- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Isolation of Mitochondria from Ustilago maydis Protoplasts
Published: Vol 12, Iss 1, Jan 5, 2022 DOI: 10.21769/BioProtoc.4277 Views: 3608
Reviewed by: Ashish RanjanAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Measurement of Energy-dependent Rhodamine 6G Efflux in Yeast Species
Yvetta Gbelska [...] Alexandra Konecna
Aug 5, 2017 8988 Views
An Assay to Determine NAD(P)H: Quinone Oxidoreductase Activity in Cell Extracts from Candida glabrata
Anamika Battu [...] Rupinder Kaur
Nov 5, 2021 3090 Views
Abstract
Ustilago maydis, a basidiomycete that infects Zea mays, is one of the top ten fungal models for studying DNA repair, signal transduction pathways, and dimorphic transitions, among other processes. From a metabolic point of view, U. maydis lacks fermentative capacity, pointing to mitochondria as a key player in central metabolism. Oxidative phosphorylation, synthesis of heme groups, Krebs cycle, β-oxidation of fatty acids, and synthesis of amino acids are some of the processes that take place in mitochondria. Given the importance of this organelle in eukaryotic cells in general, and in fungal cells in particular, we present a protocol for the isolation of U. maydis mitochondria based on the enzymatic disruption of U. maydis cell wall and differential centrifugation. The method can easily be extrapolated to other fungal species, by using appropriate lytic enzymes.
Background
As in other eukaryotes, mitochondria in fungal cells are essential for the production of many important molecules. For example, in the mitochondrial matrix occurs the synthesis of some amino acids and of the heme group that is inserted into specific subunits of the respiratory complexes. Other processes also occur in the mitochondrial matrix, like the β-oxidation of fatty acids and the tricarboxylic acid cycle. NADH and FADH2 are key products of the last two pathways, and these molecules transfer their electrons to the respiratory chain for the synthesis of ATP. Electron transfer and ATP synthesis are connected by a proton electrochemical gradient that is generated by the proton pumping activity of complexes I, III, and IV in the respiratory chain, and consumed by the ATP synthase. Membrane potential and a difference of pH across the inner mitochondrial membrane are components of the proton-motive force. For mitochondria to carry out their functions, transporters located in the outer and inner membranes catalyze the metabolic exchange between cytosol and mitochondria (Sousa et al., 2018; Reyes-Galindo et al., 2019).
U. maydis is a phytopathogen that infects Zea mays. During its life cycle, U. maydis transits through three major morphological stages: the non-pathogenic haploid basidiospore (yeast form), the infective dikaryotic filament, and the environmental resistant teliospore (Brefort et al., 2009). The microorganism is fully respiratory, pointing to mitochondria as a key player in its intermediate metabolism. Therefore, mitochondria isolation is a starting point in studies of respiratory complexes and supercomplexes, the control of ROS production by certain mitochondrial inner membrane proteins, the activities of Krebs cycle enzymes, among others (Juárez et al., 2004). The goal of this protocol is to report a method for the isolation of functional U. maydis mitochondria from protoplasts generated by cell wall lytic enzymes from Trichoderma harzianum. Membrane potential and oxygen consumption were analyzed, to measure the quality of mitochondrial preparations.
Materials and Reagents
Sterile toothpick
Petri dishes (Sigma-Aldrich, catalog number: P5606-400EA)
Glass beaker (Sigma-Aldrich, catalog number: BR90648)
NalgeneTM Polycarbonate Fernbach Culture Flask (Thermo ScientificTM, catalog number: 4105-2800)
Spectrophotometer cuvettes (Merck, catalog number: C5291)
Nalgene® 500 mL centrifuge bottles, wide-mouth with sealing caps style 3141 (Sigma-Aldrich, catalog number: Z353744-4EA)
Polycarbonate tubes, 50 mL, 29 × 102 mm (Thermo Fisher Scientific, catalog number: 03268)
Quartz absorption cuvettes of 3.5 mL (Merck, catalog number: Z803669)
Hamilton syringe (Hamilton, Gastight syringe, catalog number: 1702N)
Round point tip soft paintbrush number 8 to 12 (DUGATO-Amazon)
Distilled water
Sucrose (Sigma-Aldrich, catalog number: S5016)
Glucose (Sigma-Aldrich, catalog number: G8270)
Sorbitol (Sigma-Aldrich, catalog number: S1876)
Select Yeast Extract (Sigma-Aldrich, catalog number: y1625)
Bacteriological Peptone (Sigma-Aldrich, catalog number: PO556)
Agar (Sigma-Aldrich, catalog number: A1296)
Lysing enzymes from Trichoderma harzianum (Sigma-Aldrich, catalog number: L1412)
Dimethyl sulfoxide (DMSO) (Sigma-Aldrich, catalog number: D2650)
(NH4)2SO4 (Sigma-Aldrich, catalog number: A4915)
KH2PO4 (Sigma-Aldrich, catalog number: PO662)
HCl (Sigma-Aldrich, catalog number: H1758)
NaOH (Sigma-Aldrich, catalog number: 795429)
Glycerol (Sigma-Aldrich, catalog number: G5516)
Potassium cyanide (KCN) (Sigma-Aldrich, catalog number: 60178)
Trizma® base (Sigma-Aldrich, catalog number: T1503)
Ethylenediaminetetraacetic acid (EDTA) disodium salt dihydrate (Sigma-Aldrich, catalog number: E9884)
Bovine Serum Albumin (BSA) (Sigma-Aldrich, catalog number: A7030)
Phenylmethylsulfonyl fluoride (PMSF) (Sigma-Aldrich, catalog number: PMSF-RO)
Complete Protease Inhibitor Cocktail Tablets (Roche, catalog number: 04906837001)
Carbonyl cyanide 4-(trifluoromethoxyphenyl) phenylhydrazone (FCCP) (Sigma-Aldrich, catalog number: C2920)
HEPES (4-(2-hydroxyethyl)-1 piperazineethanesulfonic acid) (Sigma-Aldrich, catalog number: H3375)
Safranine (Sigma-Aldrich, catalog number: S2255)
NADH (Sigma-Aldrich, catalog number: N8129)
Sodium succinate dibasic hexahydrate (Sigma-Aldrich, catalog number: S2378)
N,N,N’,N’-Tetramethyl-p-phenylenediamine (TMPD) (Sigma- Aldrich, catalog number: T7394)
Sodium L-ascorbate (Sigma- Aldrich, catalog number: A7631)
Solution A (see Recipes)
Solution B (see Recipes)
Solution C (see Recipes)
PMSF (see Recipes)
Solution D (see Recipes)
Solution E (see Recipes)
YPD-culture media (see Recipes)
Solution F (see Recipes)
FCCP solution (see Recipes)
Safranine solution (see Recipes)
Succinate solution (see Recipes)
Equipment
Teflon stirring rod (FisherbrandTM, catalog number: 14-513-85)
Glass/Teflon Potter Elvehjem homogenizer (Thomas®, catalog number: C917)
UV-Visible spectrophotometer (Thermo ScientificTM, GENESYS 20)
DW-2a-TM UV/Visible spectrophotometer (AmincoTM, Modernized by OLIS, Inc.)
Water bath (PolyScience, constant temperature, basic controller)
Electric drill (Black+Decker)
Thermo Scientific Rotor A27-8 × 50
Thermo Scientific Rotor FiberliteTM F12-6 × 500 LEX
Thermo Scientific Sorvall LYNX 4000 centrifuge (Thermo Fisher Scientific)
High-Speed Refrigerated Microcentrifuge (SCILOGEX, model: D3024R-SCILOGEX)
Magnetic Stirrer (Thermolyne Cimarec 1)
Vortex Mixer (Thermo Scientific, model: M16710-33Q)
Variable autotransformer (Staco Energy Products Co. Model 3PN10108, input 120V 50/60 Hz, Output 0-140V, AMP 1.4 KVA)
Incubator shaker for yeast growth in liquid culture (SEVMEXICO, PRENDO-INO 650V-7)
Biological oxygen monitor, equipped with a Clark type polarographic electrode (COLE-PARMER®, YSI5300A-1)
Ultra-Low Temperature Freezer (RevcoTM, model: ULT1340-3-A36)
Beckman Phi 32 pH Meter
Procedure
Protoplast preparation
Streak the U. maydis strain from a 25% glycerol stock (maintained at -70°C) on solid YPD agar (Yeast extract, Peptone, Dextrose; see Recipe 7), incubate for 1-2 days at 28°C, and then store the plate at 4°C.
Select a colony with a sterile toothpick.
Culture cells at 28°C and 180 rpm in 100 mL of YPD medium for 18-24 h.
Inoculate cells in 1 L of YPD liquid medium (at an initial optical density at 600 nm = 0.04 = 9 × 105 cells/mL) and incubate for 24 h at 28°C/180 rpm. The incubation time and culture media should be adjusted to your specific needs.
Collect the cells using a pre-weighed 500 mL centrifuge bottle. Centrifuge for 5 min at 3,800 × g and 4°C in the FiberliteTM F12-6 × 500 LEX. Remove supernatant, suspend the cells in distilled water, and repeat the centrifugation step. Decant the supernatant.
Measure the wet weight of the pellet, and add 12.5 mL of solution A per gram of wet weight. Use a glass or Teflon rod to suspend the cells. At this stage, cells can be vortexed to disperse cell aggregates. Transfer the cell suspension to a glass beaker. Add 0.016 g of the T. harzianum lysing enzymes per gram of wet weight. Gently mix with a Teflon or glass rod, and incubate for 30-60 min at 30°C in a water bath.
To check the formation of protoplasts, shake the cell suspension every 10 min, withdraw 20 µL of the suspension, and mix with 980 µL of distilled water. Measure the optical density at 600 nm (Figure 1A). Due to the osmotic shock, cells without a cell wall burst, and their content are released to the medium. This results in a decrease in the optical density of the suspension. When the optical density reaches its minimal value, the formation of protoplasts is complete.
Note: From this step, all procedures should be carried out at 4°C.
Centrifuge the suspension at 3,800 × g at 4°C for 5 min to recover the protoplasts. Decant the supernatant.
Add 12.5 mL of cold solution B per gram of initial wet weight, to wash the lysing enzymes. With a Teflon or glass rod, carefully suspend the protoplasts. Do not use the vortex to disperse the protoplasts. Then, centrifuge the suspension for 10 min at 3,800 × g and 4°C. The new pellet should be compact.
Mitochondria isolation
Transfer the pellet of protoplasts to a Teflon pestle homogenizer and add cold solution C (40 mL/25-30 g of the initial wet weight of cells) containing the complete cocktail of protease inhibitors plus PMSF (1 mM final concentration).
Attach the Teflon pestle to the electric drill. Connect the drill to the variable autotransformer (rheostat), select an output intensity of 40, and homogenize for 20 cycles. Avoid the formation of foam and bubbles.
After homogenization, bring the volume to 130 mL with cold solution C (containing protease inhibitors), transfer the suspension to 50 mL polycarbonate tubes, and eliminate cell debris and nuclei by centrifuging at 3,800 × g and 4°C for 10 min.
Recover the supernatant from the polycarbonate tubes and centrifuge at 17,000 × g and 4°C for 10 min. Carefully decant the supernatant. Then, with the buffer remaining in the tube and with the help of a soft paintbrush (number 8 or 12, according to the biomass), suspend the mitochondrial pellet. Keep mitochondria at 4°C for same-day experiments, or at -70°C to be used later.
Determine the protein concentration of your sample by the Lowry method (Lowry et al., 1951). Prepare a stock solution of 1 mg/mL BSA and construct a standard curve in the range of 5 to 100 µg protein. Dilute the sample tenfold using 0.4% sodium deoxycholate solution, and use 5-10 µL of the diluted sample for protein determination.
Oxygen consumption
To measure the respiratory activity of isolated mitochondria in a thermostated chamber with magnetic stirring, use a Clark-type oxygen electrode coupled to a YSI5300A biologic oxygen monitor.
Bring the thermostated chamber, the solutions D and E, and the distilled water to 30°C.
Pour 1.135 mL of distilled water, 65 µL of solution D (final concentration 50 mM), and 100 µL of solution E (final concentration 20 mM) in the chamber with magnetic stirring. Adjust the signal to 100%, which corresponds to the dissolved oxygen.
Add 5-10 µL of isolated mitochondria; wait until the signal is stabilized and then stimulate oxygen consumption by adding 10 mM succinate (13 µL from 1 M stock solution), or 0.15 mM NADH (9.8 µL from 20 mM stock solution).
To inhibit respiratory complex IV, add 0.5 mM KCN (2.6 µL from 0.25 M stock solution), or 2 µM n-octyl gallate (1.3 µL from 2 mM stock solution), to inhibit the alternative oxidase
Wash the chamber with distilled water before adding new media.
Calculate the specific activity by dividing the oxygen consumption slope by the amount of protein (mg) or the dry weight (g).
Mitochondrial membrane potential assay
Turn on the Olis Modernized AmincoTM DW-2 spectrophotometer and allow the lamp to warm up for 30 min. Select the dual-beam option, and adjust the wavelengths of monochromator 1 (MONO1) to 511 nm, and of MONO2 to 533 nm. In the dual-beam mode, the sample is illuminated with two different wavelengths, one corresponding to the reference wavelength and the other to the measuring wavelength. The output corresponds to the electronic subtraction of the absorbance at 511 nm minus the absorbance at 533 nm.
Transfer 2.7 mL of solution F into a 3.7 mL glass cuvette containing a magnetic stirrer. Start the magnetic stirring. Add 2 µM safranine (5.4 µL from 1 mM stock solution) and then the mitochondrial sample (0.5-1.0 mg protein). Place the cuvette in the spectrophotometer and close the cover.
Adjust the baseline and add, through a small hole made in the chamber cover, the substrate of your interest (for example, 10 mM of succinate, or 0.15 mM NADH) with a Hamilton syringe.
To collapse the membrane potential generated by succinate or any other respiratory substrate, add 2-10 μM FCCP.
Data analysis
Protoplast formation
The formation of protoplasts is followed by the decrease in optical density at 600 nm (OD600nm). As shown in Figure 1A, after 60 min of incubation in the presence of the lytic enzymes, the transformation of cells into protoplasts is complete. In addition, Figure 1B shows that the formation of protoplasts is associated with cell flocculation. Next, respiratory activity and generation of membrane potential were assayed to evaluate the quality of mitochondria.
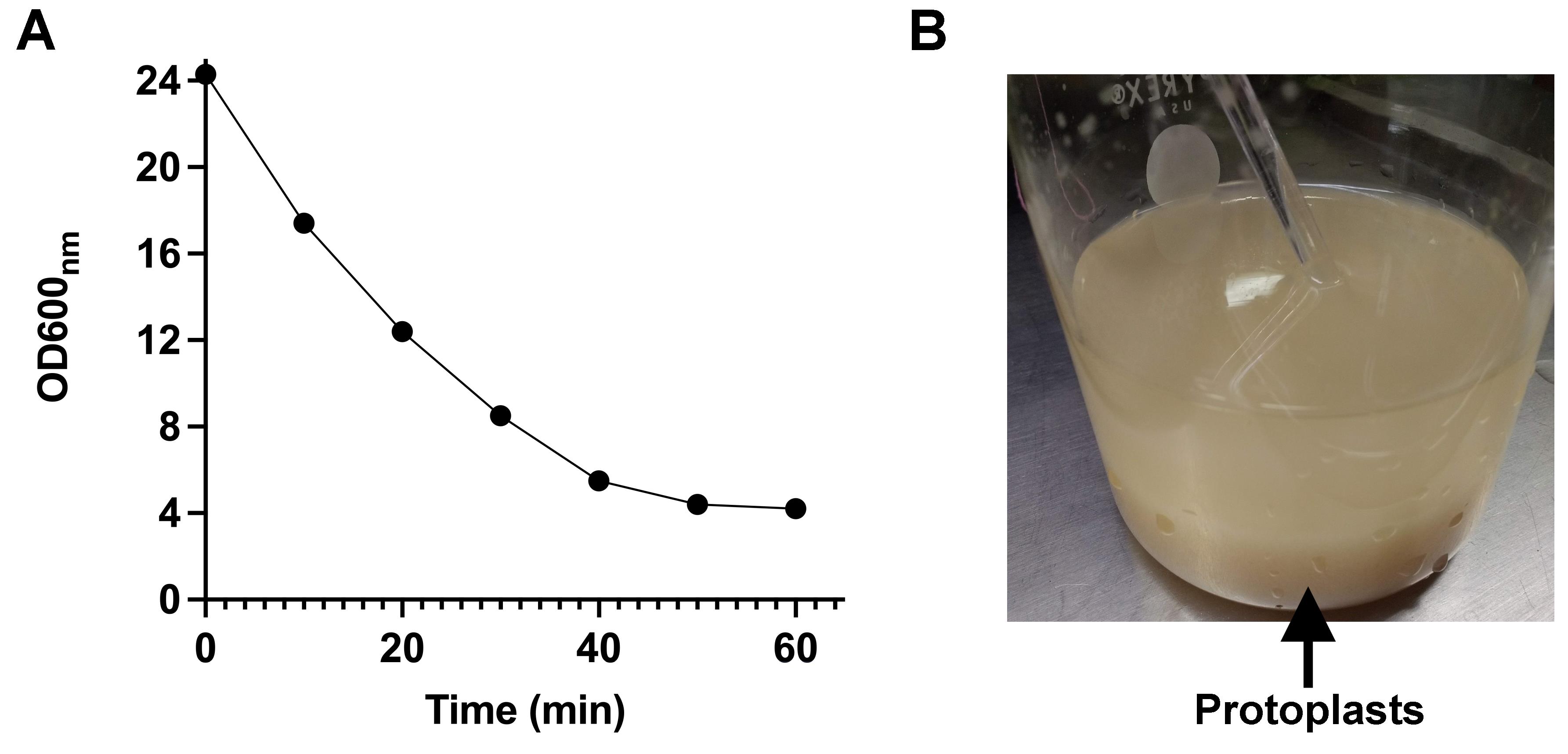
Figure 1. Formation of Ustilago maydis protoplast. During the incubation with lytic enzymes, the OD600nm decreases due to the loss of the yeast cell wall. (A) In the first 30 min of incubation, 50% of the cells are converted to protoplast. (B) Because of the degradation of the cell wall, cells begin to form clumps.Mitochondrial respiratory activity
U. maydis mitochondria contain the four classic respiratory complexes (I-IV), a glycerol-3-phosphate dehydrogenase, a cyanide resistant alternative oxidase (AOX), and external and internal type 2 NADH dehydrogenases (Matuz-Mares et al., 2018; Reyes-Galindo et al., 2019). To determine the activity of complex I (CI), pyruvate-malate (Pyr-Mal) is added; for complex II (CII), use succinate; and use TMPD-ascorbate for complex IV (CIV). The activity of the different complexes can be inhibited by adding rotenone (CI), antimycin A (CIII), or cyanide (CIV) (Juárez et al., 2004).
The addition of succinate or NADH to the mitochondrial suspension stimulated oxygen consumption (Figure 2). Respiratory activity was higher with NADH than with succinate, indicating that the external NADH dehydrogenase is more active than the succinate dehydrogenase. Because of the presence of the AOX, inhibition of respiration by cyanide was not complete. However, in the presence of succinate, there was a time-dependent inactivation of the AOX (Sierra-Campos et al., 2009), probably due to the increased production of H2O2 by some complexes under this condition. Inhibition of AOX by H2O2 was recently reported (Yamasaki et al., 2021).
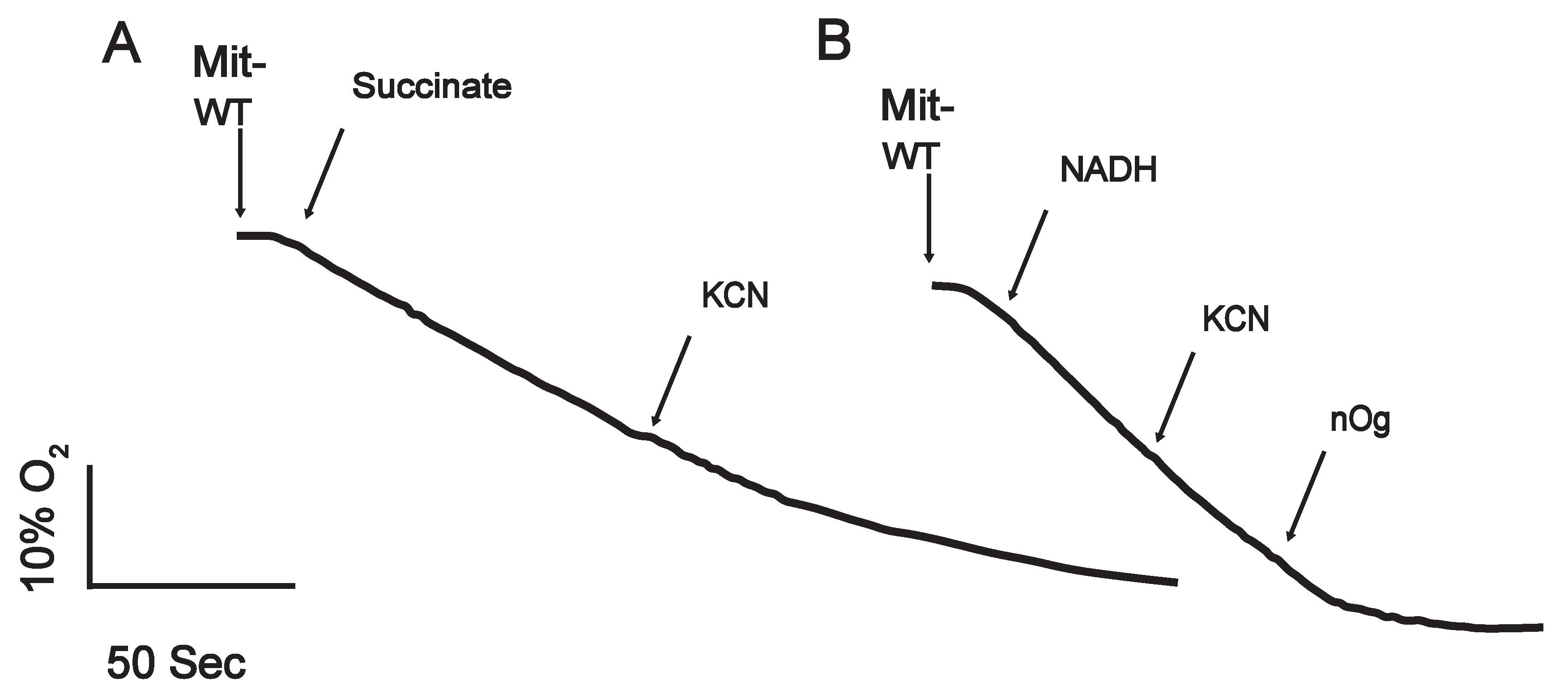
Figure 2. Oxygen consumption by U. maydis mitochondria. Cells were grown in YPD liquid medium for 24 h at 28°C and collected by centrifugation. Then mitochondria were isolated as described in the protocol. Oxygen consumption was stimulated by adding A) succinate (10 mM), or B) NADH (0.15 mM), and the respiratory activity was inhibited by KCN (1 mM), or n-octyl gallate (2 µM).Mitochondrial membrane potential
Mitochondrial membrane potential (ΔΨ) is the main component of the proton motive force generated by respiratory complexes I, III, and IV (Figueira et al., 2012). The cationic lipophilic dye safranine has been used to measure the membrane potential in mitochondria. Upon the generation of the membrane potential by the addition of respiratory substrates, there is an accumulation of safranin inside the mitochondrial matrix, which results in dye stacking and spectral shifts (Akerman and Wikström, 1976). The increase in the 511-533 nm absorbance differences is related to membrane potential generation, while the decrease in this difference induced by the protonophore carbonyl cyanide p-trifluoromethoxyphenylhydrazone (FCCP) is related to the dissipation of ΔΨ.
Figure 3A-3C shows the generation of a ΔΨ by different respiratory substrates in mitochondria obtained from U. maydis protoplasts. Entrance of electrons at the levels of complex I (Pyr-Mal), complex II (succinate), or the external NADH dehydrogenase induces the rapid formation of a ΔΨ, which is associated with an increase in the safranine signal (Figure 3). In all cases, the membrane potential was collapsed by the uncoupler FCCP (Figure 3). Qualitatively, the membrane potential was the same with the three substrates.
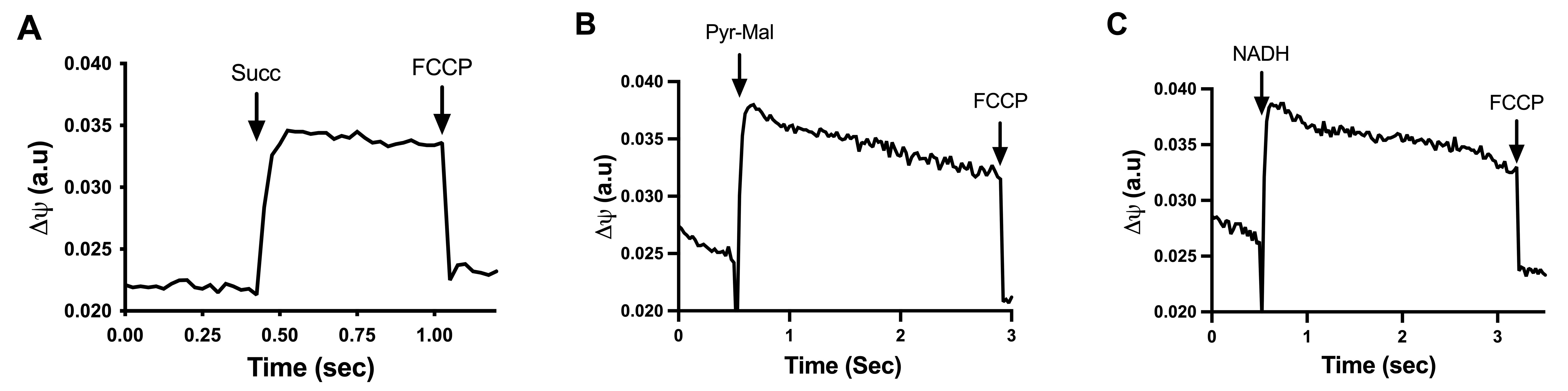
Figure 3. Spectral change of safranine caused by energization of mitochondria. Mitochondria were energized by adding (A) 10 mM succinate (Succ), (B) 5 mM pyruvate-malate, or (C) 250 µM NADH. Then potential was abolished with 10 μM FCCP. The system contained: 300 mM sorbitol, 10 mM Hepes pH 7.0, 0.33 EGTA, 0.5 mg (protein) mitochondria, 0.2% BSA, and 2 μM safranine.
Notes
We recommend incubating cells in the water bath for protoplast formation.
To obtain high-quality mitochondria preparations, we recommended starting with at least 20 g of wet weight of cells, and keeping the temperature at 4°C after protoplast formation.
We recommend a fresh mitochondrial preparation for oxygen consumption experiments.
To achieve inhibition of serine proteases, PMSF should be prepared just before use. Never use a stored PMSF.
Recipes
Solution A
(NH4)2SO4 0.6 M (79.26 g/L)
KH2PO4 20 mM (2.72 g/L)
Solution B
Sucrose 0.8 M (273.84 g/L)
Trizma base-HCl (pH 7.0) 10 mM (1.21 g/L)
EDTA 2 mM (0.74 g/L)
KH2PO4 20 mM (2.72 g/L)
BSA 0.3% (3.0 g/L)
Adjust pH to 7.0 at room temperature.
Solution C
Sucrose 0.4 M (137 g/L)
Trizma base-HCl (pH 7.0) 10 mM (1.21 g/L)
EDTA 2 mM (0.74 g/L)
KH2PO4 20 mM (2.72 g/L)
BSA 0.3% (3.0 g/L)
Adjust pH to 7.0 at room temperature.
PMSF
PMSF 1 M (0.0871 g/0.5 mL of DMSO)
Solution D
Glucose 1 M (1.80 g/10 mL)
Solution E
Tris-HCl (pH 7.0) 250 mM (1.51 g/50 mL)
YPD culture medium
Glucose 0.5% (5 g/L)
Select yeast extract 0.5% (5 g/L)
Bacteriological peptone 0.25% (2.5 g/L)
Sterilize at 120°C for 15 min.
For solid YPD, add 2% agar (20 g/L).
Solution F
Sorbitol 300 mM (54.65 g/L)
Hepes 20 mM, pH 7.0 (4.76 g/L)
EGTA 0.33 mM (125.51 mg/L)
BSA 0.2% (2 g/L)
FCCP solution
FCCP 1 mM (1.27 mg/5 mL)
Safranine solution
Safranine 1 mM (3.51 mg/10 mL)
Succinate solution
Succinate 1 M (1.3507 g/5 mL)
NADH solution
NADH 20 mM (0.07094 g/5 mL)
KCN solution
KCN (0.0815 g/5 mL)
n-Octyl gallate Solution
n Octyl gallate (0.0028 g/5 mL)
Acknowledgments
Author thank to Universidad Nacional Autónoma de México–Programa de Apoyo a Proyectos de Investigación e Inovación Tecnológica (PAPIIT IV200519 e IA200321) and Consejo Nacional de Ciencia y Tecnología (CONACYT-254904). Also, we thanks to Oscar Iván Luqueño Bocardo by the assistance in figures elaboration and Mercedes Esparza Perusquía for technical assistance.
Competing interests
The author declares no competing interest related to this work.
References
- Akerman, K. E. and Wikström, M. K. (1976). Safranine as a probe of the mitochondrial membrane potential. FEBS Lett 68(2): 191-197.
- Brefort, T., Doehlemann, G., Mendoza-Mendoza, A., Reissmann, S., Djamei, A. and Kahmann, R. (2009). Ustilago maydis as a Pathogen. Annu Rev Phytopathol 47: 423-445.
- Figueira, T. R., Melo, D. R., Vercesi, A. E. and Castilho, R. F. (2012). Safranine as a fluorescent probe for the evaluation of mitochondrial membrane potential in isolated organelles and permeabilized cells. Methods Mol Biol 810: 103-117.
- Juárez, O., Guerra, G., Martínez, F. and Pardo, J. P. (2004). The mitochondrial respiratory chain of Ustilago maydis. Biochim Biophys Acta 1658(3): 244-251.
- Lowry, O. H., Rosebrough, N. J., Farr, A. L. and Randall, R. J. (1951). Protein measurement with the Folin phenol reagent. J Biol Chem 193(1): 265-275.
- Matuz-Mares, D., Matus-Ortega, G., Cárdenas-Monroy, C., Romero-Aguilar, L., Villalobos-Rocha, J. C., Vázquez-Meza, H., Guerra-Sánchez, G., Peña-Díaz, A. and Pardo, J. P. (2018). Expression of alternative NADH dehydrogenases (NDH-2) in the phytopathogenic fungus Ustilago maydis. FEBS Open Bio 8(8): 1267-1279.
- Reyes-Galindo, M., Suarez, R., Esparza-Perusquía, M., de Lira-Sánchez, J., Pardo, J. P., Martínez, F. and Flores-Herrera, O. (2019). Mitochondrial respirasome works as a single unit and the cross-talk between complexes I, III2 and IV stimulates NADH dehydrogenase activity. Biochim Biophys Acta Bioenerg 1860(8): 618-627.
- Sierra-Campos, E., Velázquez, I., Matuz-Mares, D., Villavicencio-Queijeiro, A. and Pardo, J. P. (2009). Functional properties of the Ustilago maydis alternative oxidase under oxidative stress conditions. Mitochondrion 9(2): 96-102.
- Sousa, J. S., D’Imprima, E. and Vonck, J. (2018). Mitochondrial Respiratory Chain Complexes. Subcell Biochem 87: 167-227.
- Yamasaki, S., Shoji, M., Kayanuma, M., Sladek, V., Inaoka, D. K., Matsuo, Y., Shiba, T., Young, L., Moore, A. L., Kita, K., et al. (2021). Weak O2 binding and strong H2O2 binding at the non-heme diiron center of trypanosome alternative oxidase. Biochim Biophys Acta Bioenerg 1862(4): 148356.
Article Information
Copyright
© 2022 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Pardo, J. P., Guerra-Sánchez, G. , Flores-Herrera, O. and Romero-Aguilar, L. (2022). Isolation of Mitochondria from Ustilago maydis Protoplasts. Bio-protocol 12(1): e4277. DOI: 10.21769/BioProtoc.4277.
Category
Microbiology > Microbial biochemistry > Protein > Activity
Biochemistry > Protein > Isolation and purification
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link