, 0.17 mm thick, custom made)
Hamilton syringe 1 mL (Carl Roth, catalog number: EY44.1)
0.22 µm membrane filter (Milex-GV, catalog number: SLGVR04NL)
Silicone paste (Korasilon-Paste, Carl Roth, catalog number: 0857.1)
2 mm thick homemade Teflon spacer (length/width 40/22 mm), see Figure 2
Eppendorf® microloader 20 µL tips (Eppendorf, catalog number: 5242956003)
Glass vial for the lipid stock solution (DKW Life Science, catalog number: 11768929)
Glass vial for the PVA solution (rollrandglaeser; Carl Roth, catalog number: X661.1)
Pipette tips (capacity 1,000 µL; Merck, catalog number: AXYT1000B)
Weighing pan ROTILABO® (Carl Roth, catalog number: 2149.2)
Measuring vessels for Osmomat 3000 (Gonotec, catalog number: 30.9.0010)
Equipment
Oven (Thermo Fisher Scientific, Heraeus Vacutherm)
Vacuum pump (Vacuubrand, model: MZ 2C NT + 2AK)
Micropipette puller (Sutter Instruments, model: P-97)
Micropipette manipulator system (Sutter Instruments, model: MPC-385)
Microinjector (Eppendorf, model: FemtoJet 5247)
Confocal microscope (Leica TCS SP5 equipped with an oil-immersion objective, 63×, NA 1.4)
Vacuum desiccator ROTILABO® (Carl Roth, article No. 1008.1)
Milli-Q® system (SG water purification system, Integra UV plus, 18.2 MΩ·cm)
Variable volume pipette (0.12 µL, 0.510 µL, 1001,000 µL, Eppendorf®)
Magnetic stirrer (IKA Werke Staufen, type: Bigsquid)
Magnetic stirrer bars (Fisherbrand, catalog number: 11834792)
Osmometer (Gonotec, Osmomat 3000 freezing point osmometer)
Procedure
The complete protocol is summarized in a flow chart shown in Figure 1.
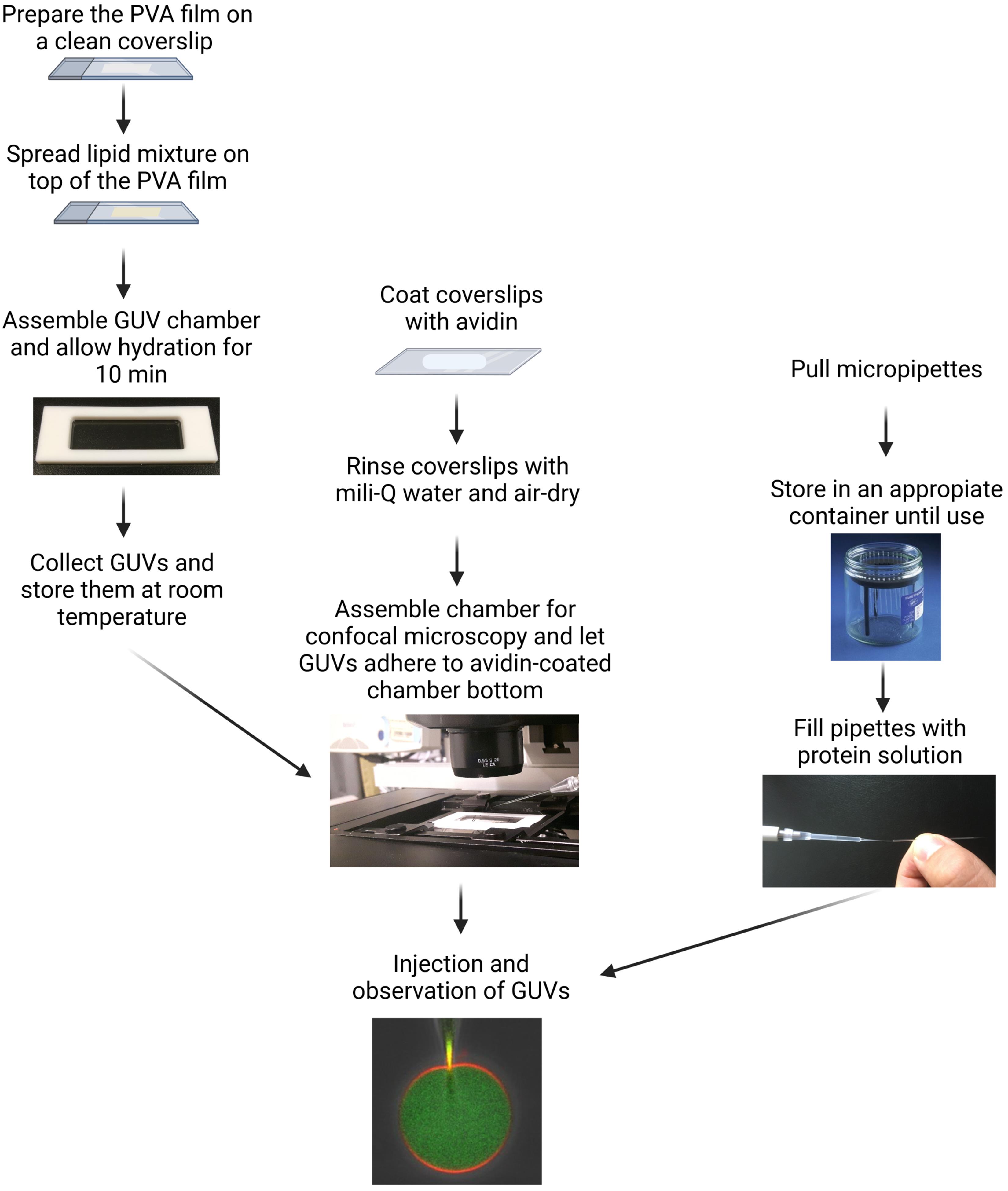
Figure 1. Flow chart of the complete protocol for femtoliter injection of ESCRT-III proteins into adhered GUVs.
Formation of GUVs by PVA gel-assisted swelling
Clean a 26 mm × 56 mm glass slide by rinsing it consecutively with water, ethanol, and water; dry it under a N2 flow. A separate slide is needed for each tested condition.
Place 50 µL of 5% w/v PVA solution (see Recipe 1) on the cleaned glass slide and spread it with the micropipette tip.
Incubate the glass at 50°C in the oven for at least 30 min.
Depending on the protein type to be examined, prepare a mixture of POPC:POPS:DSPE-PEG 2000 Biotin:DPPE-Rhodamine or POPC:POPS:chol:PI(3)P:DSPE-PEG 2000 Biotin:DPPE-Rhodamine (see Recipes 2 and 3) in chloroform at 1 mg·mL-1 final concentration.
Clean thoroughly a Hamilton syringe with chloroform and spread 10 to 15 µL of the lipid mixture on the PVA film dried on the glass (taken from the oven without cooling it down) using the needle of the syringe and until the slide appears dry.
Place the glass slide for 1 h under vacuum to eliminate the excess chloroform.
Glue the Teflon spacer (via silicone grease) to the glass with the dried lipid film (see Figure 2A).
Fill the chamber with 1,800 µL of 1× protein buffer (see Recipe 4) and place a glass coverslip on top of the Teflon spacer (see Figure 2B) to avoid unwanted evaporation.
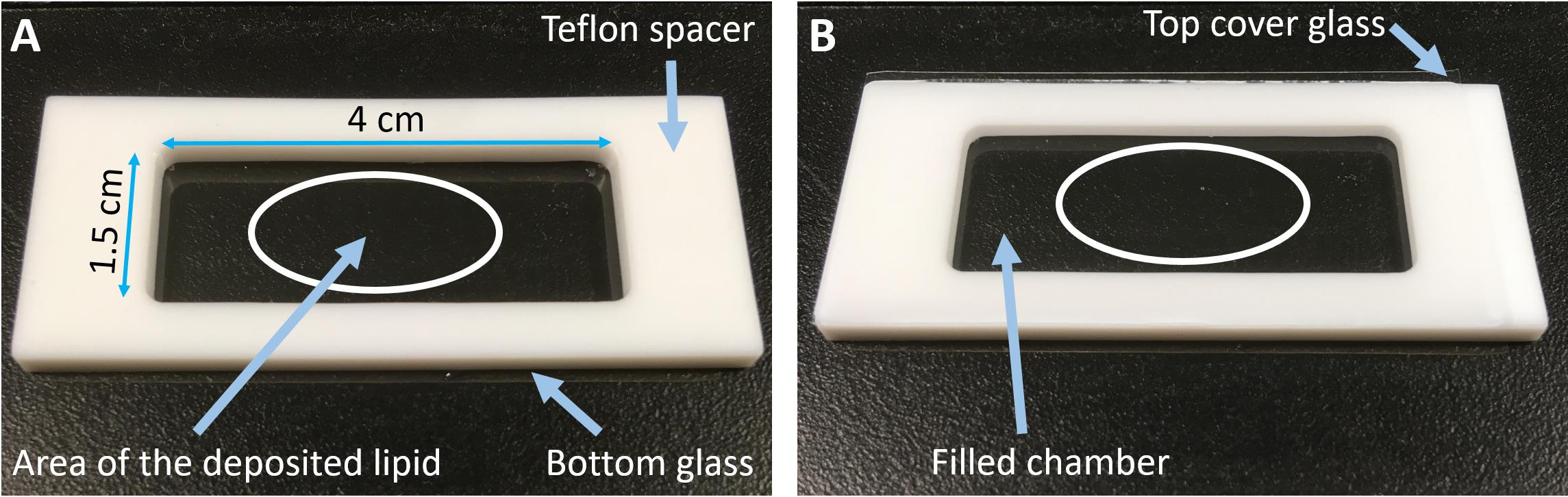

Figure 2. GUV chamber for gel-assisted swelling. The Teflon spacer is between two cover glasses (seen in panel B). The bottom glass is coated with PVA where lipids are deposited. The white oval roughly indicates the area with the deposited lipid mixture.Incubate for 10 min at room temperature, to allow swelling and GUV formation.
After this time, tap gently a few times on the bottom of the growing chamber, remove the upper coverslip by sliding it to the side and collect the GUVs using a 1,000 µL pipette tip without touching the PVA film, to avoid collecting PVA debris. Collected GUVs must be placed in a glass container, protected from light to avoid oxidation, and stored at room temperature (~21°C). The solutions should be used fresh, even though no differences in the behavior were observed for GUV solutions used on the following day after preparation.
Fabrication and loading of the micropipette
Take a borosilicate capillary and carefully apply N2 flow through it, to make sure that the capillary is not clogged.
Place the capillary at the holder of the micropipette puller (Figure 3).
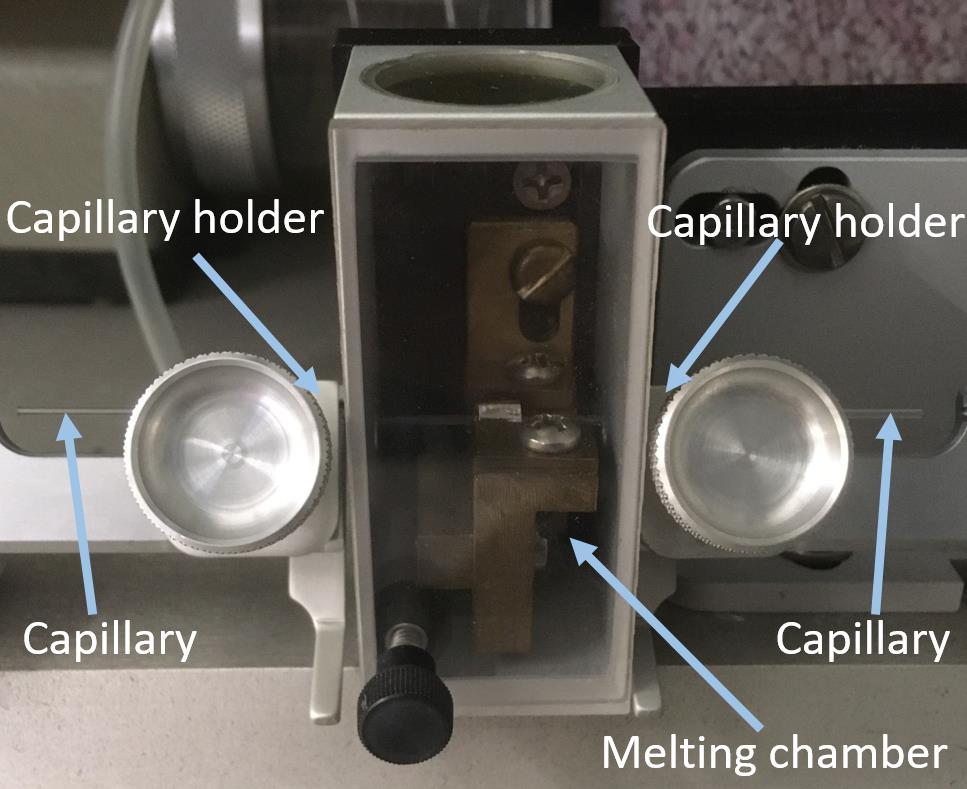

Figure 3. Capillary placed in the micropipette puller.Pull the capillary using the one-line program, to achieve a bee-needle orifice of ~250 nm (HEAT Ramp; PULL 100, VEL 10, TIME 250, PRESSURE 500´) in the micropipette puller.
Filter the injection solution (2.4 µM PfBro1, and 4.8 µM of either PfVps32 or PfVps60 dissolved in 0.8× protein buffer, or 0.6 µM EhVps20t and 2.4 µM EhVps32, and 3 × 10-2 mg·mL-1 PEG-FITC, to monitor the injection; omit the addition of PEG-FITC in case one of the proteins is fluorescently labelled) through the 0.22 µm filter.
Fill the micropipette with 10 µL of the filtered protein solution, using a microloader pipette tip (Figure 4).
Gently tap the filled micropipette and leave it for 30 min in a vertical position to eliminate air bubbles.
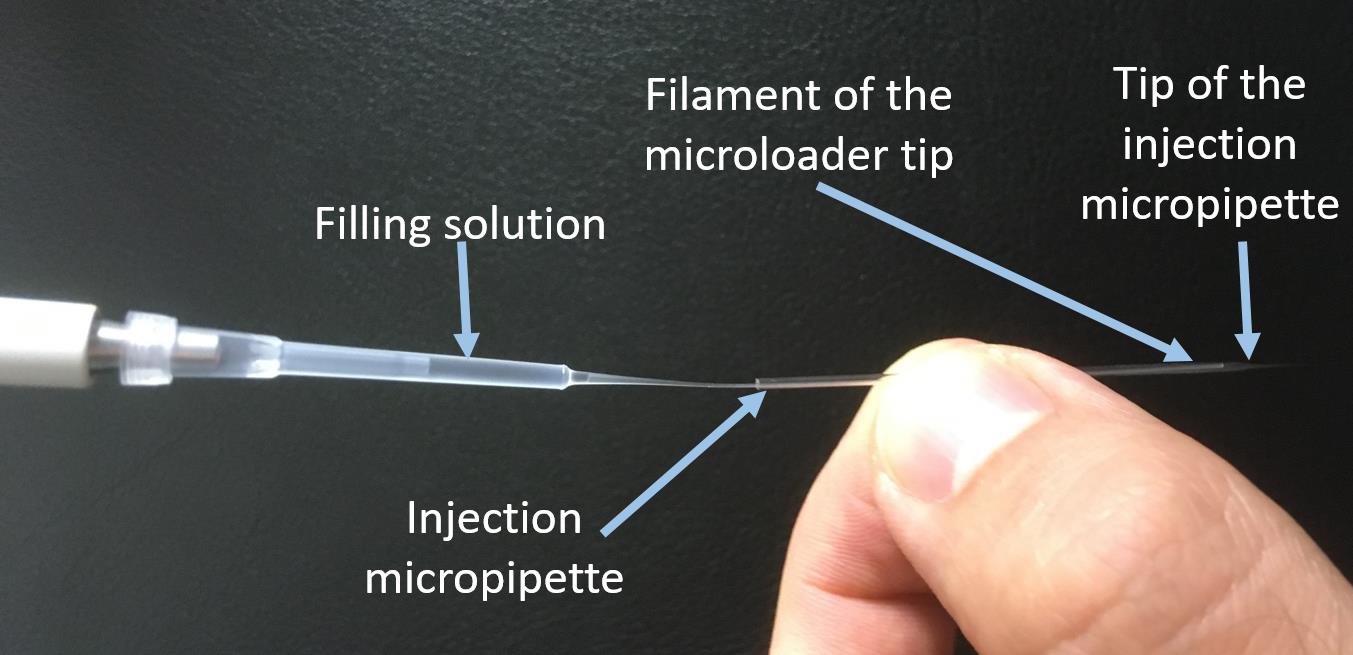

Figure 4. Loading of ESCRT proteins into the injection micropipette. The filament of the microloader tip is inserted as deep as possible into the injection micropipette. Pipette the loading solution into the micropipette and slowly pull out the microloader after loading 10 µL.
Coating of coverslip glasses with avidin and immobilization of GUVs
Clean a 26 mm × 56 mm coverslip by rinsing consecutively with water, ethanol, and water; dry it under N2 flow.
Functionalize the coverslips with 100 µL of a 1:1 (v/v) mixture of BSA (1 mg·mL-1) and biotin-BSA (1 mg·mL-1), dissolved in 1× protein buffer, following the previously reported protocol (Maan et al., 2018).
Incubate for 20 min at room temperature.
Rinse the glass with water and spread 100 µL of 5 × 10-3 mg·mL-1 avidin solution (in 1× protein buffer).
Incubate for 5 min at room temperature.
Rinse the glass with water to remove the unbound avidin.
Glue the Teflon spacer (with silicone paste) to the glass, to form an observation chamber.
Transfer the collected GUVs to the observation chamber with a pipette and let them sediment for at least 10 min. DSPE-PEG 2000 Biotin (in the vesicle membrane) binds to the avidin on the coverslip, resulting in vesicle immobilization.
Note: Increased concentrations of avidin result in higher binding of the GUV to the glass surface, which later hinders budding, as the vesicle excess area is consumed for adhesion (see Figure 5).
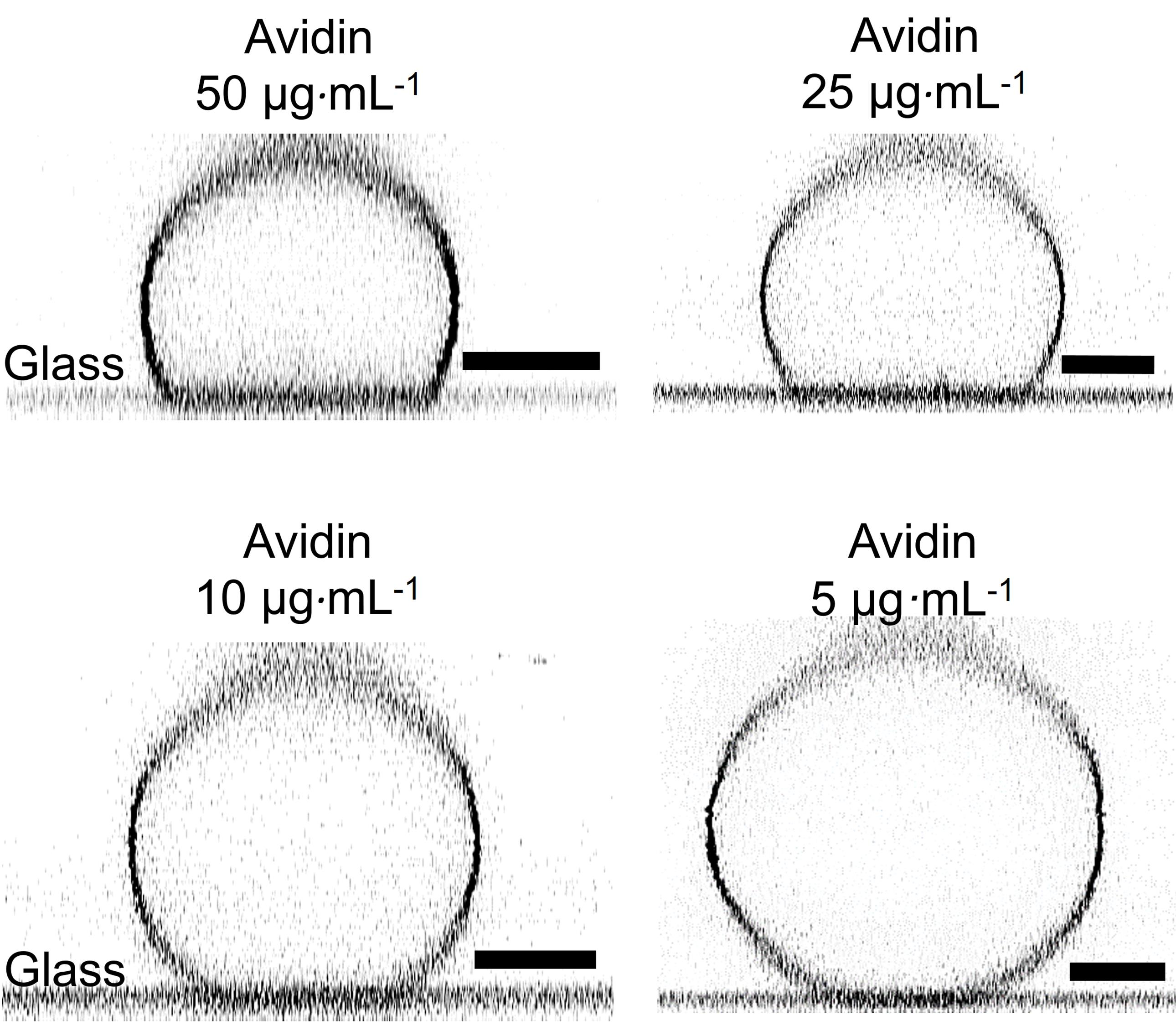
Figure 5. Side view of immobilized vesicles (here, POPC:POPS:chol:PI(3)P:DSPE-PEG 2000 Biotin:DPPE-Rhodamine in molar ratio 60.9:10:25:3:1:0.1). The coverslips were coated with a 1:1 mixture of biotin-BSA (1 mg·mL-1) and BSA (1 mg·mL-1), and then avidin was deposited. The vesicles are visualized via vertical confocal cross sections. The color is inverted. The scale bars correspond to 10 µm.
Injection and observation of GUVs
Place the observation chamber with the immobilized GUVs on the microscope stage.
Connect the filled micropipette to the holder of the microinjector (Figure 6).
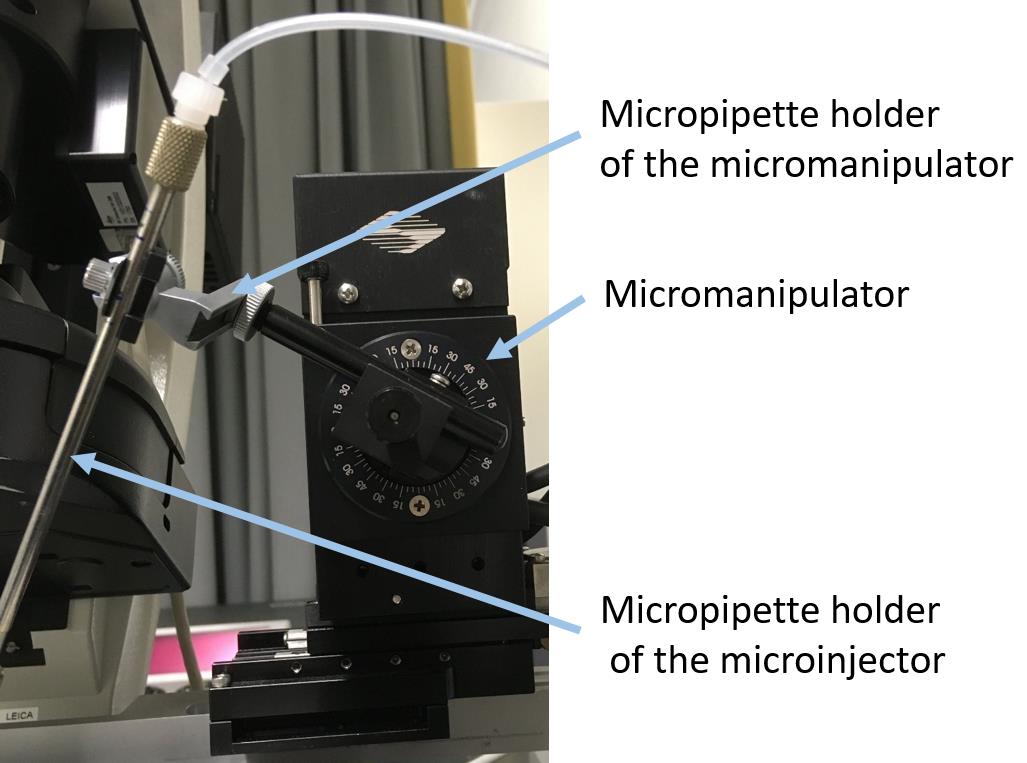

Figure 6. Attachment of the micropipette holder of the microinjector to the micromanipulator.Attach the holder with the pipette to the mechanical arm of the micromanipulator.
Set the angle of injection as large as the microscope setup allows (see Figure 7).
Note: The ideal angle of injection would be 90°, to avoid lateral displacement of the vesicles during puncture attempts. However, this cannot be achieved since the condenser of the microscope occupies the space above the sample.
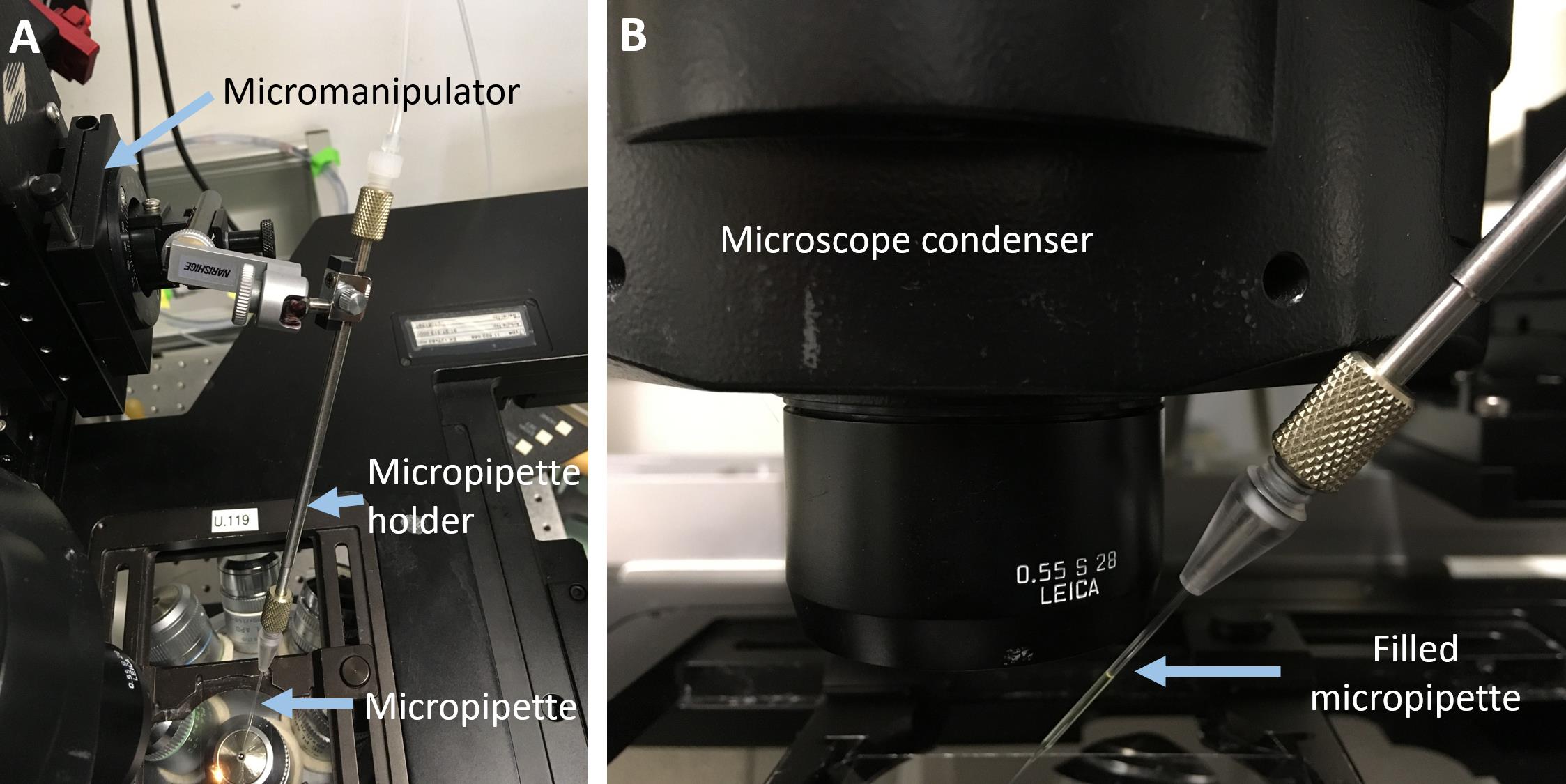

Figure 7. Micropipette setup of the injection procedure. A: Top-view of the setup; for clarity the observation chamber was removed. B: A close-up side-view of the injection angle. Note that the micropipette holder is as close to the condenser of the microscope as possible to ensure high angle of injection.With the micromanipulator, introduce the micropipette into the solution of the observation chamber and focus on the tip of the pipette.
Set the microscope to the desired observation settings. For the measurement presented here, they were the following:
The DPPE-Rhodamine dye (integrated in the membrane of the GUVs) was excited with a 561 nm diode-pumped solid-state laser, and the signal was collected in the 570–650 nm range.
The PEG-FITC dye was excited with a 488 nm argon laser, and the signal was collected in the 495–530 nm range.
To avoid crosstalk between the different fluorescence signals, sequential scanning was performed.
Place the micropipette in a site where no GUVs are observed and purge it (by pressing the “Clean” bottom of the microinjector) to confirm that the micropipette is not clogged; a signal from the fluorescent dye leaving the pipette tip should be detected.
Lower the micropipette close to, but still above the focal plane of the GUVs.
Approach a selected GUV with the tip of the micropipette from above; the selected vesicle should be clean and without defects (to ensure this, examine the selected GUV with a XYZ scan).
Lift the micropipette a few micrometers above the vesicle (the micropipette tip goes out of focus).
Puncture the GUV, by moving the micropipette towards the vesicle in both Z and X directions.
Perform a XYZ scan to make sure that the micropipette penetrates into the GUV.
Start recording a time sequence.
Inject the vesicle (see Figure 8 and Figure 9) using the following parameters of the microinjector: pressure of injection: 150 hPa; time of injection: 5 s; compensation pressure: 1 hPa. These parameters ensure that injected volumes are in the sub-picoliter range (a few hundreds of femtoliters).
Pull the micropipette out from the interior of the GUV (lift the micropipette in Z direction).
Perform a XYZ scan to detect in which focal plane the outward buds formed (Figure 8, Figure 9).
Note: Tense vesicles are easier to puncture than fluctuating ones.
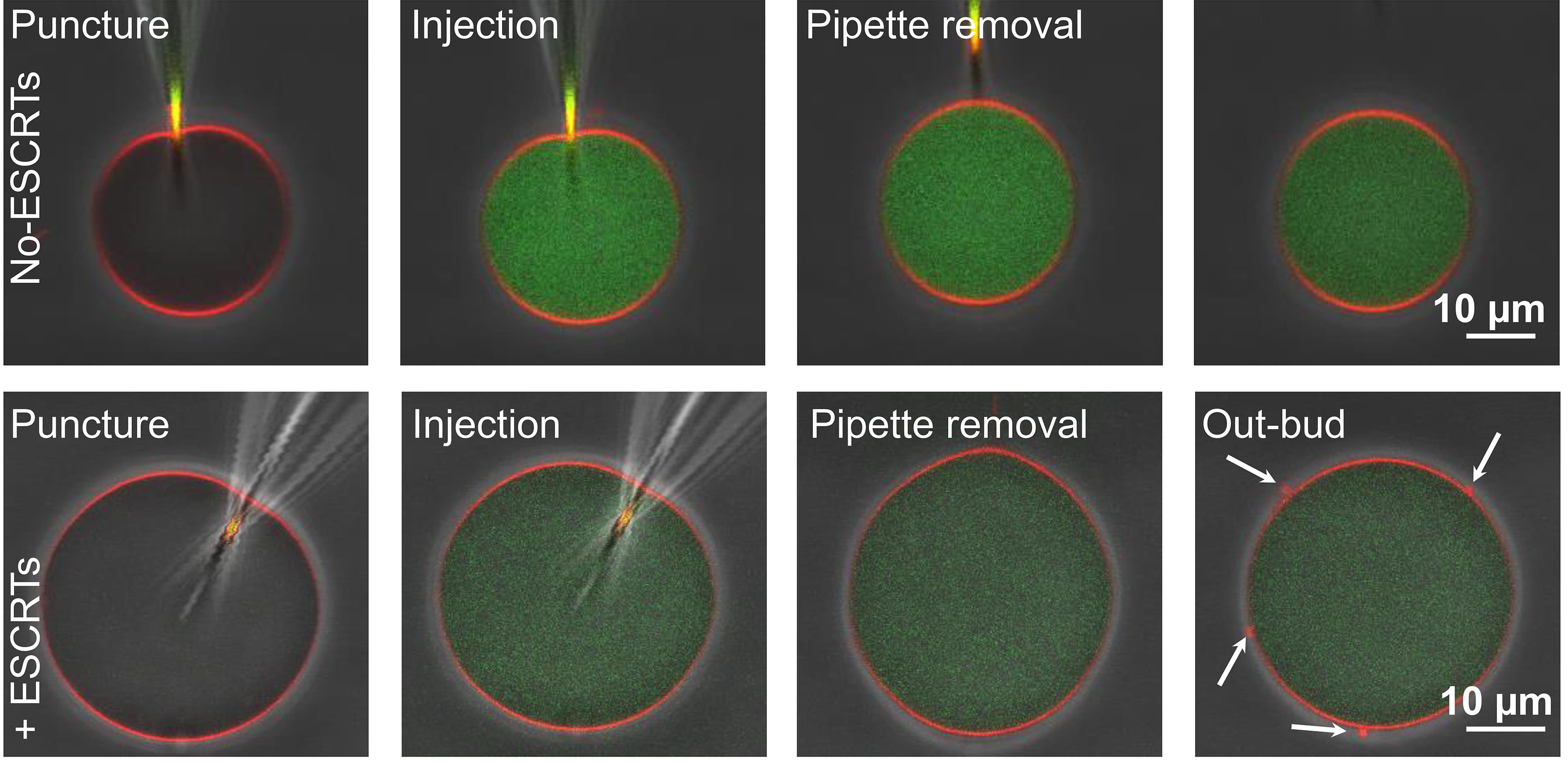
Figure 8. Horizontal (XY) confocal cross sections overlayed with phase-contrast images showing injection of GUVs with PEG-FITC solution (top row) or purified P. falciparum ESCRT-III proteins (bottom row). The membrane is presented in red and PEG-FITC in green. The tip of the injection pipette can be noticed on the membrane in the first two frames in each sequence. The upper row represents the control experiment (ESCRT-free buffer is injected, so outward buds were not observed). In contrast, when ESCRT-III recombinant proteins were injected, outward buds formed; see also Video 1. The arrows on the last frame point to the outward buds. Adapted from Avalos-Padilla et al. (2021b).Video 1. Injection of a GUV with purified P. falciparum ESCRT-III proteins.Real-time recording. The specific information describing the system is indicated in the caption of Figure 8.
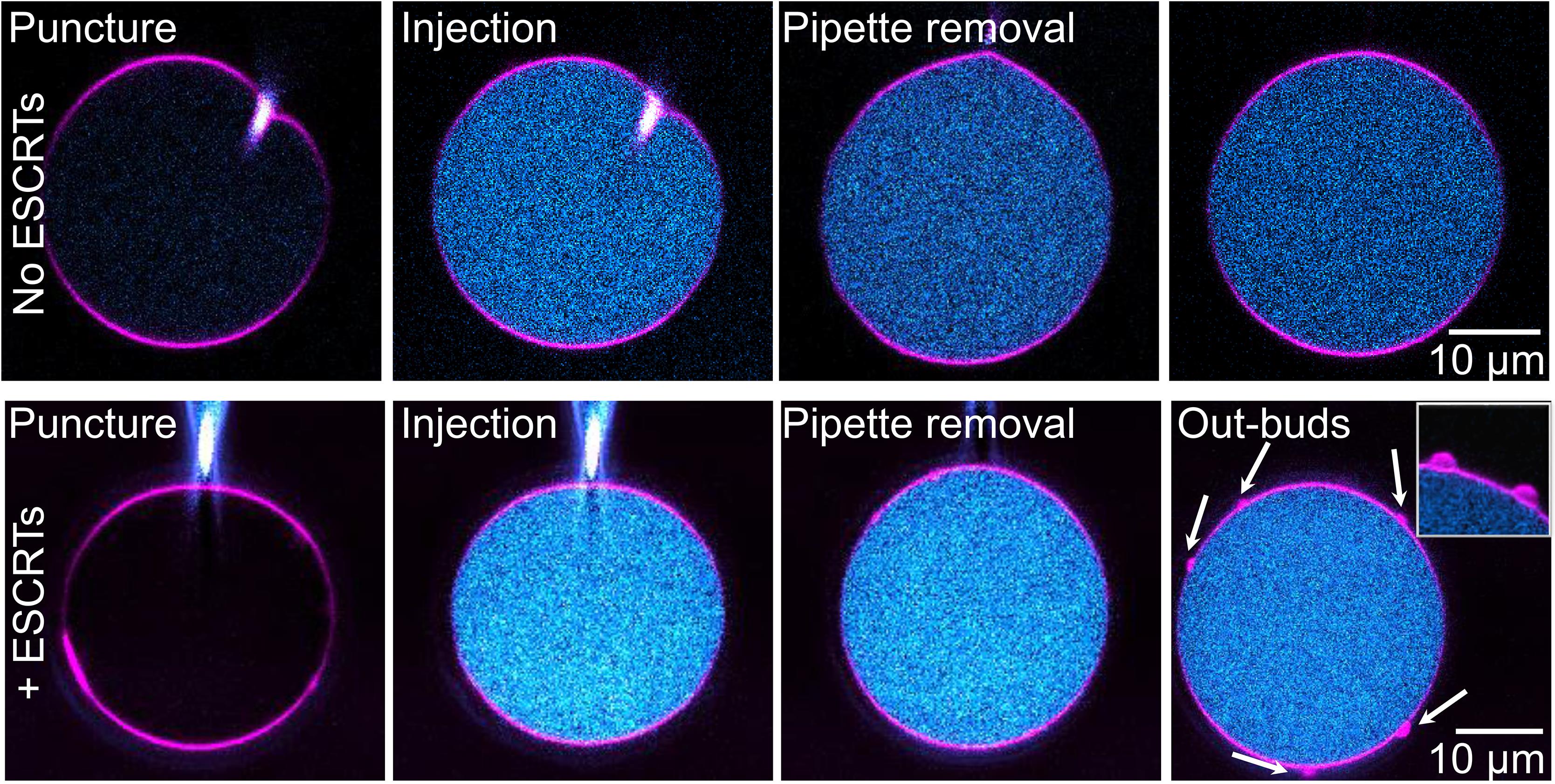
Figure 9. Horizontal (XY) confocal cross sections of injection of GUVs with PEG-FITC solution (top row) or purified E. histolytica ESCRT-III proteins (bottom row). The membrane is shown in magenta and PEG-FITC in cyan. The tip of the injection pipette can be seen on the first two frames in each sequence. The arrows on the last frame point to the outward buds; see also Video 2. The inset in the last snapshot shows a zoomed-in view of outward buds from the same vesicle but at another XY plane.Video 2. Injection of a GUV with purified E. histolytica ESCRT-III proteins.Real-time recording. The specific information describing the system is indicated in the caption of Figure 9.
Recipes
PVA solution (5% w/v)
Weigh 0.5 g PVA and place it in a glass vial.
Add 10 mL of 1× protein buffer (to maintain osmolarity) and place a clean magnetic stirrer in the vial.
Keep the solution in a 90°C water bath under constant stirring (300–400 rpm) until PVA dissolves completely. The solution becomes clear.
Lipid mix (1 mg·mL-1) for the injection of recombinant P. falciparum PfBro1 and PfVps32 in vesicles made of POPC:POPS:DSPE-PEG 2000 Biotin:DPPE-Rhodamine (Final volume: 200 µL):
Lipids (stock solutions) & Reagents Concentration (mg·mL-1) Volume (µL) Final molar fraction % POPC 10 15.78 78.9 POPS 10 4.00 20.0 DSPE-PEG 2000 Biotin 1 2.00 1.0 DPPE-Rhodamine 0.25 1.00 0.1 Chloroform 177.22 Lipid mix (1 mg·mL-1) for the injection of recombinant E. histolytica EhVps20t and EhVps32 in vesicles made of POPC:POPS:chol:PI(3)P:DSPE-PEG 2000 Biotin:DPPE-Rhodamine (Final volume: 200 µL):
Lipids (stock solutions) & Reagents Concentration (mg·mL-1) Volume (µL) Final molar fraction % POPC 10 12.18 60.9 POPS 10 2.00 10.0 chol 10 5.00 25.0 PI(3)P (dissolved in chloroform/methanol 60:40) 0.25 24.00 3.0 DSPE-PEG 2000 Biotin 1 2.00 1.0 DPPE-Rhodamine 0.25 1.00 0.1 Chloroform 153.82 Protein buffer 1×, pH 7.4:
Compound Concentration (mM) Osmolarity (mOsm) Trizma® hydrochloride 25 325 NaCl 150 Adjust pH with 5 M HCl
Acknowledgments
This work is part of the MaxSynBio consortium, which was jointly funded by the Federal Ministry of Education and Research of Germany and the Max Planck Society. The work was funded by the Ministerio de Ciencia, Innovación y Universidades, Spain (which included FEDER funds), grant number RTI2018-094579-B-I00. ISGlobal and IBEC are members of the CERCA Programme, Generalitat de Catalunya. Y. A. P. acknowledges the financial support provided by the European Commission under Horizon 2020’s Marie Skłodowska-Curie Actions COFUND scheme (712754) and by the Severo Ochoa programme of the Spanish Ministry of Science and Competitiveness [SEV-2014-0425 (2015-2019). This research is part of ISGlobal's Program on the Molecular Mechanisms of Malaria which is partially supported by the Fundación Ramón Areces. Illustrations created with BioRender.com. This protocol was originally used and published in Avalos-Padilla et al. (2021a).
Competing interests
The authors declare no competing interests.
References
- Alqabandi, M., de Franceschi, N., Maity, S., Miguet, N., Bally, M., Roos, W. H., Weissenhorn, W., Bassereau, P. and Mangenot, S. (2021). The ESCRT-III isoforms CHMP2A and CHMP2B display different effects on membranes upon polymerization. BMC Biol 19(1): 66.
- Angelova, M. I. and Dimitrov, D. S. (1986). Liposome Electroformation. Faraday Discuss 81: 303-311.
- Avalos-Padilla, Y., Georgiev, V. N. and Dimova, R. (2021a). ESCRT-III induces phase separation in model membranes prior to budding and causes invagination of the liquid-ordered phase. Biochim Biophys Acta Biomembr 1863(10): 183689.
- Avalos-Padilla, Y., Georgiev, V. N., Lantero, E., Pujals, S., Verhoef, R., L, N. B.-C., Albertazzi, L., Dimova, R. and Fernandez-Busquets, X. (2021b). The ESCRT-III machinery participates in the production of extracellular vesicles and protein export during Plasmodium falciparum infection. PLoS Pathog 17(4): e1009455.
- Avalos-Padilla, Y., Knorr, R. L., Javier-Reyna, R., García-Rivera, G., Lipowsky, R., Dimova, R. and Orozco, E. (2018). The conserved ESCRT-III machinery participates in the phagocytosis of Entamoeba histolytica. Front in Cell Infect 8: 53.
- Booth, A., Marklew, C. J., Ciani, B. and Beales, P. A. (2019). In vitro membrane remodeling by ESCRT is regulated by negative feedback from membrane tension. Iscience 15: 173-184.
- Campos, F. M. F., Franklin, B. S., Teixeira-Carvalho, A., Filho, A. L. S., de Paula, S. C. O., Fontes, C. J., Brito, C. F. and Carvalho, L. H. (2010). Augmented plasma microparticles during acute Plasmodium vivax infection. Malar J 16: 327.
- Clos-Garcia, M., Loizaga-Iriarte, A., Zuniga-Garcia, P., Sanchez-Mosquera, P., Rosa Cortazar, A., Gonzalez, E., Torrano, V., Alonso, C., Perez-Cormenzana, M., Ugalde-Olano, A., et al. (2018). Metabolic alterations in urine extracellular vesicles are associated to prostate cancer pathogenesis and progression. J Extracell Vesicles 7(1): 1470442.
- Combes, V., Taylor, T. E., Juhan-Vague, I., Mege, J. L., Mwenechanya, J., Tembo, M., Grau, G. E. and Molyneux, M. E. (2004). Circulating endothelial microparticles in Malawian children with severe falciparum malaria complicated with coma. JAMA 291(21): 2542-2544.
- Dao, T. P. T., Fauquignon, M., Fernandes, F., Ibarboure, E., Vax, A., Prieto, M. and Le Meins, J. F. (2017). Membrane properties of giant polymer and lipid vesicles obtained by electroformation and PVA gel-assisted hydration methods.Colloids Surf A 553: 347-353.
- Dimova, R. (2019). Giant Vesicles and Their Use in Assays for Assessing Membrane Phase State, Curvature, Mechanics, and Electrical Properties. Annu Rev Biophys 48(1): 93-119.
- Dimova, R. and Marques, C. (2019). The Giant Vesicle Book. Taylor & Francis Group, LLC. Boca Raton. ISBN: 9781498752176.
- Dores, M. R., Chen, B., Lin, H., Soh, U. J., Paing, M. M., Montagne, W. A., Meerloo, T. and Trejo, J. (2012). ALIX binds a YPX(3)L motif of the GPCR PAR1 and mediates ubiquitin-independent ESCRT-III/MVB sorting. J Cell Biol 197(3): 407-419.
- Doyotte, A., Russell, M. R., Hopkins, C. R. and Woodman, P. G. (2005). Depletion of TSG101 forms a mammalian "Class E" compartment: a multicisternal early endosome with multiple sorting defects. J Cell Sci 118(Pt 14): 3003-3017.
- Glover, S. C., Nouri, M. Z., Tuna, K. M., Mendoza Alvarez, L. B., Ryan, L. K., Shirley, J. F., Tang, Y., Denslow, N. D. and Alli, A. A. (2019). Lipidomic analysis of urinary exosomes from hereditary alpha-tryptasemia patients and healthy volunteers. FASEB Bioadv 1(10): 624-638.
- Herrmann, I. K., Wood, M. J. A. and Fuhrmann, G. (2021). Extracellular vesicles as a next-generation drug delivery platform. Nat Nanotechnol 16(7): 748-759.
- Hurtig, J. and Orwar, O. (2008). Injection and transport of bacteria in nanotube-vesicle networks. Soft Matter 4(7): 1515-1520.
- Im, Y. J., Wollert, T., Boura, E. and Hurley, J. H. (2009). Structure and function of the ESCRT-II-III interface in multivesicular body biogenesis. Dev Cell 17(2): 234-243.
- Keklikoglou, I., Cianciaruso, C., Guc, E., Squadrito, M. L., Spring, L. M., Tazzyman, S., Lambein, L., Poissonnier, A., Ferraro, G. B., Baer, C., et al. (2019). Chemotherapy elicits pro-metastatic extracellular vesicles in breast cancer models. Nat Cell Biol 21(2): 190-202.
- Lefrançois, P., Goudeau, B. and Arbault, S. (2018). Electroformation of phospholipid giant unilamellar vesicles in physiological phosphate buffer. Integr Biol (Camb) 10(7): 429-434.
- Li, Q., Wang, X., Ma, S., Zhang, Y. and Han, X. (2016). Electroformation of giant unilamellar vesicles in saline solution. Colloids Surf B Biointerfaces 147: 368-375.
- Maan, R., Loiseau, E. and Bausch, A. R. (2018). Adhesion of active cytoskeletal vesicles. Biophys J 115(12): 2395-2402.
- Martin-Jaular, L., Nakayasu, E. S., Ferrer, M., Almeida, I. C. and Del Portillo, H. A. (2011). Exosomes from Plasmodium yoelii-infected reticulocytes protect mice from lethal infections. PLoS One 6(10): e26588.
- Mfonkeu, J. B. P., Gouado, I., Kuate, H. F., Zambou, O., Zollo, P. H. A., Grau, G. E. R. and Combes, V. (2010). Elevated Cell-Specific Microparticles Are a Biological Marker for Cerebral Dysfunctions in Human Severe Malaria. Plos One 5(10): e13415.
- Montes, L. R., Alonso, A., Goni, F. M. and Bagatolli, L. A. (2007). Giant unilamellar vesicles electroformed from native membranes and organic lipid mixtures under physiological conditions. Biophys J 93(10): 3548-3554.
- Moyano, A. L., Li, G., Boullerne, A. I., Feinstein, D. L., Hartman, E., Skias, D., Balavanov, R., van Breemen, R. B., Bongarzone, E. R., Mansson, J. E., et al. (2016). Sulfatides in extracellular vesicles isolated from plasma of multiple sclerosis patients. J Neurosci Res 94(12): 1579-1587.
- Nabhan, J. F., Hu, R., Oh, R. S., Cohen, S. N. and Lu, Q. (2012). Formation and release of arrestin domain-containing protein 1-mediated microvesicles (ARMMs) at plasma membrane by recruitment of TSG101 protein. Proc Natl Acad Sci U S A 109(11): 4146-4151.
- Nantakomol, D., Dondorp, A. M., Krudsood, S., Udomsangpetch, R., Pattanapanyasat, K., Combes, V., Grau, G. E., White, N. J., Viriyavejakul, P., Day, N. P., et al. (2011). Circulating red cell-derived microparticles in human malaria. J Infect Dis 203(5): 700-706.
- Nickerson, D. P., West, M. and Odorizzi, G. (2006). Did2 coordinates Vps4-mediated dissociation of ESCRT-III from endosomes. J Cell Biol 175(5): 715-720.
- Pott, T., Bouvrais, H. and Meleard, P. (2008). Giant unilamellar vesicle formation under physiologically relevant conditions. Chem Phys Lipids 154(2): 115-119.
- Raposo, G. and Stahl, P. D. (2019). Extracellular vesicles: a new communication paradigm? Nat Rev Mol Cell Biol 20(9): 509-510.
- Schofield, L. and Grau, G. E. (2005). Immunological processes in malaria pathogenesis.Nat Rev Immunol 5(9): 722-735.
- Schöneberg, J., Pavlin, M. R., Yan, S., Righini, M., Lee, I. H., Carlson, L. A., Bahrami, A. H., Goldman, D. H., Ren, X. F., Hummer, G., et al. (2018). ATP-dependent force generation and membrane scission by ESCRT-III and Vps4.Science 362(6421): 1423-1428.
- Tang, S. G., Buchkovich, N. J., Henne, W. M., Banjade, S., Kim, Y. J. and Emr, S. D. (2016). ESCRT-III activation by parallel action of ESCRT-I/II and ESCRT-0/Bro1 during MVB biogenesis. Elife 13(5): e15507.
- Tao, L., Zhou, J., Yuan, C., Zhang, L., Li, D., Si, D., Xiu, D. and Zhong, L. (2019). Metabolomics identifies serum and exosomes metabolite markers of pancreatic cancer. Metabolomics 15(6): 86.
- van Niel, G., D'Angelo, G. and Raposo, G. (2018). Shedding light on the cell biology of extracellular vesicles. Nat Rev Mol Cell Biol 19(4): 213-228.
- Vietri, M., Radulovic, M. and Stenmark, H. (2020). The many functions of ESCRTs. Nat Rev Mol Cell Biol 21(1): 25-42.
- Virtanen, J. A., Cheng, K. H. and Somerharju, P. (1998). Phospholipid composition of the mammalian red cell membrane can be rationalized by a superlattice model. Proc Natl Acad Sci U S A 95(9): 4964-4969.
- Wang, Q. and Lu, Q. (2017). Plasma membrane-derived extracellular microvesicles mediate non-canonical intercellular NOTCH signaling. Nat Commun 8(1): 709.
- Weinberger, A., Tsai, F. C., Koenderink, G. H., Schmidt, T. F., Itri, R., Meier, W., Schmatko, T., Schroder, A. and Marques, C. (2013). Gel-Assisted Formation of Giant Unilamellar Vesicles. Biophys J 105(1): 154-164.
- Wick, R., Angelova, M. I., Walde, P. and Luisi, P. L. (1996). Microinjection into giant vesicles and light microscopy investigation of enzyme-mediated vesicle transformations. Chem Biol 3(2): 105-111.
- Wolfers, J., Lozier, A., Raposo, G., Regnault, A., Thery, C., Masurier, C., Flament, C., Pouzieux, S., Faure, F., Tursz, T., et al. (2001). Tumor-derived exosomes are a source of shared tumor rejection antigens for CTL cross-priming. Nat Med 7(3): 297-303.
- Xu, R., Rai, A., Chen, M., Suwakulsiri, W., Greening, D. W. and Simpson, R. J. (2018). Extracellular vesicles in cancer - implications for future improvements in cancer care.Nat Rev Clin Oncol 15(10): 617-638.
Article Information
Copyright
© 2022 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Georgiev, V. N., Avalos-Padilla, Y., Fernàndez-Busquets, X. and Dimova, R. (2022). Femtoliter Injection of ESCRT-III Proteins into Adhered Giant Unilamellar Vesicles. Bio-protocol 12(4): e4328. DOI: 10.21769/BioProtoc.4328.
Category
Biophysics > Microscopy
Biological Engineering > Synthetic biology
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link











