- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Preparation of Giant Escherichia coli spheroplasts for Electrophysiological Recordings
(*contributed equally to this work) Published: Vol 11, Iss 24, Dec 20, 2021 DOI: 10.21769/BioProtoc.4261 Views: 3711
Reviewed by: Gal HaimovichJerome LacroixChen Fan

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
In-house Fabrication of Nanoplastics of Tunable Composition and Application: Assessment of Bioelectric Changes in Primary Rat Lung Alveolar Epithelial Cell Monolayers Exposed to Nanoplastics
Ricki Chairil [...] Kwang-Jin Kim
Jun 5, 2025 2210 Views
Method for Extracellular Electrochemical Impedance Spectroscopy on Epithelial Cell Monolayers
Athena J. Chien [...] Craig R. Forest
Jun 20, 2025 3292 Views
In Vitro Co-culture of Bacterial and Mammalian Cells to Investigate Effects of Potential Probiotics on Intestinal Barrier Function
Ajitpal Purba [...] Dulantha Ulluwishewa
Jun 20, 2025 2615 Views
Abstract
Prokaryotic ion channels have been instrumental in furthering our understanding of many fundamental aspects of ion channels’ structure and function. However, characterizing the biophysical properties of a prokaryotic ion channel in a native membrane system using patch-clamp electrophysiology is technically challenging. Patch-clamp is regarded as a gold standard technique to study ion channel properties in both native and heterologous expression systems. The presence of a cell wall and the small size of bacterial cells makes it impossible to directly patch clamp using microelectrodes. Here, we describe a method for the preparation of giant E. coli spheroplasts in order to investigate the electrophysiological properties of bacterial cell membranes. Spheroplasts are formed by first inhibiting bacterial cell wall synthesis, followed by enzymatic digestion of the outer cell wall in the presence of a permeabilizing agent. This protocol can be used to characterize the function of any heterologous ion channels or ion transporters expressed in E. coli membranes.
Background
Approximately 30% of all proteins encoded in the human body are membrane proteins (Overington et al., 2006; Murray et al., 2012; He et al., 2014). They play vital roles in regulating cellular homeostasis, neuronal activity, cell metabolism, etc. (Hille, 2001). However, eukaryotic ion channels can be challenging model systems for a variety of reasons, which makes the prokaryotic orthologs a preferred model system for many biophysical studies. In the ion channel field, proteins such as KcsA (Doyle et al., 1998; Cuello et al., 2010; Cordero-Morales et al., 2011), NaK (Shi et al., 2006) and MthK (Jiang et al., 2002 and 2020; Fan et al., 2020) have been used as model systems to gain invaluable information about the structure and function of ion channels.
Studying the biophysical properties of prokaryotic ion channels through electrophysiology is not trivial. Heterologous expression of prokaryotic ion channels in eukaryotic expression systems such as human embryonic kidney cells (HEK 293) and Xenopus laevis (X. Laevis) oocytes is challenging. This is in part due to differences in trafficking mechanisms between prokaryotic and eukaryotic systems (Wagner et al., 2006; Rapoport et al., 2007). The prokaryotic expression system in Escherichia coli (E. coli) can circumvent this issue and it is likely to be a better mimic of the native lipid environment of a prokaryotic ion channel (Wagner et al., 2006).
At present, it is not possible to directly patch clamp E. coli. Like all gram-negative bacteria, E. coli is protected by a cell wall which must be removed in order to access the plasma membrane. Furthermore, a single E. coli is smaller than the tip opening of a typical patch clamp microelectrode. Previously, it has been shown that inhibition of cell wall synthesis in bacteria by cephalexin-treatment leads to the formation of long filamentous ‘snake-like’ structures (Ruthe and Adler, 1985; Martinac et al., 1987; Renner and Weibel, 2011; Sun et al., 2017). Digestion of cephalexin-treated bacteria with lysozyme in the presence of EDTA leads to the formation of giant bacterial spheroplasts (5 to 10 µm), which are suitable for patch clamping studies and have been used successfully to study mechanosensitive ion channels (Martinac et al., 1987). However, this method is not well described and, thus, not routinely used in electrophysiological studies. The following protocol is a summary of previously published protocols with minor modifications (Ruthe and Adler, 1985; Martinac et al., 1987; Mittman et al., 1987; Renner and Weibel, 2011; Randall et al., 2013; Sun et al., 2017). This protocol details the methodology for preparing E. coli spheroplasts for electrophysiological characterization of prokaryotic ion channels. This method can be used for other measurements that require access to the inner bacterial membrane.
Materials and Reagents
1.5 ml microcentrifuge tubes (Fisher, catalog number: 05-408-129)
Pipettes and tips (Eppendorf, catalog number: 2231300002)
Drummond calibrated micropipettes (100 µl) (Drummond Scientific Company, catalog number: 2-000-100)
PCR tubes (Fisher, catalog number: 14-230-225)
50 ml centrifuge tubes (Corning, catalog number: 430829)
Cell culture tubes (Fisher Scientific, catalog number: 14-959-1B)
Serological pipettes (Midsci, catalog numbers: TP94010, TP94005)
Various sizes of Baffled cell culture flasks (Thermo Scientific, catalog numbers: 41100250, 41100250)
10 ml syringes (Becton Dickinson (BD), catalog number: 302995)
E. coli OverExpress C43 (DE3) competent cells (Lucigen, catalog number: 60442-1)
pQE82L expression vector
Liquid nitrogen
Luria Broth Media (RPI Research Products, catalog number: L24400)
Ampicillin Sodium Salt (Sigma Aldrich, catalog number: A0166)
Cephalexin hydrate (Alfa Aesar, catalog number: J63672-06)
Isopropyl β-D-1-thiogalactopyranoside (Gold Bio, catalog number: I2481C)
Sucrose (Sigma-Aldrich, catalog number: S0389)
Tris base (Sigma-Aldrich, catalog number: 1070897600)
Ready-Lyse Lysozyme solution (Lucigen, catalog number: R1804M)
OmniCleave Endonuclease (Lucigen, catalog number: OC7850K)
Ethylenediaminetetraaceticacid (Sigma- Aldrich, catalog number: E9884)
Magnesium Chloride (Sigma-Aldrich, catalog number: M8266)
Potassium Chloride (Sigma-Aldrich, catalog number: 529552)
Calcium Chloride (Sigma-Aldrich, catalog number: C8106)
Potassium Hydroxide (Sigma-Aldrich, catalog number: 221473)
Hydrochloric acid (Acros organics, catalog number: AC450550025)
LB medium (see Recipes)
Stop buffer (see Recipes)
Pipette solution (Intracellular solution) (see Recipes)
Bath solution (see Recipes)
Equipment
-80°C freezer
Water bath
Incubating shaker with temperature control (New Bruinswick Scientific, Model C25KC classic series incubator shaker)
Spectrophotometer/Nanodrop
Autoclave
Centrifuge (Eppendorf, model: 5810R)
Deionized water (18.2 µΩ resistance)
pH meter (Thermo Scientific Orion Star A214)
Electrophysiology setup:
Air table (TMC)
Fluorescence microscope (Olympus, model: IX70)
Micromanipulator (Scientifica, model: ACCi UI)
Digidata 1440A (Axon Instruments)
Multiclamp (Axon Instruments)
Perfusion system (home-made gravity perfusion system)
Chamber (Warner Instruments, catalog number: RC26)
Incubator for bacterial growth
Pipette puller (Sutter Instruments, model: P-97)
Phase contrast Microscope (Invitrogen, model: EVOS M5000)
Software
pCLAMP and Clampfit (Molecular devices)
Procedure
Making the spheroplasts
Bacterial transformation
Take an aliquot (40 µl) of E. coli OverExpress C43 (DE3) competent cells from -80°C freezer and place on ice for approximately 5 min to let it thaw.
Then, add 1 ng of plasmid containing gene of interest. We used pQE82L expression vector containing MthK-IR (Inactivation removed calcium activated potassium channel) gene to the bacteria and gently flick the tube to mix them well. Let it incubate on ice for about 30 min.
After incubation, perform heat-shock on the DNA-competent cell mixture by placing in a 42°C water bath for 45 s and then put the tube back on ice for another 5 min before spreading it on LB Agar plates containing appropriate antibiotic (ampicillin in this case).
Incubate the plate at 37°C overnight.
Bacterial growth
The next day (16 h after transformation), pick a single colony from the plate and inoculate it into 20 ml LB medium with antibiotic held in a 60 ml cell culture flask.
Incubate the culture in a shaker at 37°C and 220 rpm. After 3 h, check the optical density (OD600) of the culture periodically (every 20-30 min) with a spectrophotometer or nanodrop until OD600 reaches ~0.7.
Cephalexin treatment
Pre-heat 55 ml of autoclaved LB medium at 42°C. Once the culture reaches OD600 = 0.7, place the culture on ice, quickly take 5 ml of this liquid culture and add to 55 ml of pre-heated LB medium containing antibiotic. Add 50 μg/ml cephalexin.
Put the culture in an incubator at 42°C and reduce the speed of the shaker to 120 rpm (‘snake-like’ E. coli filaments are really fragile).
Remove 100 μl of the culture every 20 min and observe under a microscope to check the efficiency of cephalexin treatment (Figure 1A).
Based on our experience, this process usually takes about 2 h. Once the snake-like filaments are longer than 100 µm, leave the culture at room temperature. Drop the temperature of the shaker to 37°C (typically takes about 15 min).
Add 0.8 mM IPTG to the culture and place it back in the incubator at 37°C and 120 rpm for another hour.
Spheroplast preparation
Following induction, divide the 60 ml cell culture in equal volumes into three 50 ml centrifuge tubes. Centrifuge the culture at low speed 453 × g at 4°C for 15 min. Discard the supernatant carefully, as the bacteria does not attach to the bottom of the tube firmly. Place the three tubes with bacterial pellets on ice and resuspend each of the pellets gently in 3 ml of 1 M sucrose.
Add 240 μl 1 M Tris, pH 8.0, 67,500 units of Ready-Lyse Lysozyme solutions (Lucigen), 1,600 units OmniCleave Endonuclease (Lucigen), and 20 μl of 500 mM EDTA to the resuspended cells and incubate at room temperature. After 10 min of incubation, add 1 ml of stop buffer (see Recipes) to one of the three tubes and mix by gently swirling the tube. Place the tube on ice. Similarly, add the stop solution to the other two tubes after 20 min and 40 min of incubation, respectively, and mix by gently swirling the tubes. Place all tubes on ice immediately after adding the stop buffer. Take 20 μl of the solution from each spheroplast preparation and examine under the microscope to identify the optimal spheroplasts preparation out of the three, as shown in Figure 1B.
Under a phase contrast microscope, the well prepared spheroplasts appear like dark gray circles (Figure 1C). Choose the preparation with the highest number of dark gray spheroplasts. Aliquot the spheroplasts into PCR tubes (50 μl per tube), flash freeze in liquid nitrogen, and store at -80°C until further use. If stored at -80°C, the spheroplasts can be used for recording for 6 months.
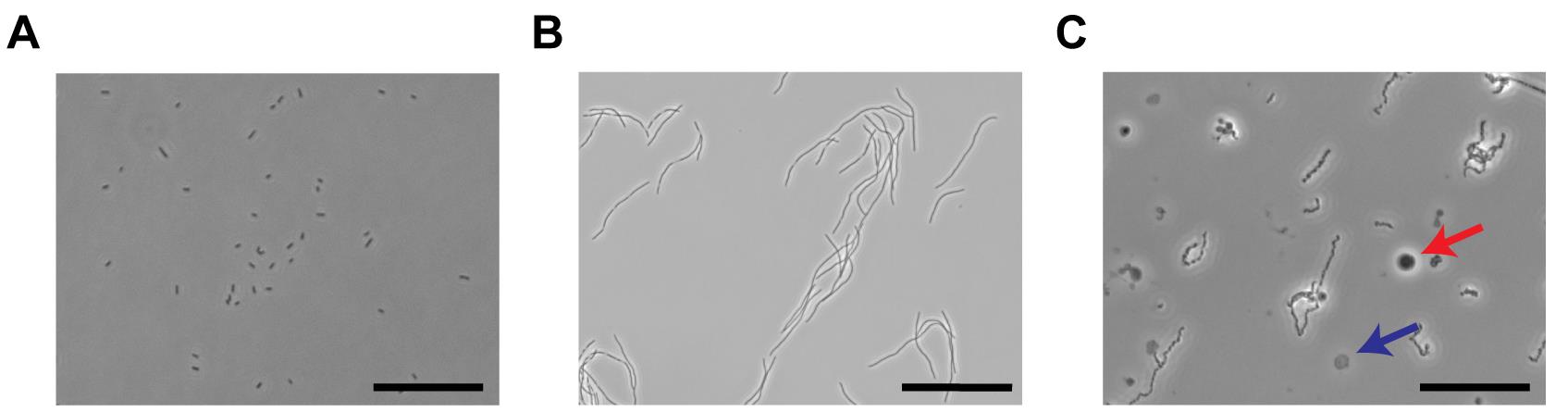
Figure 1. Illustration of spheroplasts preparation. A. An image of E. coli cell culture before cephalexin treatment. The scale bar equals 50 µm. B. Illustration of ‘snake-like’ bacterial filaments after 2 h of 50 μg/ml cephalexin treatment under phase contrast microscope. Scale bar is 100 µm. C. Phase contrast image showing spheroplasts after the digestion of E. coli outer membrane with lysozyme and EDTA. The dark gray circle (red arrow) is a good spheroplast and can form giga-ohm seals while the transparent one (blue arrow) is not suitable for patching. The scale bar indicates 50 µm.
Patch clamping the spheroplasts
Solutions: Bacteria have a higher cytosolic osmolarity compared to cell lines such as HEK293 or X. Laevis oocytes. In our experiments, we recorded currents from Mthk, which is a calcium-activated potassium channel, and therefore the pipette solution (Intracellular solution) (see Recipes) and the bath solution (see Recipes) have differernt calcium concentrations. The osmolarity for different concentrations of calcium should be adjusted with sucrose accordingly.
Glass electrodes: We chose 100 µl Drummond calibrated micropipettes. The micropipettes are made of N51A borosilicate glass and have an internal diameter of 1.3 mm and outer diameter of 1.68 mm. Pull the pipettes with an electrode puller (Sutter Instruments Model P97), so that the bubble number (described below) is between 5.0-5.5.
As the E. coli spheroplasts are fairly small in size (5-10 µm), the tip size of the glass electrodes needs to be approximately 1 μm, which is too small to be accurately evaluated under a light microscope. Thus, to calibrate the tip size of the glass electrodes, which is extremely important for the success rate of patching spheroplasts, ‘bubble number’ should be used as a standard (Mittman et al., 1987). Pull a glass electrode and connect it to a 10 ml syringe with an aspirator for forming an airtight seal as shown in Figure 2A. Submerge the micropipettes into methanol to about 1 cm deep and slowly press the plunger from 10 ml graduation until a fine stream of bubbles appears in the methanol (Figure 2B, 2C). The volume read out at this moment is the ‘bubble number’ (or the volume of air remaining in the syringe is the bubble number) of the micropipette and this number should ideally be between 5.0 to 5.5. In our experience, a ‘bubble number’ between 5.5 to 6 is also usable (see Video 1). If the bubble number is outside this range, the electrode pulling program should be modified accordingly until the tip size falls in this range.
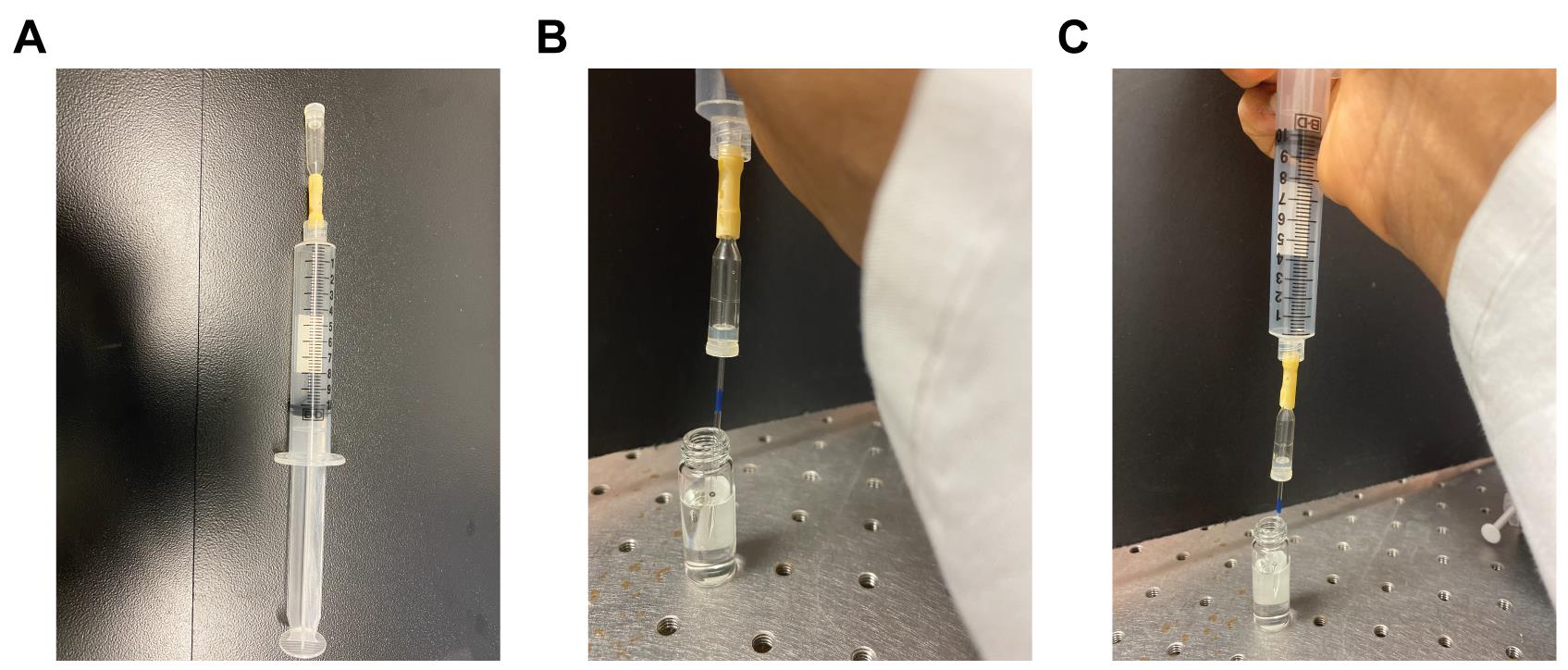
Figure 2. Bubble number measurement. A. Illustration of a 10 ml syringe connected with an aspirator. B. Submerge the tip of the micropipette into a vial containing methanol with the plunger set at 10 ml. C. Apply positive pressure on the plunger. Watch for the first appearance of bubbles in the methanol. The corresponding reading on the syringe is the bubble number. Pipettes with a bubble number between 5.0 and 5.5 are ideal for patching spheroplasts.Video 1. Bubble number measurement.Spheroplast patching
Note: It is very important to keep the glass tip clean to form a giga-ohm seal, so a positive air pressure should be applied before the electrode is immersed into the bath solution.
Spheroplasts do not stick to the bottom of the glass surface of the chamber. If the positive pressure applied is too large, then the spheroplasts are pushed away. To overcome this problem, use a very low positive pressure (5 to 10 millibars of pressure is sufficient). Alternatively, release the pressure right before you approach the spheroplast. Another related issue is that it is difficult to see the dent on the spheroplast caused by the positive pressure from the glass electrode tip; while we do not have a solution for this, with some practice researchers can get a feeling of when to apply a negative pressure to form a giga-ohm seal with the electrode.
Finding a good spheroplast
When imaged with a phase contrast microscope, a small percentage of spheroplasts will appear to have lost their interior phase contrast. Based on our experience, these types of spheroplasts could not form a giga-ohm seal with the glass electrode.
Forming an inside-out patch: Once the giga-ohm seal is formed, move the glass microelectrode away from the bottom of the chamber (Caution: do not move it out of the bath solution). Apply a sharp but gentle movement on the head-stage by flicking it. This would apply a dramatic shear force between the tip of the microelectrode and the spheroplast. Sometimes this step will cause a loss of the giga-ohm seal.
Recipes
LB medium
25 g granulated LB
1 L of ddH2O
Stop buffer
0.8 M sucrose
80 mM Tris, pH 8.0
12 mM MgCl2
Pipette solution (Intracellular solution)
150 mM KCl
20 mM MgCl2
15 mM Tris, pH 8.0
0.1 mM CaCl2
450 mM sucrose
Bath solution
150 mM KCl
20 mM MgCl2
15 mM Tris, pH 8.0
Acknowledgments
We would like to thank Dr. V.A. Klenchin for his help on various aspects of this protocol. We would like to thank Dr. Sergei Sukharev at the University of Maryland for his help with patch clamping bacterial spheroplasts. We thank National Institutes of Health (NS-116850, NS-101723, NS-081293, GM131662), McDonnell Center for Neuroscience, CIMED and Department of Anesthesiology for research support. This protocol was derived from the original research paper “Activation of the archaeal ion channel MthK is exquisitely regulated by temperature” (Jiang et al., 2020).
Competing interests
There are no conflicts of interest or competing interests.
References
- Cordero-Morales, J. F., Jogini, V., Chakrapani, S. and Perozo, E. (2011). A multipoint hydrogen-bond network underlying KcsA C-type inactivation. Biophys J 100(10): 2387-2393.
- Cuello, L. G., Jogini, V., Cortes, D. M. and Perozo, E. (2010). Structural mechanism of C-type inactivation in K+ channels. Nature 466(7303): 203-208.
- Doyle, D. A., Morais Cabral, J., Pfuetzner, R. A., Kuo, A., Gulbis, J. M., Cohen, S. L., Chait, B. T. and MacKinnon, R. (1998). The structure of the potassium channel: molecular basis of K+ conduction and selectivity. Science 280(5360): 69-77.
- Fan, C., Sukomon, N., Flood, E., Rheinberger, J., Allen, T.W. and Nimigean, C. M. (2020). Ball-and-chain inactivation in a calcium-gated potassium channel. Nature 580(7802): 288-293.
- He, Y., Wang, K. and Yan, N. (2014). The recombinant expression systems for structure determination of eukaryotic membrane proteins. Protein Cell 5(9): 658-672.
- Hille, B. (2001). Ion channels of excitable membranes. Sunderland, Mass: Sinauer.
- Jiang, Y., Idikuda, V., Chowdhury, S. and Chanda, B. (2020). Activation of the archaeal ion channel MthK is exquisitely regulated by temperature. Elife 9: e59055
- Jiang, Y., Lee, A., Chen, J., Cadene, M., Chait, B.T. and MacKinnon, R. (2002). Crystal structure and mechanism of a calcium-gated potassium channel. Nature 417(6888): 515-522.
- Murray, C.W., Verdonk, M.L. and Rees, DC. (2012). Experiences in fragment-based drug discovery. Trends Pharmacol Sci 33(5): 224-232.
- Martinac, B., Buechner, M., Delcour, A.H., Adler, J. and Kung, C. (1987). Pressure-sensitive ion channel in Escherichia coli. Proc Natl Acad Sci U S A 84(8): 2297-2301.
- Mittman, S., Flaming, D. G., Copenhagen, D. R. and Belgum, J. H. (1987). Bubble pressure measurement of micropipet tip outer diameter. J Neurosci Methods 22(2):161-166.
- Randall, C. P., Mariner, K. R., Chopra, I. and O'Neill, A. J. (2013). The target of daptomycin is absent from Escherichia coli and other gram-negative pathogens. Antimicrob Agents Chemother 57(1): 637-639.
- Rapoport, T. A. (2007). Protein translocation across the eukaryotic endoplasmic reticulum and bacterial plasma membranes. Nature 50(7170): 663-669.
- Renner, L. D. and Weibel, D. B. (2011). Cardiolipin microdomains localize to negatively curved regions of Escherichia coli membranes. Proc Natl Acad Sci U S A 108(15): 6264-6269.
- Ruthe, H. J. and Adler, J. (1985). Fusion of bacterial spheroplasts by electric fields. Biochim Biophys Acta 819(1):105-113.
- Overington, J. P., Al-Lazikani, B. and Hopkins, A. L. (2006). How many drug targets are there? Nat Rev Drug Discov 5(12): 993-996.
- Shi, N., Ye, S., Alam, A., Chen, L. and Jiang, Y. (2006). Atomic structure of a Na+- and K+-conducting channel. Nature 440(7083): 570-574.
- Sun, Y., Sun, T. L. and Huang, H. W. (2014). Physical properties of Escherichia coli spheroplast membranes. Biophys J 107(9): 2082-2090.
- Wagner, S., Bader, M. L., Drew, D. and de Gier, J. W. (2006). Rationalizing membrane protein overexpression. Trends Biotechnol 24(8): 364-371.
Article Information
Copyright
Jiang et al. This article is distributed under the terms of the Creative Commons Attribution License (CC BY 4.0).
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Jiang, Y., Idikuda, V. and Chanda, B. (2021). Preparation of Giant Escherichia coli spheroplasts for Electrophysiological Recordings. Bio-protocol 11(24): e4261. DOI: 10.21769/BioProtoc.4261.
- Jiang, Y., Idikuda, V., Chowdhury, S. and Chanda, B. (2020). Activation of the archaeal ion channel MthK is exquisitely regulated by temperature. Elife 9: e59055
Category
Cell Biology > Cell-based analysis > Electrophysiological technique
Biophysics > Electrophysiology > Patch-clamp technique
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










