- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Plasmodium cynomolgi Berok Growth Inhibition Assay by Thiol-reactive Probe Based Flow Cytometric Measurement
Published: Vol 11, Iss 17, Sep 5, 2021 DOI: 10.21769/BioProtoc.4147 Views: 3483
Reviewed by: Alexandros AlexandratosYee Ling LauAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Babesia duncani in Culture and in Mouse (ICIM) Model for the Advancement of Babesia Biology, Pathogenesis and Therapy
Vandana Kumari [...] Choukri Ben Mamoun
Nov 20, 2022 1957 Views
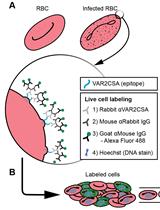
VAR2CSA Ectodomain Labeling in Plasmodium falciparum Infected Red Blood Cells and Analysis via Flow Cytometry
Olivia M.S. Carmo and Matthew W.A. Dixon
Aug 5, 2023 1332 Views
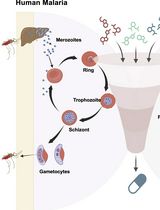
A Standardized Culture Medium for Comparative Drug Efficacy Evaluation Across Plasmodium and Babesia Species
Pratap Vydyam and Choukri Ben Mamoun
Mar 5, 2026 107 Views
Abstract
The relapsing malaria species, Plasmodium vivax, is the most widely distributed and difficult-to-treat cause of human malaria. The merozoites of P. vivax preferentially invade ephemeral human CD71+ reticulocytes (nascent reticulocytes), thereby limiting the development of a robust continuous culture in vitro. Fortunately, P. vivax’s sister species, P. cynomolgi Berok, can be cultured continuously, providing the ability to screen novel therapeutics drug and vaccine candidates in a reliable and high-throughput manner. Based on well-established growth inhibition activity (GIA) assays against P. falciparum and P. knowlesi, this protocol adopts the current flow cytometry assay methodology and investigates P. vivax inhibitory antibodies using the P. cynomolgi Berok invasion model based on the thiol-reactivity and DNA abundance of viable parasites in macaque erythrocytes. Established GIA assays screen antibodies at either a single concentration or high/low dose concentrations to provide quick insights for prioritizing potential antibodies capable of specifically interrupting parasite ligand and host receptor binding with minimal concentrations. Hence, this protocol expands on the existing GIA assay by using serially diluted antibodies and generating a dose-response curve to better quantify the inhibitory efficacy amongst selected vaccine candidates.
Keywords: Plasmodium vivaxBackground
The growth inhibition assay examines the re-invasion and growth of the Plasmodium spp. over two cycles of parasite replication (one cycle is approximately 24 h for P. knowlesi and 48 h for P. vivax, P. cynomolgi, and P. falciparum). After egress, asexual merozoites are released to continuously invade and multiply within host erythrocytes, increasing parasitaemia. During merozoite invasion, a cascade of parasitic proteins interacts with host cell receptors to manipulate host cell entry. In asymptomatic P. vivax patients, monoclonal antibodies recognise a wide range of parasitic ligands such as the duffy binding proteins (DBPs), reticulocyte binding proteins (RBPs), merozoite surface proteins (MSPs), and apical membrane antigen (AMA) (Han et al., 2019; Rawlinson et al., 2019). These antibodies have been shown to be strain-transcendent, confer protective immunity, and reduce merozoite invasion to reticulocytes in vitro short-term culture (Russell et al., 2011). However, the lack of a successfully multiplying continuous culture system of P. vivax reduces the detection signal in the growth inhibition assay (GIA) (Gunalan et al., 2020). To overcome this limitation, past P. vivax studies utilized surrogate species such as P. knowlesi, P. cynomolgi, and P. falciparum to screen inhibitory antibodies (Mitran et al., 2019; Collins et al., 1999; Muh et al., 2020).
The conventional GIA quantifies parasite viability and density using the SYBR green dye and lactate dehydrogenase (LDH) enzyme via flow cytometry and spectrophotometry, respectively (Mohring et al., 2020). Our approach explores thiol reactivity quantification in P. cynomolgi, which reliably assesses P. falciparum parasitaemia in antibody-dependent cellular inhibition assays (Jogdand et al., 2012). The quantification is based on the accumulation and fluorescent covalent biconjugates formed by the thiol-reactive probe carbocyanin, which acts as an indicator for the parasite’s viability in the sarcoplasmic reticulum and mitochondria of eukaryotes (Habicht and Brune 1980; Amaratunga et al., 2014). As the Mitotracker product is available in a wide range of fluorescent dyes and probes, this method also presents an opportunity to investigate other fluorescent channels in flow cytometry. Furthermore, our approach adapts the mean inhibitory concentration approach commonly used in drug assays, allowing researchers to select amongst highly effective inhibitory antibodies with low inhibitory concentrations.
Materials and Reagents
1.5 ml microcentrifuge tube (Axygen®, catalog number: MCT-150-C)
15 ml centrifuge tube (Corning, Costar, catalog number: CLS430052)
50 ml centrifuge tube (Corning, Costar, catalog number: CLS430291)
24-well flat-bottom plates (Corning, Costar, catalog number: CLS3738)
96-well round-bottom plates (Corning, Costar, catalog number: CLS3367)
96-well flat-bottom plates (Corning, Costar, catalog number: CLS3370)
Non-vented T75 flasks (Corning, Costar, catalog number: CLS3814)
3 ml round bottom polystyrene tubes (Becton Dickinson, catalog number: 156758)
Sterile V-channel reservoirs (VWR, catalog number: VWRI613-1174)
Sterile 14 mm Petri dish (VWR, catalog number: VWRI390-1373)
Non-woven filters (Antoshin)
Aluminium foil
Lithium heparin vacuteiners (Becton Dickinson, catalog number: 367880)
Clot activator vacuteiners (Becton Dickinson, catalog number: 367820)
P. cynomolgi Berok erythrocyte culture (Bruce Russell, University of Otago) (Chua et al., 2019)
Macaca fascicularis erythrocytes (Monash Animal Research Platform, Monash University)
M. fascicularis serum (Monash Animal Research Platform, Monash University)
Pre-immune and P. vivax antibodies (Yang Cheng, Jiangnan University) (Shen et al., 2021)
RPMI 1640 medium supplemented with GlutaMAX (Gibco, catalog number: 61870-036)
HEPES (Sigma-Aldrich, catalog number: H4034)
D-glucose (Sigma-Aldrich, catalog number: G7021)
Hypoxanthine (Calbiochem, catalog number: 4010BC)
PBS tablets (Gibco, catalog number: 18912014)
Giemsa (VWR, catalog number: 352603R)
Methanol (Fisher Scientific, catalog number: A412-4)
Hoechst (Sigma-Aldrich, catalog number: 63493)
Mitotracker Deep Red (Thermo Fisher Scientific, FM Molecular Probes, catalog number: 22426)
Complete culture media (see Recipes)
Equipment
Class II Microbiological Safety Cabinet
Microplate orbital shaker
Multichannel pipette (12-Channel pipette, 10-100 μl) (Thermo ScientificTM, catalog number: 4661070N)
Benchtop plate centrifuge (Eppendorf, model: 5804R)
Light microscope with 100× oil immersion lens (NIKON, model: YS100)
Canto II flow cytometer (Becton Dickinson, Canto II)
Tri-mix gas with 5% O2, 5% CO2, and the rest with N2
Hypoxia incubator chamber (Stemcell, catalog number: 27310)
37°C incubator
4°C fridge
-20°C freezer
Software
FACSDiva 6.1.3 software (Becton Dickinson)
Flowjo, LLC (Becton Dickinson, Tree Star Inc, https://www.flowjo.com/contact)
Microsoft Excel (Microsoft, https://www.microsoft.com/en-us/microsoft-365/excel)
Prism 8 (GraphPad, https://www.graphpad.com/scientific-software/prism/)
Procedure
Parasite culture
Thaw a frozen vial containing 1 ml of P. cynomolgi Berok parasitized erythrocytes using a series of sodium chloride solutions in a dropwise manner, according to Moll et al. (2008).
Resuspend the pelleted parasitized erythrocytes in complete culture media in a 24-well plate and incubate the parasite culture at 5% hematocrit in a Hypoxia incubator chamber filled with tri-mix gas in a 37°C incubator.
After overnight incubation, check for successful parasite maturation before expanding the parasitized erythrocytes in a non-vented flask in a 25-cm2 cell culture flask at 5% hematocrit using M. fascicularis erythrocytes, according to Chua et al. (2019).
If parasite culture shows a mix of early trophozoites and schizont stage (1:1) parasitized erythrocytes 2 days before the experiment day, perform a “quick and dirty” culture synchronisation.
“Quick and dirty” culture synchronisation
About 30 h before the experiment, pellet the parasite culture (at least 2 ml culture at 5% hematocrit) at 1,114 × g for 2 min at room temperature.
Set aside the supernatant (complete culture media) in a separate 50 ml Falcon tube.
Carefully aspirate and remove the brownish-black coloured top layer of pelleted erythrocytes (schizonts that have not egressed) manually with a pipette.
Using a fresh pipette tip, gently resuspend the pelleted red blood cells with the supernatant set aside previously, and transfer the mixture into a non-vented T75 flask. Gas with tri-mix gas for 5 s, and incubate flask at 37°C overnight.
Check that the overnight parasitized erythrocytes have matured to the schizont stage (at least 70%) the following afternoon.
Antibody preparation for GIA
Thaw frozen vials of antibodies on ice, and prepare fresh after confirmation of schizont maturation in parasitized erythrocyte culture.
In the antibody dilution plate, fill all outer wells of the 96-well round-bottomed plate with 200 μl of sterile water.
Note: If a flat-well bottomed plate is used, place the plate on an angle to ensure consistent pipetting at the bottom of each well.
Label the lid of the plate to indicate rows with the antibody name and columns from “1” to “10”.
Seed wells labelled “2” to “10” with 20 μl of complete media (see Figure 1A for plate layout).
Dilute the corresponding antibodies to their respective wells labelled with “1” with a final volume of 30 μl at 1 mg/ml with culture media.
Perform a threefold serial dilution by transferring 10 μl from well “1” to well “2”. Mix by pipetting.
Repeat the serial dilution of antibodies up to well “10”.
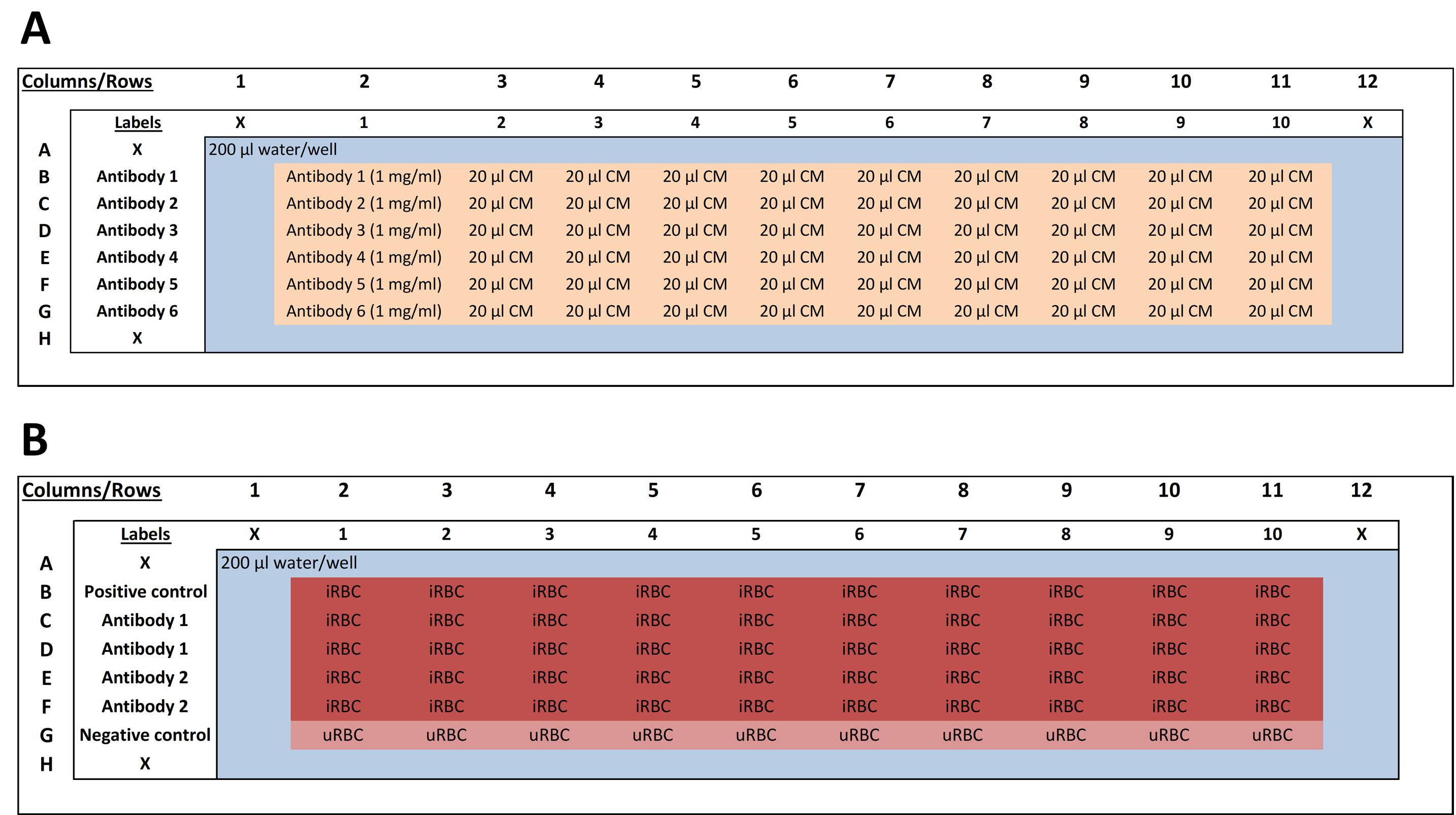
Figure 1. Antibody and experimental plates layout. The blue coloured wells show wells filled with 200 μl sterile water. Blanks are indicated as ‘X’. A. An illustration of the antibody plate layout with labels indicating antibodies from Rows A to H. Columns 1 to 12 indicate the decreasing antibody concentration. Orange coloured wells indicate 20 μl complete media (CM) as antibody diluent. B. An illustration of the experimental plate layout with labels indicating antibodies from Rows A to H and concentrations from 1 to 10. A positive control is included in Row B, with a negative control in Row G. Dark red coloured wells indicate wells containing infected red blood cells (iRBC), and the light red coloured wells indicate uninfected red blood cells (uRBC).
P. cynomolgi Berok GIA – Set up
Prior to set up, ensure that the majority of parasitized erythrocytes are in the mid-late schizont stage (at least 70% of parasitized erythrocytes in culture, with each schizont containing at least 8 merozoites).
To minimise media evaporation, fill all outer wells of the 96-well round-bottomed or flat-bottomed plate with 200 μl of sterile water, and place a covered 14 mm Petri dish containing sterile water in the hypoxia incubator chamber.
Label the lid of the plate to indicate in duplicate the antibody name and their corresponding test dilution with wells “1” to “10”. Include a row each of positive and negative control wells (see Figure 1B for plate layout).
Seed negative control wells with 70 μl of uninfected red blood cells at 1% hematocrit.
Seed positive control wells with 70 μl of parasitized red blood cells at 1% hematocrit and 0.5% parasitaemia.
The final volume for each test well is 70 μl. Calculate the volume of pelleted erythrocytes for a final volume of 70 μl/well at 1% hematocrit and 0.5% parasitaemia. However, as antibodies will be diluted 10× to attain a final concentration of 100 μg/ml, resuspend pelleted erythrocytes with complete culture media volume at 63 μl/well instead.
Transfer the mixture into a sterile V-channel reservoir.
Mix the solution in the reservoir by gently pipetting up and down to ensure a homogenous solution before pipetting 63 μl of the parasite culture into each test well.
Add 7 μl of antibodies (from Step C6) to each well in duplicate using a multi-channel pipette.
Incubate the plates in the hypoxia incubator chamber flushed with tri-mix gas at 37°C.
To maintain a healthy parasitized erythrocyte culture at the second cycle of merozoite invasion in this 2 cycle assay, add 7 μl of complete culture media (with freshly supplemented serum) to each well at 48 h post incubation.
P. cynomolgi Berok GIA – Staining
At 96 h post incubation, gently resuspend the parasite culture before extracting 20 μl into a new 96-well round-bottomed plate for staining.
Pellet red blood cells by spinning the plate in a centrifuge at 400 × g for 5 min at room temperature with low deceleration.
Prepare a staining MasterMix for a final volume of 50 μl/well with 8 μM Hoechst 34580 and 150 nM Mitotracker Deep Red FM in 1× PBS.
After the spin is complete, remove as much supernatant as possible from each well.
Resuspend the pelleted red blood cells with 50 μl of staining MasterMix.
Incubate the plate on the microplate orbital shaker at 120 rpm for 20 min at room temperature. To prevent quenching of the dye, cover the shaker with a big sheet of aluminium foil (reusable).
Pellet red blood cells by spinning the plate in a centrifuge at 400 × g for 5 min at room temperature with low deceleration.
Remove as much supernatant as possible, and wash cells with 200 μl of PBS.
Repeat the centrifugation and washing steps twice.
Resuspend cells in 200 μl of PBS, and transfer mixture to a 3 ml round bottom polystyrene tube for analysis on the flow cytometer.
Analysis of Hoechst 34580+/Mitotracker Deep Red FM+ parasitized erythrocytes by flow cytometry
Measure parasitaemia using FACS.
Record 50,000 to 100,000 cells for each sample.
Export data to fsc file format.
Data analysis
Using Flowjo software, gate single cells (x axis: SSC-H, y axis: FSC-H), then select the corresponding detector channels based on the single-cell population and gate the double-positive Hoechst 34580+/Mitotracker Deep Red+ population.
Export gated population values to Excel software.
Subtract the baseline value (negative control) from test samples.
Copy values from Excel software to GraphPad Prism 8 software.
To compare the percentage of Hoechst 34580+/Mitotracker Deep Red+ viable parasites across different drug concentrations, drug concentrations are first log2 transformed and the data values are then normalized.
Use the four-parameter logistic model or log (inhibitor) vs. response – variable slope function for dose-response curve fitting.
The mean IC50 values for various antibodies are compared using one-way ANOVA with the Tukey post hoc test function.
Recipes
Complete culture media
RPMI-1640 medium, GlutaMAXTM supplement, HEPES with the following additions:
2.5 ml HEPES (1 M)
1.0 g/L D-(+)-Glucose
0.05 g/L Hypoxanthine
20% (vol/vol) M. fascicularis serum
Sterile filter and store at 4°C
Acknowledgments
This work was supported by the Research Grant from the Institute of Medical Sciences, Kangwon National University 2020 (J-H. H), and by the Marsden Fund 17-U00-241 from the B.R. laboratory (B. R.). The protocol is adapted from Shen et al. (2021).
Competing interests
The authors declare no competing financial interests.
References
- Amaratunga, C., Neal, A. T. and Fairhurst, R. M. (2014). Flow cytometry-based analysis of artemisinin-resistant Plasmodium falciparum in the ring-stage survival assay. Antimicrob Agents Chemother 58(8): 4938-4940.
- Chua, A. C. Y., Ong, J. J. Y., Malleret, B., Suwanarusk, R., Kosaisavee, V., Zeeman, A. M., Cooper, C. A., Tan, K. S. W., Zhang, R., Tan, B. H., Abas, S. N., Yip, A., Elliot, A., Joyner, C. J., Cho, J. S., Breyer, K., Baran, S., Lange, A., Maher, S. P., Nosten, F., Bodenreider, C., Yeung, B. K. S., Mazier, D., Galinski, M. R., Dereuddre-Bosquet, N., Le Grand, R., Kocken, C. H. M., Renia, L., Kyle, D. E., Diagana, T. T., Snounou, G., Russell, B. and Bifani, P. (2019). Robust continuous in vitro culture of the Plasmodium cynomolgi erythrocytic stages. Nat Commun 10(1): 3635.
- Collins, W. E., Warren, M. and Galland, G. G. (1999). Studies on infections with the Berok strain of Plasmodium cynomolgi in monkeys and mosquitoes. J Parasitol 85(2): 268-272.
- Gunalan, K., Rowley, E. H. and Miller, L. H. (2020). A Way Forward for Culturing Plasmodium vivax. Trends Parasitol 36(6): 512-519.
- Habicht, J. and Brune, K. (1980). Carbocyanine dyes stain the sarcoplasmic reticulum of beating heart cells. Exp Cell Res 125(2): 514-518.
- Han, J. H., Cheng, Y., Muh, F., Ahmed, M. A., Cho, J. S., Nyunt, M. H., Jeon, H. Y., Ha, K. S., Na, S., Park, W. S., Hong, S. H., Shin, H. J., Russell, B. and Han, E. T. (2019). Inhibition of parasite invasion by monoclonal antibody against epidermal growth factor-like domain of Plasmodium vivax merozoite surface protein 1 paralog. Sci Rep 9(1): 3906.
- Jogdand, P. S., Singh, S. K., Christiansen, M., Dziegiel, M. H., Singh, S. and Theisen, M. (2012). Flow cytometric readout based on Mitotracker Red CMXRos staining of live asexual blood stage malarial parasites reliably assesses antibody dependent cellular inhibition. Malar J 11: 235.
- Moll, K., L. M. I., Perlmann, H., Scherf, A. and Wahlgren, M. (2008). Methods In Malaria Research. Malaria Research and Reference Reagent Resource Center (MR4) American Type Culture Collection (ATCC). 5th edition.
- Mitran, C. J., Mena, A., Gnidehou, S., Banman, S., Arango, E., Lima, B. A. S., Lugo, H., Ganesan, A., Salanti, A., Mbonye, A. K., Ntumngia, F., Barakat, K., Adams, J. H., Kano, F. S., Carvalho, L. H., Maestre, A. E., Good, M. F. and Yanow, S. K. (2019). Antibodies to Cryptic Epitopes in Distant Homologues Underpin a Mechanism of Heterologous Immunity between Plasmodium vivax PvDBP and Plasmodium falciparum VAR2CSA. mBio 10(5): ):e02343-19.
- Mohring, F., Rawlinson, T. A., Draper, S. J. and Moon, R. W. (2020). Multiplication and Growth Inhibition Activity Assays for the Zoonotic Malaria Parasite, Plasmodium knowlesi. Bio-protocol 10(17): e3743.
- Muh, F., Kim, N., Nyunt, M. H., Firdaus, E. R., Han, J. H., Hoque, M. R., Lee, S. K., Park, J. H., Moon, R. W., Lau, Y. L., Kaneko, O. and Han, E. T. (2020). Cross-species reactivity of antibodies against Plasmodium vivax blood-stage antigens to Plasmodium knowlesi. PLoS Negl Trop Dis 14(6): e0008323.
- Rawlinson, T. A., Barber, N. M., Mohring, F., Cho, J. S., Kosaisavee, V., Gerard, S. F., Alanine, D. G. W., Labbe, G. M., Elias, S. C., Silk, S. E., Quinkert, D., Jin, J., Marshall, J. M., Payne, R. O., Minassian, A. M., Russell, B., Renia, L., Nosten, F. H., Moon, R. W., Higgins, M. K. and Draper, S. J. (2019). Structural basis for inhibition of Plasmodium vivax invasion by a broadly neutralizing vaccine-induced human antibody. Nat Microbiol 4(9): 1497-1507.
- Russell, B., Suwanarusk, R., Borlon, C., Costa, F. T., Chu, C. S., Rijken, M. J., Sriprawat, K., Warter, L., Koh, E. G., Malleret, B., Colin, Y., Bertrand, O., Adams, J. H., D'Alessandro, U., Snounou, G., Nosten, F. and Renia, L. (2011). A reliable ex vivo invasion assay of human reticulocytes by Plasmodium vivax. Blood 118(13): e74-81.
- Shen, F. H., Ong, J. J. Y., Sun, Y. F., Lei, Y., Chu, R. L., Kassegne, K., Fu, H. T., Jin, C., Han, E. T., Russell, B., Han, J. H. and Cheng, Y. (2021). A chimeric Plasmodium vivax merozoite surface protein antibody recognizes and blocks erythrocytic P. cynomolgi berok merozoites in Vitro. Infect Immun 89(2): e00645-20.
Article Information
Copyright
© 2021 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Ong, J. J. Y., Russell, B. and Han, J. (2021). Plasmodium cynomolgi Berok Growth Inhibition Assay by Thiol-reactive Probe Based Flow Cytometric Measurement. Bio-protocol 11(17): e4147. DOI: 10.21769/BioProtoc.4147.
Category
Immunology > Antibody analysis > Antibody function
Microbiology > Microbial cell biology > Cell isolation and culture
Cell Biology > Cell-based analysis > Flow cytometry
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









