- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Isolation of Plant Nuclei Compatible with Microfluidic Single-nucleus ATAC-sequencing
Published: Vol 11, Iss 23, Dec 5, 2021 DOI: 10.21769/BioProtoc.4240 Views: 5014
Reviewed by: Wenrong HeYuan WangTravis Lee

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Rice Meiotic Chromosome Spread Preparation of Pollen Mother Cells
Xingwang Li and Changyin Wu
Jul 20, 2014 11724 Views
Telomere-mediated Chromosomal Truncation via Agrobacterium tumefaciens or Particle Bombardment to Produce Engineered Minichromosomes in Plants
Nathaniel D. Graham [...] James A. Birchler
Sep 20, 2015 9232 Views
CAPS-Based SNP Genotyping for Nitrogen-Response Phenotypes in Maize Hybrids
Jannis Jacobs [...] Peter K. Lundquist
Dec 20, 2025 543 Views
Abstract
Gene expression depends on the binding of transcription factors with DNA regulatory sequences. The level of accessibility for these sequences varies between cells and cell types. Until recently, using the Tn5 assay for transposase-accessible chromatin for sequencing (ATAC-seq) technology allowed assessing the profiles of chromatin from an entire organ or, when coupled with the isolation of nuclei tagged in specific cell types (INTACT) method, from a cell-type. Recently, ATAC-seq experiments were conducted at the level of individual plant nuclei. Applying single nuclei ATAC-seq (sNucATAC-seq) technology to thousands of individual cells revealed more finely tuned profiles of chromatin accessibility. In this manuscript, we describe a method to isolate nuclei fom plant roots and green tissues, permeabilize the nuclear membrane using detergent to allow the penetration of the Tn5 transposase, and re-suspend them in a nuclei resuspension buffer compatible with the construction of sNucATAC-seq libraries using the 10× Genomic’s Chromium technology. This protocol was successfully applied on Arabidopsis thaliana and Glycine max root nuclei.
Background
The assay for transposase-accessible chromatin using sequencing (ATAC-seq) is a popular technology to map DNA accessible sites and to identify putative regulatory elements (Lu et al., 2017; Huang et al., 2021). The accessibility of these elements plays an important role in controlling gene expression because they are transcription factor-binding sites. One major limitation associated with the use of ATAC-seq technology is the cellular complexity of the sample used. When working at the entire organ level, the profile of chromatin accessibility cannot reveal the differential and unique contributions of each cell and cell type composing the organ. To overcome this challenge, recent studies successfully tested ATAC-seq technology on thousands of individual plant nuclei isolated from Arabidopsis thaliana roots and various Zea mays organs (Dorrity et al., 2021; Farmer et al., 2021; Marand et al., 2021).
The integration of this molecular information with RNA-seq datasets generated from isolated protoplasts and nuclei revealed the impact of chromatin accessibility and regulatory elements in controlling gene expression (Denyer et al., 2019; Jean-Baptiste et al., 2019; Nelms and Walbot, 2019; Ryu et al., 2019; Shulse et al., 2019; Zhang et al., 2019; Dorrity et al., 2021; Farmer et al., 2021; Liu et al., 2020; Marand et al., 2021). Therefore, single-cell/nucleus RNA-seq and single nuclei ATAC-seq (sNucATAC-seq or snATAC-seq) have recently changed our assessment of the molecular mechanisms controlling gene expression. These technologies will also contribute to a better understanding of gene function to control plant cell differentiation, development, and plant cell response to environmental stresses (Rich-Griffin et al., 2020; Shaw et al., 2021).
The SNucATAC-seq technology relies on the permeabilization of the membrane of isolated plant nuclei to enable the penetration of the Tn5 transposase, the tagmentation of the accessible genomic DNA sequences, and the construction of Illumina-compatible libraries. In this manuscript, we describe the methodology used to conduct sNucATAC-seq. This method complements the protocol describing the implementation of single nuclei RNA-seq technology on plant samples (Thibivilliers et al., 2020). Here, we describe a protocol to isolate and purify nuclei from model plants and crop species before permeabilizing the nuclear membrane to enable the penetration of the Tn5 transposase (Figure 1). Examples of successful sNucATAC-seq profiles from Arabidopsis thaliana and Glycine max are also provided.
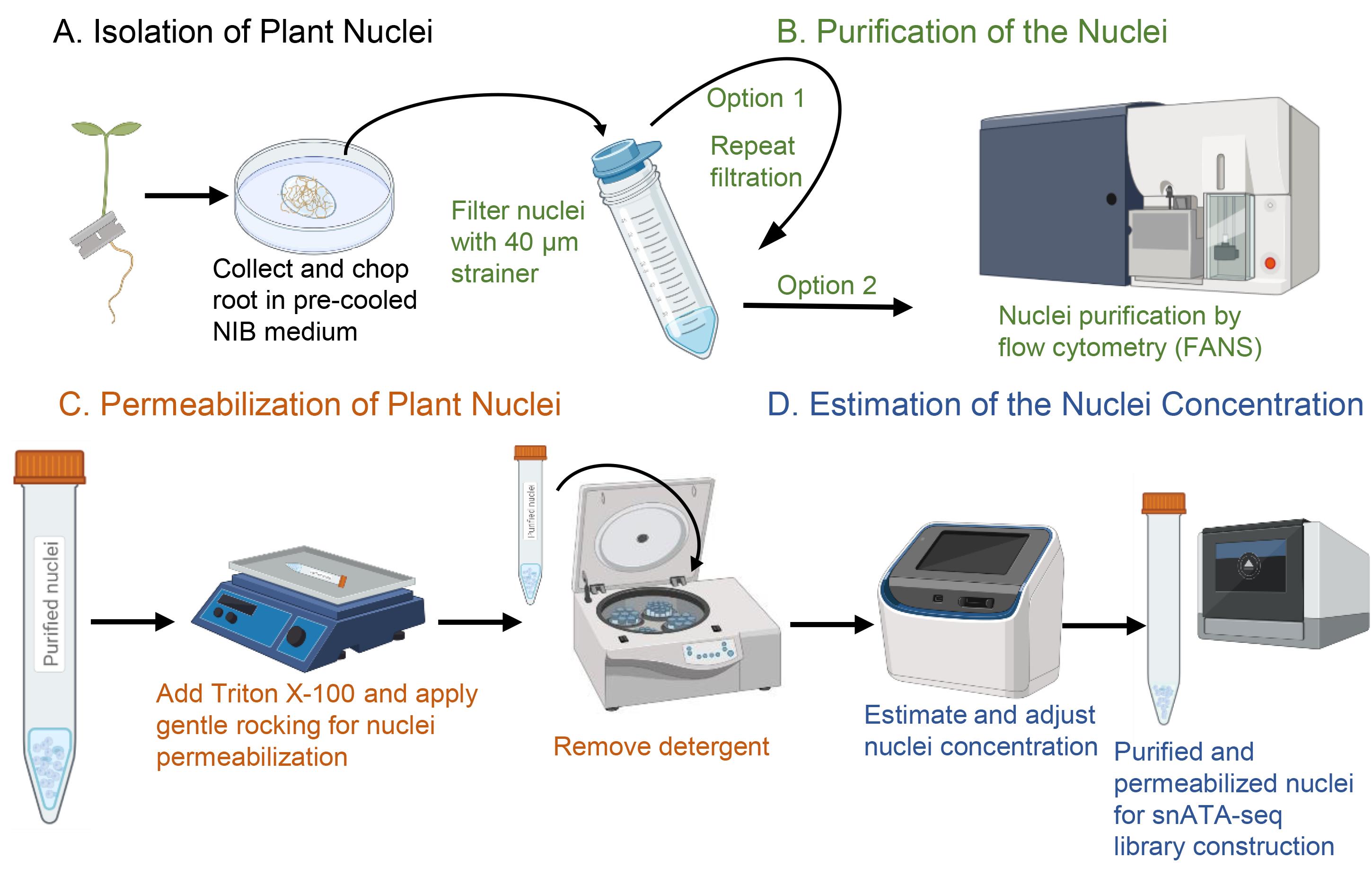
Figure 1. Workflow of nuclei isolation, purification, and permeabilization compatible with single-nucleus ATAC-seq library construction (created in BioRender.com).
Materials and Reagents
60 mm Petri dishes (Corning Life Sciences, catalog number: 08757100B)
Single edge extra keen razor blade (Electron Microscopy Sciences, catalog number: 71962)
50 ml Falcon conical tubes (Corning, catalog number: 352070)
15 ml Falcon conical tubes (Corning, catalog number: 430790)
MACS SmartStrainers 30 µm (Miltenyi Biotec, catalog number: 130-098-458)
40 µm sterile cell strainers (Corning, catalog number: 07-201-430)
Mini 20 µm cell strainers (pluriStrainer, catalog number: 43-10020-50)
Micro slide (Corning, catalog number: 2940-75x25)
Cover glass (Thermo Scientific, catalog number: 12450S)
Eppendorf microcentrifuge tube (Fisher Scientific, catalog number: 05-402-25)
Pipette tips [Rainin RT LTS, catalog numbers: 30389226 (10 µl), 30389240 (200 µl), 30389213 (1,000 µl), 30389218 (1,000 µl wide-orifice)]
CelLyticTM PN Isolation/Extraction buffer 4× (Sigma-Alrich, catalog number: CELLYTPN1) is diluted with DNAse free water to reach a final working concentration of 1×. We name this diluted buffer “NIB medium.”
Nuclease-free water (Invitrogen, catalog number: AM9937)
Phosphate-buffered saline (PBS) without calcium and magnesium (Corning, catalog number: 21-040-CV)
Bovine serum albumin 50 mg/ml (BSA; Invitrogen, catalog number: AM2616)
7-AAD (eBioscience, catalog number: 00-6993-50)
SYTOX Green nucleic acid stain (Invitrogen, catalog number: S7020)
Triton X-100 (Sigma, catalog number: T8787-100ml)
Nuclei buffer 20× (10× Genomics, catalog number: 2000207; part of the Chromium Next GEM Single Cell ATAC Library Kit v1.1, PN-1000163/4)
Trypan blue stain 0.4% (Invitrogen, catalog number: T10282)
1% BSA-PBS (see Recipes)
NIB medium supplemented with 2% Triton X-100 (see Recipes)
1% BSA-PBS supplemented with 2% Triton X-100 (see Recipes)
Equipment
Dissecting Forceps, Fine Tip (VWR, catalog number: 82027-386)
Countess cell counting chamber slide (Invitrogen, catalog number: C10228)
BD FACSAria II cell sorter (BD Biosciences, San Jose, CA)
Microvolume pipettes (Rainin, models: Lite – XLS, P1000, P200, P20)
Platform Rocker (Fisherbrand, catalog number: 13-687-704)
Sorvall Legend X1R Centrifuge with Swinging Bucket Rotor (Thermo Scientific, catalog number: 75004261)
Countess II Automated Cell Counter (Invitrogen, catalog number: AMQAX1000)
Procedure
Thirty minutes before collecting the plant sample(s), cool down all materials and reagents to 4°C (i.e., Petri dishes, razor blades, tips, tubes, cell strainers). It is critical to keep the samples and the isolated nuclei at 4°C throughout the entire process, to reduce stress levels and maximize the integrity of the nuclear membrane. Ultimately, a decrease in nuclear membrane integrity would generate a higher rate of nuclei bursting during library construction, a lower recovery rate of nuclei, and a decrease in the yield of sNucATAC-seq DNA fragments per nucleus.
Isolation of plant nuclei
Collect between 100 to 1,000 mg of fresh plant sample(s) and place it in a 60 mm Petri-dish with 500 μl of pre-cooled NIB medium.
Note: This assay was successfully tested on 1 g Arabidopsis roots grown for 7 days on MS medium under continuous light, and on soybean roots grown for 6 days on B&D medium in the dark (for more details about growth conditions, see Pingault et al., 2018).
Chop the plant tissue in the NIB medium with a razor blade for about 30 s.
Cover the chopped plant tissues with pre-cooled NIB medium to keep the sample moist (on average, 1 ml of NIB medium is needed for 1 g of chopped plant tissue). However, avoid using excessive volumes of NIB medium because it would make chopping the sample difficult.
Finely chop the plant tissue for an additional 4 min (i.e., as a result, the chopped sample should be easy to pipette using a 1 ml wide-bore tip)
Place a 40 μm strainer on top of a 50 ml conical tube tilted on ice. Pre-moisten it with 500 μl of pre-cooled NIB medium. This volume will also serve as a cushion at the bottom of the conical tube for the nuclei.
Using 1 ml wide-bore tips, gently transfer the nuclei-NIB medium mixture to the top of the strainer.
Add 1 ml of pre-cool NIB in the 60mm Petri dish to wash and collect the remaining nuclei. Gently transfer this volume with 1 ml wide-bore tips to the 40 μm strainer.
Purification of the nuclei
When working with plant nuclei that are less than 30 µm in diameter (e.g., A. thaliana root nuclei), use Option 1. If the nuclei diameter is above 30 µm, use Option 2. These additional purification steps are strongly recommended to remove large sized debris before initiating the construction of the sNucATAC-seq libraries. Debris may clog the microfluidic system during the formation of oil-based microbeads, preventing the emulsion phase and leading to a partial or complete loss of the library.
Option 1: No Fluorescent-Activated Nuclei Sorting (FANS) purification
Prepare another 50 ml conical tube as described in Step A5, but using a 30 μm strainer.
Using 1ml wide-bore tips, gently transfer the filtered nuclei-NIB medium mixture from Step A7 onto the 30 μm strainer.
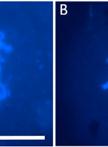
Perform a first nuclei-quality check under the microscope; check for contamination by debris (Figure 2):
Gently swirl the 50 ml conical tube containing the nuclei mixture and pipette 10 μl of this onto a Micro slide.
Add SYTOX Green DNA staining at a final concentration of 0.5 μM. Cover the Micro slide with a cover glass.
Using an epifluorescent microscope, look for potential contamination of the nuclei suspension with cellular debris. To remove the remaining debris, filter the nuclei resuspension using another 30 μm or, when possible, a 20 μm strainer as described in Option 1 – steps 1 and 2.
Upon validating the quality and purity of the isolated nuclei, proceed to Step C. Permeabilization of the nuclei.
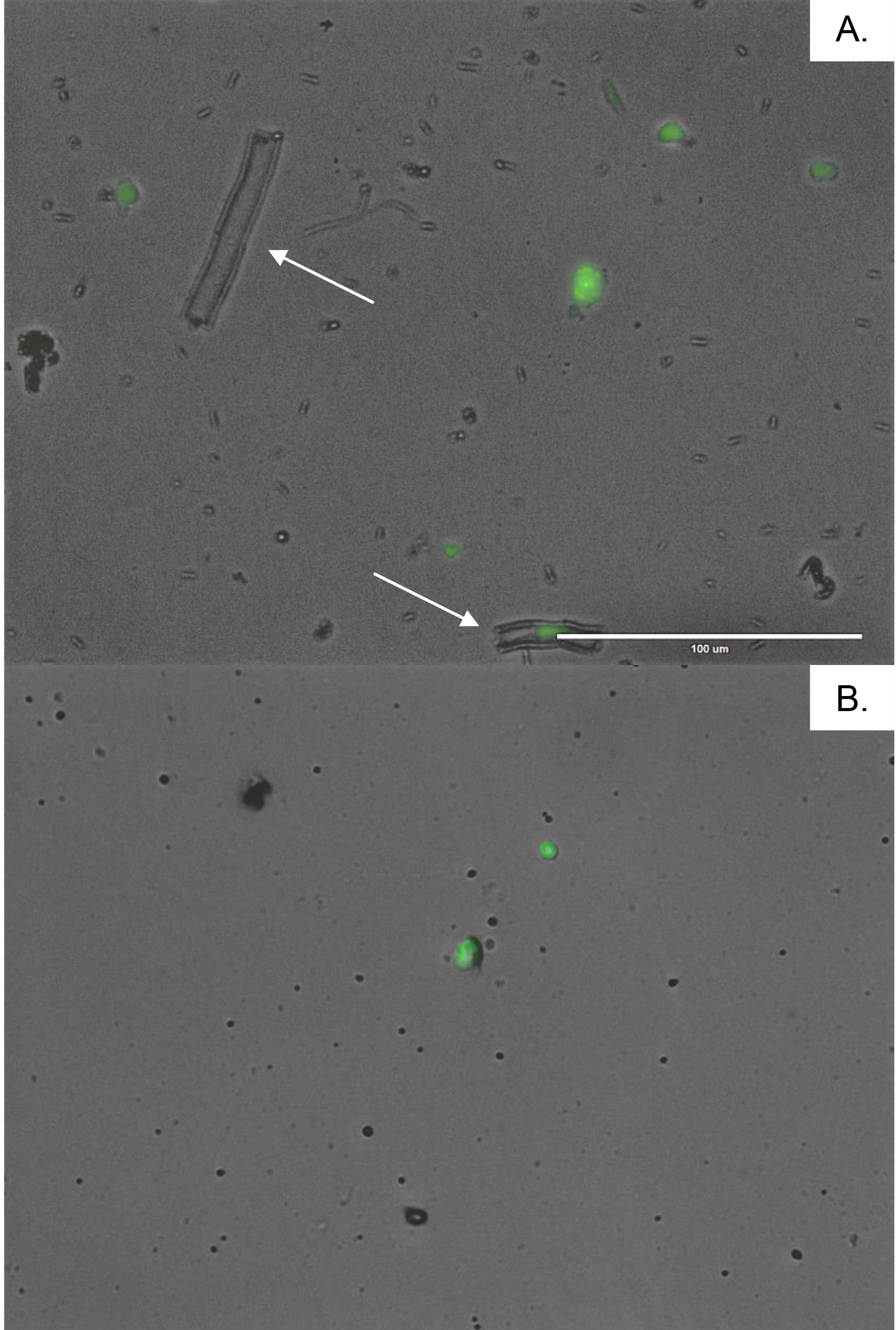
Figure 2. Microscopic pictures (objective 40×) of plant nuclei before (A) and after (B) purification with 20 µm cell strainer. The nuclei were stained with SYTOX Green. The white arrows indicate debris over 40 μm that might clog the microfluidic system. Scale bar: 100 μm.
Option 2: FANS selection
Set up the cell sorter.
Use a 100 µm nozzle and the default system pressure of 20 psi to minimize physical stresses on the nuclei during their sorting.
Cool down the sample and collection chambers to 4°C before and during the sort.
Transfer and sort 200 µl of unstained nuclei to establish the baseline of the nuclei sorting.
To sort the plant nuclei, add the DNA intercalant 7-AAD to the nuclei suspension at the ratio v/v 1/40 (i.e., 25 μl of 7-AAD stock solution per 1 ml of nuclei mixture). Incubate on ice for at least 30 min before sorting.
Note: To date, 7-AAD is the only dye validated by 10× Genomics to conduct sNucATAC-seq experiments. According to 10× Genomics, 7-ADD does not interfere with the activity of the Tn5 transposase, as opposed to other DNA staining dyes (https://kb.10xgenomics.com/hc/en-us/articles/360027640311-Can-I-sort-nuclei-for-Single-Cell-ATAC-sequencing-or-Single-Cell-Multiome-ATAC-GEX-).
Sort the sample using a BD FACSAria II cell sorter (BD Biosciences, San Jose, CA; Figure 3) or cell sorters with similar spectral properties.
Excite the 7-AAD signal with a 561 nm laser, detecting its emission with a combination of a 635 long pass (LP) filter and a 660/20 band pass (BP) filter.
Display and gate the initial signal on a forward versus side scatter (FSC vs. SSC), to remove unwanted non-nuclei events and debris. Subsequently, display the data on a 7-AAD height (7-AAD – H) vs. 7-AAD width (7-AAD – W) plot to allow for the removal of additional contaminants (Figure 3A-3B).
It can be difficult to differentiate 7-AAD stained nuclei from the autofluorescing organelles. Using a bivariate plot displaying 7-AAD vs. an adjacent fluorescence (i.e., 488 nm excitation laser and 600LP and a 610/20BP filter set), we were able to better discriminate the 7-AAD positive nuclear fraction of the sample (Figure 3C) vs. organelles (Figure 3D). In this configuration, the nuclear fraction shifts slightly away from the organelles (Figure 3C vs. 3D), allowing the stained nuclei to be gated and sorted. An example of the profile of DNA content of the sorted nuclei using this strategy is provided in Figure 3E.
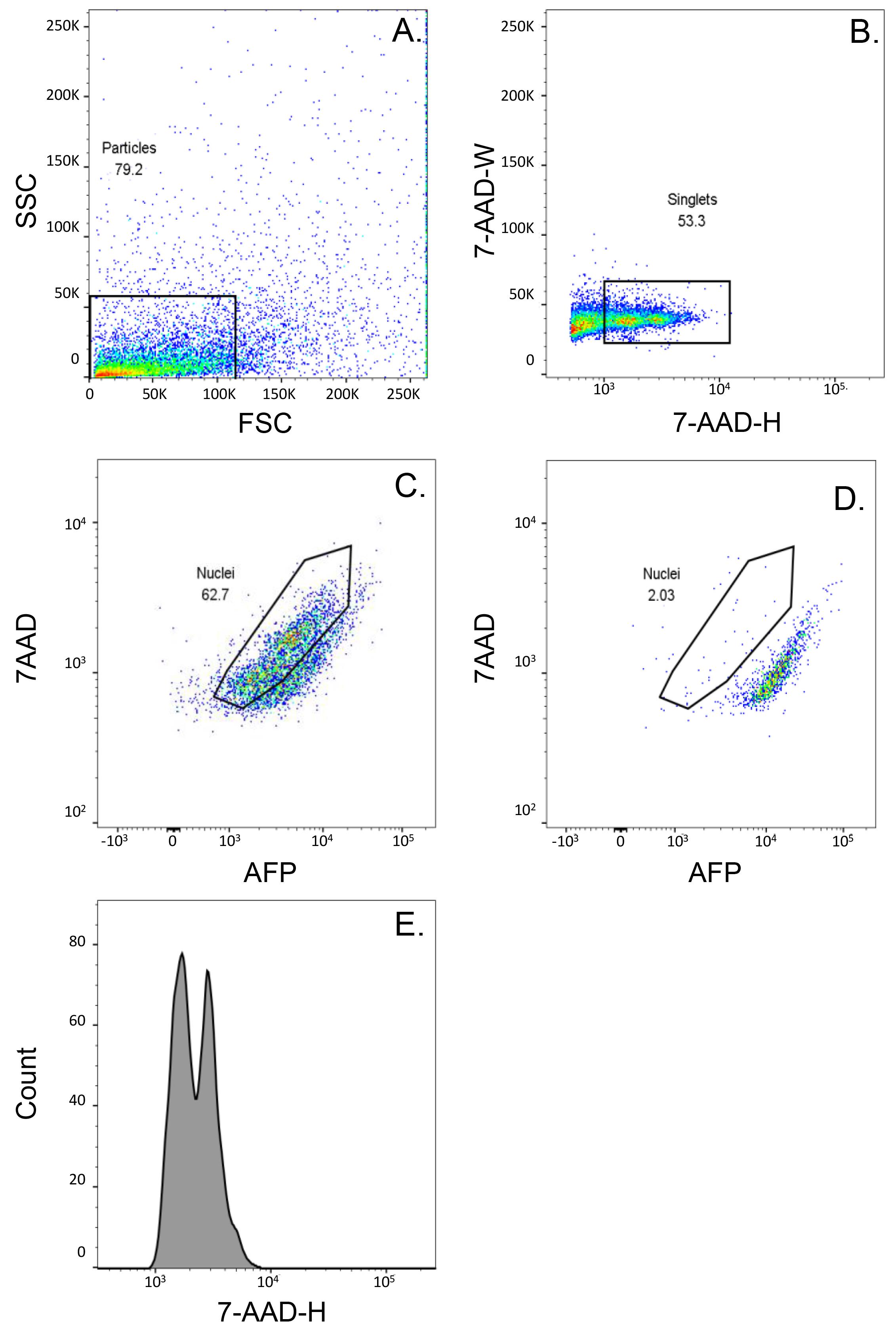
Figure 3. Gating strategy for the isolation of nuclei using the BD FACSAria II cell sorter. A. Forward Scatter (FSC) and Side Scatter (SSC) were utilized to visualize the primary particle cluster. B. Singlets were selected using 7-AAD Area (W) vs. Height (H). C and D. Plots showing 7-AAD and the adjacent fluorescent parameter (AFP) to better discriminate the stained nuclei (C) vs. the autofluorescing organelles (D). E. Histogram displaying the DNA content of the sorted nuclei. The number in each graph (A to D) represents the percentage of selected particles/nuclei (black box) for each step.Collect the sorted nuclei in a 15 ml conical tube containing 1 ml of pre-cooled 1% BSA-PBS.
To confirm the purity of the collected nuclei, sort an aliquot of the sample after SYTOX Green staining (final concentration of 5 nM). Sytox Green fluorescence was excited with a 488 nm laser and collected with a 505 LP and a 530/30 BP filter set.
Centrifuge the 7-AAD-sorted nuclei at 500 × g for 10 min at 4°C.
Discard the supernatant and gently re-suspend the nuclei pellet in 150 μl of 1% BSA-PBS.
Permeabilization of the nuclei
Add NIB medium (if using Step B. Option 1) or 1% BSA-PBS (if using Procedure B. Option 2) supplemented with 2% Triton X-100 to the nuclei suspension to reach a final Triton X-100 concentration of 0.5% (e.g., add 0.5 ml of NIB medium + 2% Triton X-100 for each 1.5 ml of NIB filtred-nuclei suspension or 50 μl of 1% BSA-PBS + 2% Triton X-100 per 150 μl of 1% BSA-PBS sorted-nuclei suspension).
Apply a gentle rocking for 5 min at 4°C (speed 2 of the Platform Rocker) for Arabidopsis thaliana (Farmer et al., 2021).
Note: Triton X-100 concentration and time of incubation may vary between plant sample and species. This step allows the penetration of the Tn5 transposase in the nuclei. However, overexposure to Triton X-100 might lead to the bursting of the nuclei in the microfluidic system. To test the integrity of the nuclei, conduct a microscopic observation on a subset of the isolated nuclei as described in Option 1, step 3. We recommend validating the condition of permeabilization of a sample before implementing sNucATAC-seq experiments.
Centrifuge the nuclei for 8 min at 500 × g and 4°C to pellet Arabidopsis nuclei and 5 min at 500 × g and 4°C to pellet Soybean nuclei (time and speed could vary depending on plant species and the size of the nuclei). Discard the supernatant.
Wash the pellet with 4 ml of 1% BSA-PBS. Re-suspend the pellet of nuclei very gently by swirling.
Centrifuge the nuclei suspension as described in Step C3. Discard the supernatant.
Resuspend the pellet of nuclei with 20 μl of 1× nuclei buffer provided by 10× Genomics. The volume can be adjusted based on the nuclei concentration and to fullfil the 10× Genomics requirements (i.e., the maximum volume to use per sample is limited to a maximum of 5 μl with a nuclei concentration between 155 and 7,700 nuclei/μl, to recover between 500 and 10,000 nuclei pending user’s choice).
Estimation of nuclei concentration
Transfer 9 μl of the nuclei suspension (from Step C6) to a new Eppendorf tube.
Add 1 µl of 0.4% Trypan blue and mix gently.
Load the entire volume of stained nuclei in the Cell Counting Chamber Slide.
Select Trypan blue positive nuclei with sizes between 2 and 30 µm (https://assets.fishersci.com/TFS-Assets/BID/manuals/MAN0014293_Countess_II_Automated_Cell_Counter_UG.pdf).
Note: Look for any clumps. Nuclei may form aggregates in the 10× Genomics 1× nuclei buffer. Clumping nuclei can be dissociated by a gentle swirling.
Estimate the nuclei density. Ideally, this density should be between 500 to 1,500 nuclei/µl.
Proceed to the sNucATAC-seq library construction following 10× Genomics protocol and using the Chromium Next GEM Single Cell ATAC Library & Gel Bead Kit (PN-1000175/6), the Chromium Next GEM Chip H Single Cell Kit, (PN-1000161/2), the Single Index Kit N, the Set A (PN-1000212), and the Chromium Controller (1000202/4) (https://assets.ctfassets.net/an68im79xiti/2NEwsG0Yu3RuxvtQiWXZo3/1e939394fa43a4f0bd88d79383833b16/CG000209_Chromium_NextGEM_SingleCell_ATAC_ReagentKits_v1.1_UserGuide_RevE.pdf).
Data analysis
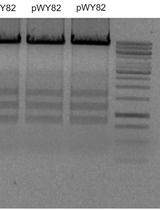
Upon synthesis of the sNucATAC-seq libraries, the profile of the cloned fragments must be evaluated using an Agilent Bioanalyzer or similar technology. Depending on the plant species, the profile of chromatin fragments may change, as shown in Figure 4. An example of processed sNucATAC-seq dataset upon using the method described in this manuscript has recently been published (Farmer et al., 2021).
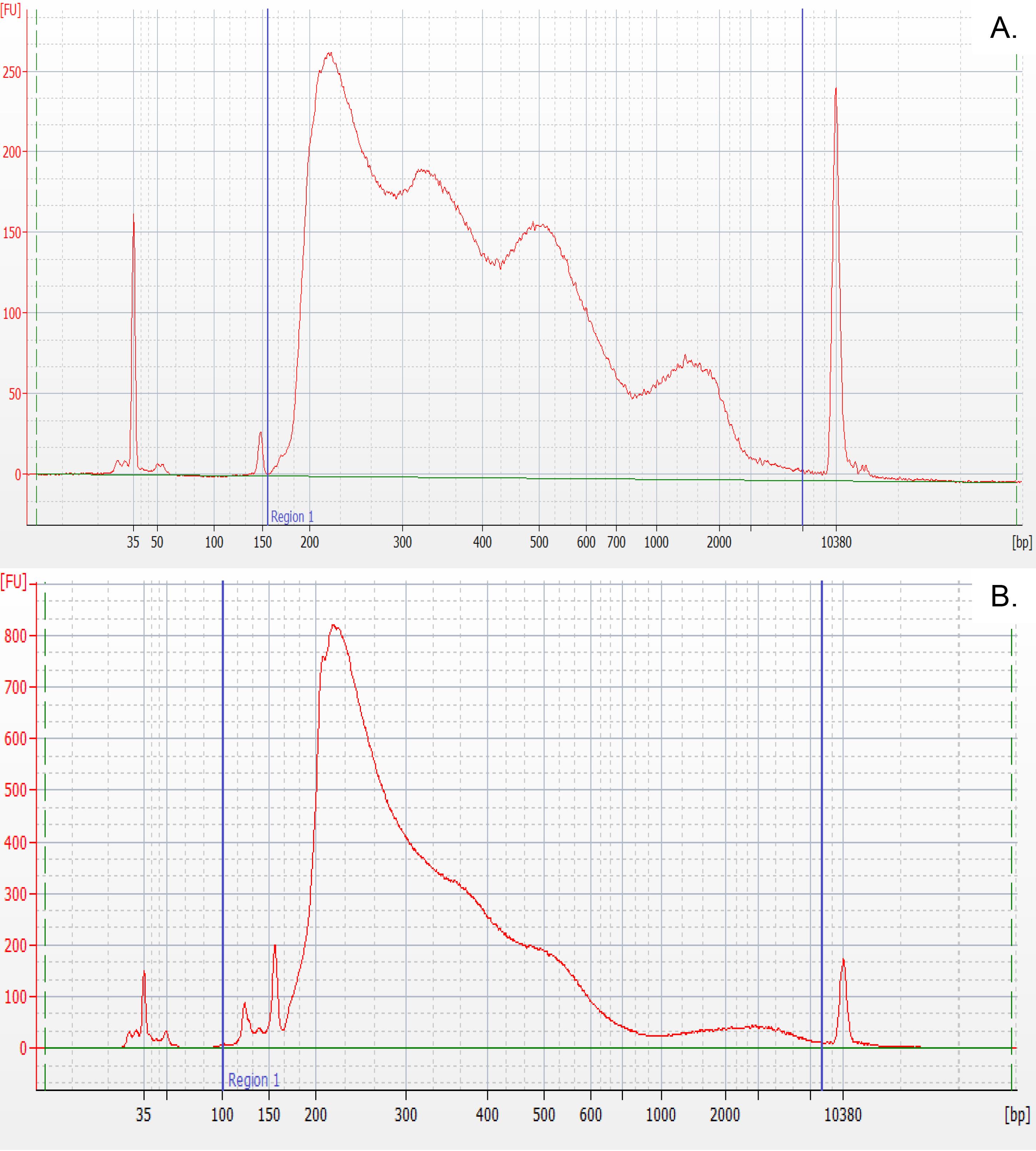
Figure 4. Representative profiles of A. Arabidopsis and B. Soybean cloned DNA fragments after tagmentation and scATAC-seq library construction. These profiles illustrate the difference in nucleosome periodicity between plant species. These profiles were generated using the High Sensitivity DNA kit and the Agilent 2100 Bioanalyzer.
Recipes
1% BSA-PBS
Add 200 µl of BSA (stock concentration: 50mg/ml) to 800 µl of PBS.
NIB medium supplemented with 2% Triton X-100
Add 20 µl of Triton X-100 to 980 µl of NIB medium 1×.
Pipette Triton X-100 slowly as it is very viscous.
1% BSA-PBS supplemented with 2% Triton X-100
Add 20 µl of Triton X-100 to 980 µl of 1% PBS-BSA.
Pipette Triton X-100 slowly as it is very viscous.
Acknowledgments
This work was supported by the U.S. National Science Foundation (IOS# 1339194 and 1854326), by the Center for Plant Science Innovation, and by the Department of Agronomy and Horticulture at the University of Nebraska-Lincoln. This protocol was adapted from a previous work (Thibivilliers et al., 2020).
Competing interests
The authors have no competing interests.
References
- Denyer, T., Ma, X., Klesen, S., Scacchi, E., Nieselt, K. and Timmermans, M. C. P. (2019). Spatiotemporal Developmental Trajectories in the Arabidopsis Root Revealed Using High-Throughput Single-Cell RNA Sequencing. Dev Cell 48(6): 840-852.e845.
- Dorrity, M. W., Alexandre, C. M., Hamm, M. O., Vigil, A. L. and Fields, S., Queitsch, C., and Cuperus, J. T. (2021). The regulatory landscape of Arabidopsis thaliana roots at single-cell resolution. Nature Commun 12(1): 3334.
- Farmer, A., Thibivilliers, S., Ryu, K. H., Schiefelbein, J. and Libault, M. (2021). Single-nucleus RNA and ATAC sequencing reveals the impact of chromatin accessibility on gene expression in Arabidopsis roots at the single-cell level. Mol Plant 14(3): 372-383.
- Huang, M. K., Zhang, L., Zhou, L. M., Yung, W. S., Li, M. W. and Lam, H. M. (2021). Genomic Features of Open Chromatin Regions (OCRs) in Wild Soybean and Their Effects on Gene Expressions. Genes (Basel) 12(5):640.
- Jean-Baptiste, K., McFaline-Figueroa, J. L., Alexandre, C. M., Dorrity, M. W., Saunders, L., Bubb, K. L., Trapnell, C., Fields, S., Queitsch, C. and Cuperus, J. T. (2019). Dynamics of Gene Expression in Single Root Cells of Arabidopsis thaliana. Plant Cell 31(5): 993-1011.
- Liu, Z., Zhou, Y., Guo, J., Li, J., Tian, Z., Zhu, Z., Wang, J., Wu, R., Zhang, B., Hu, Y., et al. (2020). Global Dynamic Molecular Profiling of Stomatal Lineage Cell Development by Single-Cell RNA Sequencing. Mol Plant 13(8): 1178-1193.
- Lu, Z., Hofmeister, B. T., Vollmers, C., DuBois, R. M. and Schmitz, R. J. (2017). Combining ATAC-seq with nuclei sorting for discovery of cis-regulatory regions in plant genomes. Nucleic Acids Res 45(6): e41.
- Marand, A. P., Chen, Z., Gallavotti, A. and Schmitz, R. J. (2021). A cis-regulatory atlas in maize at single-cell resolution. Cell 184(11): 3041-3055.e3021.
- Nelms, B. and Walbot, V. (2019). Defining the developmental program leading to meiosis in maize. Science 364(6435): 52-56.
- Pingault, L., Zogli, P., Brooks, J. and Libault, M. (2018). Enhancing Phenotyping and Molecular Analysis of Plant Root System Using Ultrasound Aeroponic Technology. Curr Protoc Plant Biol 3(4):e20078.
- Rich-Griffin, C., Stechemesser, A., Finch, J., Lucas, E., Ott, S. and Schafer, P. (2020). Single-Cell Transcriptomics: A High-Resolution Avenue for Plant Functional Genomics. Trends Plant Sci 25(2): 186-197.
- Ryu, K. H., Huang, L., Kang, H. M. and Schiefelbein, J. (2019). Single-Cell RNA Sequencing Resolves Molecular Relationships Among Individual Plant Cells. Plant Physiol 179(4):1444-1456.
- Shaw, R., Tian, X. and Xu, J. (2021). Single-Cell Transcriptome Analysis in Plants: Advances and Challenges. Mol Plant 14(1): 115-126.
- Shulse, C. N., Cole, B. J., Ciobanu, D., Lin, J., Yoshinaga, Y., Gouran, M., Turco, G. M., Zhu, Y., O'Malley, R. C., Brady, S. M. et al. (2019). High-Throughput Single-Cell Transcriptome Profiling of Plant Cell Types. Cell Rep 27(7): 2241-2247 e2244.
- Thibivilliers, S., Anderson, D. and Libault, M. (2020). Isolation of Plant Root Nuclei for Single Cell RNA Sequencing. Curr Protoc Plant Biol 5(4): e20120.
- Zhang, T. Q., Xu, Z. G., Shang, G. D. and Wang, J. W. (2019). A Single-Cell RNA Sequencing Profiles the Developmental Landscape of Arabidopsis Root. Mol Plant 12(5): 648-660.
Article Information
Copyright
© 2021 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Thibivilliers, S. B., Anderson, D. K. and Libault, M. Y. (2021). Isolation of Plant Nuclei Compatible with Microfluidic Single-nucleus ATAC-sequencing. Bio-protocol 11(23): e4240. DOI: 10.21769/BioProtoc.4240.
Category
Plant Science > Plant molecular biology > DNA > DNA structure
Molecular Biology > DNA > Chromatin accessibility
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









