- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Agrobacterium-mediated Transformation of Japonica Rice Using Mature Embryos and Regenerated Transgenic Plants
Published: Vol 11, Iss 18, Sep 20, 2021 DOI: 10.21769/BioProtoc.4143 Views: 6216
Reviewed by: Bedabrata SahaAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
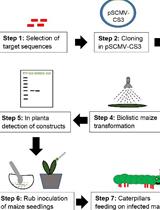
Inoculation of Maize with Sugarcane Mosaic Virus Constructs and Application for RNA Interference in Fall Armyworms
Iram Gull and Georg Jander
Jul 20, 2023 2203 Views
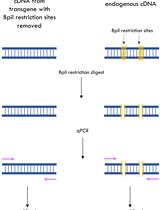
A qPCR Method to Distinguish between Expression of Transgenic and Endogenous Copies of Genes
William Bezodis [...] Hugh Dickinson
Aug 5, 2023 1734 Views
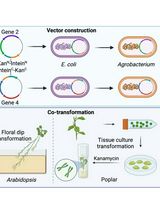
A Novel Gene Stacking Method in Plant Transformation Utilizing Split Selectable Markers
Guoliang Yuan [...] Xiaohan Yang
Feb 20, 2025 1968 Views
Abstract
Identification of novel genes and their functions in rice is a critical step to improve economic traits. Agrobacterium tumefaciens-mediated transformation is a proven method in many laboratories and widely adopted for genetic engineering in rice. However, the efficiency of gene transfer by Agrobacterium in rice is low, particularly among japonica and indica varieties. In this protocol, we elucidate a rapid and highly efficient protocol to transform and regenerate transgenic rice plants through important key features of Agrobacterium transformation and standard regeneration media, especially enhancing culture conditions, timing, and growth hormones. With this protocol, transformed plantlets from the embryogenetic callus of the japonica cultivar ‘Taichung 65’ may be obtained within 90 days. This protocol may be used with other japonica rice varieties.
Keywords: ElectroporationBackground
Genetic transformation and expression of recombinant proteins in plant cells are major factors for plant genetic engineering. This is also a powerful tool for discovering novel genes and exploring genetically controlled traits. Agrobacterium tumefaciens-mediated transformation has been known for its unique ability to transfer a DNA segment from a specialized plasmid into a host plant cell (Gelvin, 2010). This feature allows efficient insertion of stable, unrearranged, single-copy DNA into plant genomes, which may lead to more stable expression than multiple gene copies or scrambled inserts (Iglesias et al., 1997). Since the initial reports in the early 1980s using Agrobacterium to generate transgenic plants, efforts were made to improve the tool in plant transformation. In most instances, major improvements involved alterations in plant tissue culture transformation and regeneration conditions rather than manipulation of host or bacterial genes. The first effective method for Agrobacterium-mediated transformation of japonica rice is now a common protocol in many laboratories (Chan et al., 1993; Hiei et al., 1994; Park et al., 1996). Many functional analyses, for example, PCR, GUS assay, and southern blot analysis, have confirmed the integration of foreign genes into transgenic rice plants obtained by Agrobacterium-mediated transformation (Dai et al., 2001); moreover, Mendelian inheritance of the transgenes was also reported (Pawlowski and Somers, 1996; Hiei et al., 1997).
The transfer of T-DNA and its integration into the rice genome is influenced by numerous factors, such as genotype, explants, Agrobacterium strains, plasmid vectors, addition of vir-gene inducing synthetic phenolic compounds, selection marker, and various conditions of tissue culture and regeneration. In tissue culture systems, selection of actively growing regenerable calli is a principal factor for efficient plant transformation. Likewise, optimization of culture conditions for co-cultivation of rice calli with Agrobacterium (Ozawa and Takaiwa, 2010) and suppression of Agrobacterium overgrowth may be applied to improve the transformation efficiency. Scientists have attempted to improve the plasmid constructions and rice transformation by Agrobacterium-mediated transformation to investigate the function of seed storage protein gene factors as well as cloning genomic candidate regions of more than 10 kb, which may have low restriction enzymes sites required for cloning. In particular, previous limitations of efficient promoter expression were overcome (Gupta et al., 2001; Furtado et al., 2008), especially for endosperm-specific expression (Zhou et al., 2013).
Several transformation methods have been established by many laboratories using mature japonica and indica seeds. Taichung 65 is a japonica rice variety, selected from a cross between the two Japanese varieties, Shinriki and Kameji, in 1923 (Iso, 1957). Taichung 65 is considered an important cultivar for studying rice genetics and breeding. Several point mutations were induced to the genetic stock of Taichung 65 using the methylnitrosourea (MNU) mutagen and have been maintained at the institute of plant genetic resources of Kyushu university. Taichung 65 has been extensively used as a model cultivar for rice biology, breeding research, and genomic studies initiated in many laboratories. However, transformation efficiency is still low in most indica and many japonica varieties due to the difficult regeneration; consequently, the procedure to obtain transgenic plants takes an average of 5 months (Nishimura et al., 2006). Thus, additional improvements in the transformation potentiality are still possible.
Here, we describe a protocol for Agrobacterium-mediated transformation in rice using mature embryos. This protocol is based on the method described by Toki (Toki, 1997), with several modifications; in particular, meropenem is used for plant regeneration (Ogawa and Mii, 2007). Meropenem exhibits the highest antibacterial activity and also achieves a high shoot formation rate in rice and tomato (Ogawa and Mii, 2007). Using this protocol, we succeeded to produce hundreds of independent transgenic lines and transformed several novel genes over the past decade, including Endosperm Storage Protein (ESP1), encoding a eukaryotic chain release factor1 (eRF1) (Elakhdar et al., 2019); Esp2, encoding the protein disulfate isomerase 1-1 (PDI 1-1) (Satoh-Cruz et al., 2010); Glup3, encoding a vacuole processing enzyme (VPE) (Kumamaru et al., 2010); Glup4, encoding the small GTPase Rab5a (Fukuda et al., 2011); and Glup6, encoding the guanine nucleotide exchange factor (GEF) (Fukuda et al., 2013).
In summary, this protocol is a step-by-step approach to obtain stably transformed plants from mature rice embryos by optimizing several stages of Agrobacterium transformation and standard regeneration medium components, including culture conditions, timing, and growth hormones. In this method, we use a 13.8 kb construct carrying the hygromycin phosphotransferase gene (hpt), a firefly luciferase reporter gene (LUC), spectinomycin (Sp), an attR1 site, the chloramphenicol resistance gene (CmR), the ccdB gene, the attR2 site, and the ZmUbi1 promoter (Figure 1 and Figure 2).

Figure 1. Flow chart for Agrobacterium-mediated transformation and regeneration of rice calli using mature embryogenic seed (cv; Taichung 65). (a) Seed embryos. (b) Calli pieces with granular structure, yellowish-white. (c) Growth of transformed Agrobacterium harboring the pSMAH638OX/Ubilp binary vector. (d) Co-cultivation of infected calli. (e) De-colonization of Agrobacterium tumefaciens. (f) Shoot regeneration. (g) Rooting in regenerated shoots. (h) Hardening of rooted plants. (i) Transplanted hardened plants.
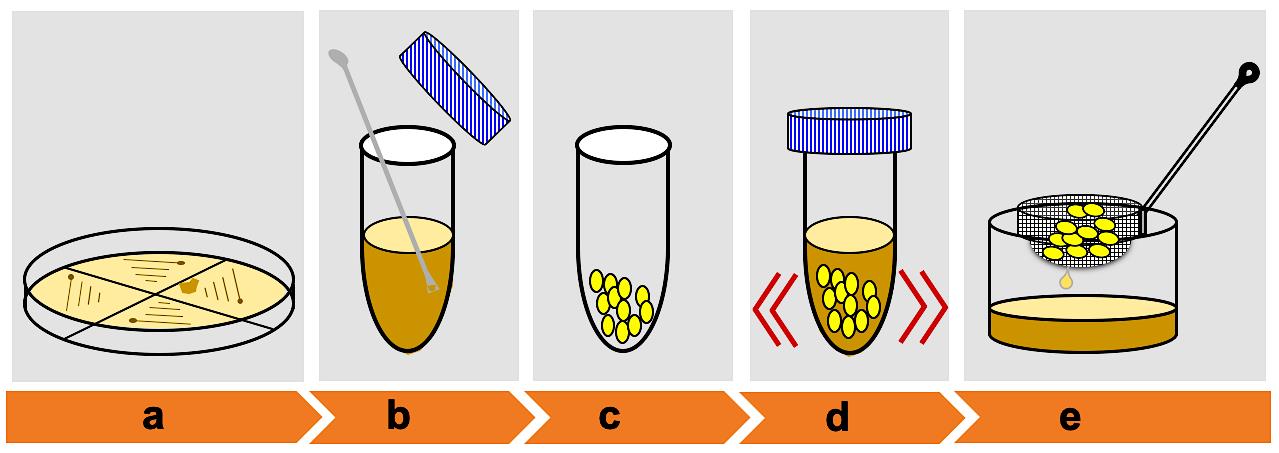
Figure 2. Schematic showing embryo calli infection by Agrobacterium. (a) Growth of Agrobacterium-mediated harboring pSMAH638OX/Ubilp vector on LB medium. (b) Mix a small portion of the positive colonies. (c) Collect healthy calli for infection. (d) Calli infection. (e) Remove excess Agrobacterium suspension.
Materials and Reagents
Plasticware and Glass
Petri dish, 90 × 20 mm
Petri dish, 90 × 15 mm
24-well multi-well plates
Sterile plastic falcon tubes, 50 ml
Filter paper, 70 mm
Parafilm tape (Parafilm Pechiney PM996; 125' L X 4"; www.parafilm.com)
Sterile syringe filters 0.22 μm
Sterile syringe, 30 ml
Stainless-steel sieve
Microspatula
Plant culture pots for rooting (6.5 × 6.5 cm)
Plant soil pots (23 × 23 × 19 cm)
Biological materials
Mature dry seeds of japonica cv. Taichung 65
The target cultivar of the current transformation method.
Binary vector containing gene of interest
In the following protocol, the binary vector pSMAH638OX/Ubilp (13.8 Kb) construct was derived by the Gateway recombination cloning technology (InvitrogenTM, Thermo Fisher Scientific, Inc.), harboring firefly luciferase (LUC) reporter gene, the hygromycin phosphotransferase (hpt) selectable gene marker, spectinomycin (Sp), an attR1 site, the chloramphenicol resistance gene (CmR), the ccdB gene, and the attR2 site. We used the ZmUbi1 promoter for LUC expression (Hakata et al., 2010; Elakhdar et al., 2019)
Agrobacterium tumefaciens strains EHA101 or EH105 (Hood et al., 1993).
Reagents and chemicals
Ethanol CH3CH2OH (Sigma-Aldrich, catalog number: N0640), 70% (v/v) in distilled water, store at room temperature (RT)
Sodium hypochlorite 10% NaClO (Sigma-Aldrich, catalog number: 105614), store at RT
Sodium chloride, NaCl (Sigma-Aldrich, catalog number: 7647-14-5), store at RT
Sodium hydroxide, NaOH (Sigma-Aldrich, catalog number: 1310-73-2), store at RT
Tween® 20 EMD, (CALBIOCHEM, catalog number: 655205), store at RT
Sucrose C12H22O11 (Wako, catalog number: 196-00015), store at RT
D-Glucose C6H12O6 (Sigma-Aldrich, catalog number: 07-0680-5), store at RT
D-Sorbitol C16H14O6 (Sigma-Aldrich, catalog number: 28-4770-5), store at RT
Casamino acid (Sigma-Alorich, catalog number: 65072-00-6), store at RT in dry condition
L-Proline (PEPTIOE, catalog number: 2718), store at RT
CHU (N6) Basal salt mixture (Sigma-Aldrich, catalog number: C1416-10L), store at 2-8°C
Murashige & Skoog Basal Medium (Sigma-Aldrich, catalog number: M552450L), store at 2-8°C
CHU N6-Vitamin solution X1000 (Phyto, catalog number: PHT:C149), store at 2-6°C
LB Broth, Miller (Luria-Bertani; BD Difco, catalog number: 244620), store at RT
Bacto Peptone (BD Difco, catalog number: 211677), store at RT
Bacteriological agar (Bacto Agar; BD Difco, catalog number: 214010), store at RT
2,4-Dichlorophenoxyacetic acid (Wako, catalog number: 040-18532), store at RT
1-Naphthaleneacetic acid NAA C10H7CH2COOH (Wako, catalog number: 86-87-3), store at RT
Kinetin C10H9N5O (Wako, catalog number: 110-00331), store at 2-10°C
Gelrite (0.4%) (Wako, catalog number: 067-04035), store at RT
Carbenicillin sodium salt (Sigma-Aldrich, catalog number: C1389-5G), store at 2-8°C
Hygromycin B (Wako, catalog number: 085-06153), store at -20°C
Acetosyringone (Sigma-Aldrich, catalog number: D134406-5G), store at -20°C
Meropenem (Dainippon Sumitomo Pharma, Osaka, Japan), store at -20°C
Dimethyl sulfoxide DMSO (CH3)2SO (Wako, catalog number: 046-21981), store at RT
KOD-FX (TOYOBO, catalog number: KFX-101), store at -20°C
Sterilizing solution (see Recipes)
Solution A
Solution B
Antibiotics (see Recipes)
Carbenicillin (500 mg/ml)
Hygromycin (50 mg/ml)
Kanamycin (50 mg/ml)
Acetosyringone (100 mg/ml)
Acetosyringone (10 mg/ml)
Meropenem (12.5 mg/ml)
2,4-D (0.2 mg/ml)
NAA (0.2 mg/ml)
Kinetin (1 mg/ml)
Cultivation medium (see Recipes; Table 1)
LB sold medium
LB liquid medium
YEP medium
Notes:
All liquids and equipment containing Agrobacterium must be appropriately sterilized.
Essential elements that may influence the effectiveness of transformation are reagents and chemicals. Therefore, we strongly suggest following the methodology, particularly when starting a new protocol. The usage and storage conditions of chemicals are different; therefore, please make sure to follow the recommendations of each company, with particular attention to hazard information on the substances.
Equipment
-86°C Ultra-Low temperature freezer (SANYO, catalog number: MDF-U538)
-30°C Ultra-Low temperature freezer (SANYO, catalog number: MDF-792AT)
MicroPluserTM (Bio-Rad, Hercules)
Spectrophotometer (Thermo ScientificTM, model: NanoDrop 2000)
Incubator/ shaker 28°C for Agrobacterium
Plant growth chamber (SANYO)
Shaker for seed sterilization
Laminar flow hood (SANYO)
Autoclave
pH meter
Spinbar® magnetic stir bar (Sigma-Aldrich, catalog number: Z127116)
Polycarbonate vacuum desiccator (SANSYO, catalog number: SPD-WVGT240)
Thermal cycler (Bio-Rad, model: Tetrad 2 Thermal Cycler (4 × 96-well)
Locking system greenhouse
Rice seedling raising soil (JA Kumiai King Soil; Agr. Japan Co., Ltd.)
Procedure
Agrobacterium culture and transformation
Transformation of competent Agrobacterium cells (EHA101 or EHA105) with binary vector pSMAH638OX/Ubilp (13.8 Kb) constructs (Figure 3).
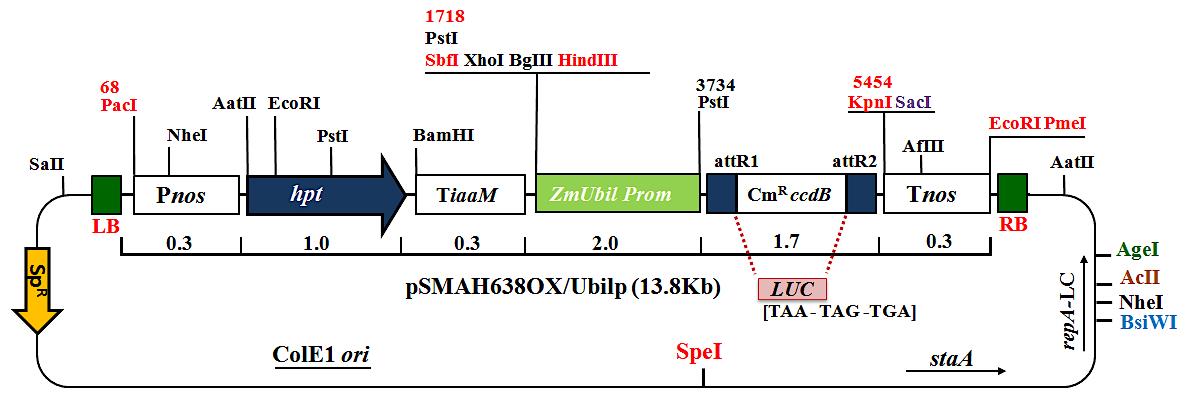
Figure 3. Binary vector pSMAH638OX/Ubilp (13.8 Kb) construct carrying the hygromycin phosphotransferase gene (hpt), firefly luciferase reporter gene (luc), spectinomycin (Sp), an attR1 site, the chloramphenicol resistance gene (CmR), the ccdB gene, the attR2 site, and the ZmUbi1 promoter.Thaw 50 μl Agrobacterium competent cells (EHA101 or EH105) on ice (Figure 4A).
In 1.5 ml tube, mix 2 µl plasmid DNA (100 picogram/µl) with competent cells.
Incubate the cell mixture on ice for 10 min.
Perform transformation using the MicroPluserTM (Bio-Rad) electroporation method with the “Agr” mode (Figure 4B).
After 10 min, move the cell mixture to the cuvette tube and then the cuvette to chamber slide.
Press the Pluse button until a tone sounds indicating that the pulse has been given.
Immediately add 250 µl YEP medium to cells.
Transfer cells to 25 ml tube containing 3.75 ml YEP medium (Figure 4C).
Warp the tube with foil and incubate for 2 h in the dark at 28°C.
Centrifuge the mixture at 1,500-3,000 × g for 5 min at RT.
Discard around 3.5 ml of the supernatant.
Mix the remaining cells gently.
Plate 20-100 µl of cells on LB agar amended with 50 mg/L hygromycin and 50 mg/L kanamycin.
Make sure that cells are completely dry, then wrap the Petri dish with parafilm and incubate 28°C for 2-3 days.
Pick four single colonies and verify the transformation by PCR (Figure 4D).
Streak out the positive colonies of Agrobacterium strain EHA101 that harbor the gene of interest in a 1 × 4 grid pattern on a single LB agar containing 50 mg/L hygromycin and 50 mg/L kanamycin (Figure 4E).
Incubate the culture for 2-3 days at 28°C.
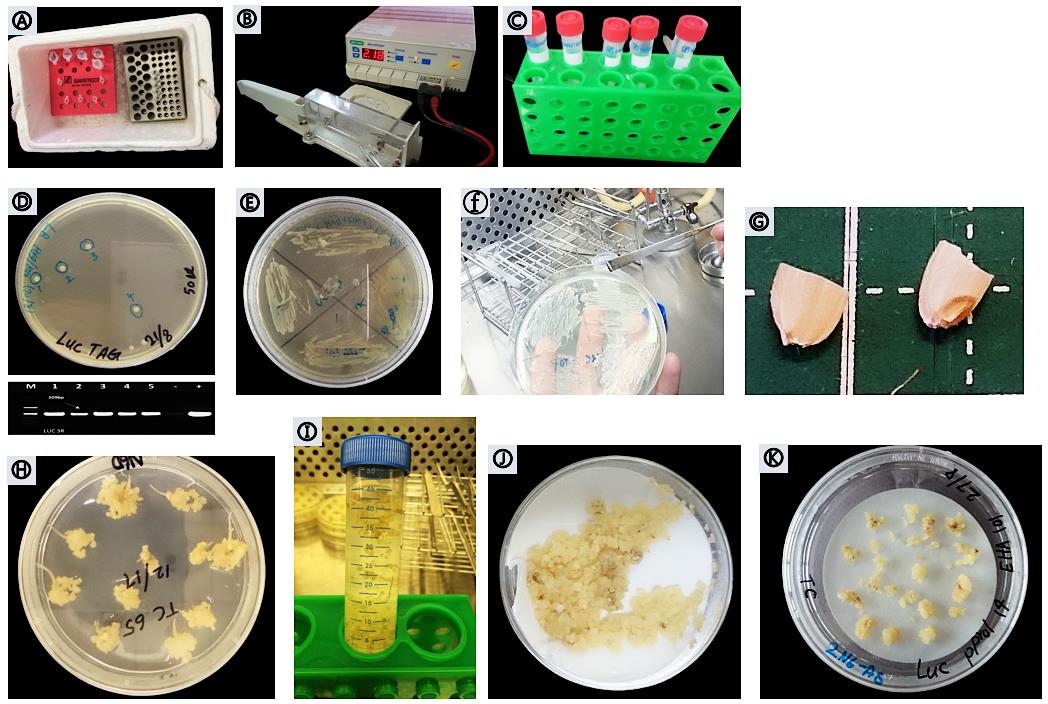
Figure 4. Procedure of Agrobacterium-mediated transformation. (A) Agrobacterium competent cells. (B) MicroPluserTM (Bio-Rad). (C) YEP medium. (D-F) Positive transformed Agrobacterium verified by PCR. (G) Mature embryos of Taichung 65. (H) Callus induction after 23 days on N6D medium. (I) Callus infection. (J) Drying calli on sterilized filter paper. (K) Co-cultivated calli on 2N6AS solid medium.
Preparing mature embryos
Seed surface disinfection
Dehusk healthy mature seeds, then hand-cut with a razor. Save the embryo seed halves and discard the endosperm seed halves (Figure 4G).
Disinfect the surface of embryogenic seed parts after soaking and stirring once in 70% ethanol for 10 s in a 50 ml tube (up to 150 seeds).
Rinse the embryo seeds halves in distilled water for 30 s.
Shake the tube vigorously (45 rpm) for 15-20 min in bleach Solution A and then rinse 3-5 times using distilled water.
Wash the seed parts with Solution B under vigorous shaking for 15-20 min, followed by five rinses in distilled water up to 30 s each time.
Place the sterilized embryo seed halves on sterile filter paper over Petri dish (see Note 3).
Callus induction (3-4 weeks)
Partially submerge 8-12 disinfected embryo seed halves per dish within N6D callus induction medium.
Seal the dishes with surgical tape, then wrap every five dishes with aluminum foil and incubate at 28°C in the dark for 3-4 weeks (see Note 4).
Preparation of calli, Agrobacterium suspension, and infection (48 to 72 h)
Mix a small portion of four colonies of Agrobacterium strain EH101 on a micro-spatula and resuspend in 40 ml 2N6D-AS liquid medium containing 16 μl acetosyringone (100 mg/ml stock solution) and mix gently (Agrobacterium colonies must be analyzed by PCR using specific primers to verify the integration of the construct with the bacterial gDNA) (Figure 4F).
Discard brown calli and seed coleoptile from the N6D medium (at the beginning of week 4). Collect the active embryogenic calli (yellowish-white, relatively dry, 1-3 mm in diameter (Nishimura et al., 2006) and globular (Hiei and Komari, 2008) into a 50-ml tube. At this stage, the calli can be used directly for Agrobacterium infection (Figure 4H).
Pour the bacterial suspension prepared in Step B3a into the 50-ml tube.
Soak the calli in the Agrobacterium suspension and gently mix for 90 s (Figure 4I).
Decant the bacterial suspension into a sterilized stainless-steel sieve (12 mesh) and hold in 500-ml sterile beaker.
Blot the sieve on sterilized filter paper placed in a 90 mm × 20 mm Petri dish to remove excess Agrobacterium suspension (Figure 4J).
Place a sterilized filter paper on solid 2N6D-AS medium. Saturate the filter paper by dripping 0.5 ml 2N6D-AS liquid medium and then co-cultivate an appropriate amount of calli (20-25 calli) on the filter paper (Figure 4K).
Seal plates with parafilm tape and wrap with aluminum foil and incubate at 25°C in the dark for 3 days (see Note 5).
Wash calli and select transformed cells (3-4 weeks)
Collect infected calli in a 50 ml sterile tube and wash with sterile water 3-5 times to remove Agrobacterium (until water is clear). Gently shake the tube and pour out the water.
Rinse calli with 40-ml sterile water containing 40 μl carbenicillin 500 mg/ml stock solution to kill Agrobacterium cells.
Pour the water suspension and blot the calli on sterile filter paper held over Petri dish. Dry the calli well (see Note 6).
Transfer calli onto N6D-HC selection medium; 15 to 20 calli can be placed on a single plate.
Seal the plate with surgical tape, and incubate the calli at 28°C in the dark [according to the Toki method; 33°C (light for 10 h)/30°C (dark for 14 h); Saika and Toki (2010)].
Check the culture regularly for contamination. In case of contamination, transfer uncontaminated calli to fresh medium immediately.
Subculture the yellowish-white calli to new fresh N2D-HC medium after 10 to 12 days, seal plates with surgical tape, and incubate at 28°C in the dark. Resistant calli (yellowish-white) can be observed after 20-25 days (Figure 5A).
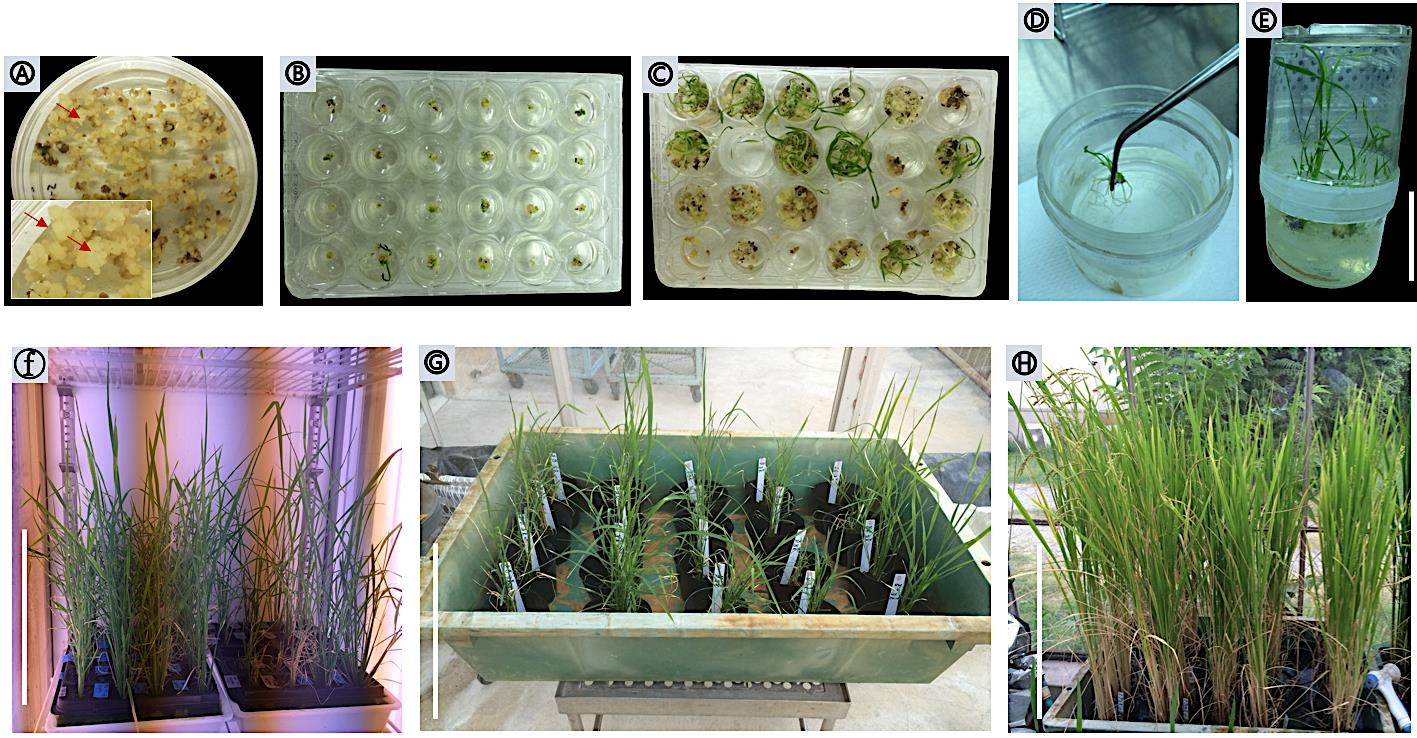

Figure 5. Calli selection and regeneration. (A) Hygromycin-resistant calli on N6D-HC selection medium. Arrows indicate resistant calli after 25 days from the infection. (B) Proliferated calli with greening spots. (C) Subculture green pots on the same fresh medium. (D) Rooting. (E) Regenerated plantlets on MS HF medium. (F) Transgenic plants transplanted to soil in the growth chamber. (G-E) Transgenic plants transferred to locking system greenhouse until harvesting. Scale bars: 5 cm in (E), 30 cm in (F), 30 cm in (G), 30 cm in (H).
Regeneration of rice transgenic plants (3-4 weeks)
Transfer resistant calli to REIII regeneration medium containing suitable antibiotics. Using sterilized forceps, subculture a single callus to each well of 24-multi-well plate. Seal the plate with surgical tape and incubate at 30°C under continuous illumination (Figure 5B). Re-transfer transgenic calli to fresh REIII medium every 10 days with the same conditions. The transformed shoot begins to differentiate after 3-4 weeks (Figure 5C).
Transfer 3-4 cm shoots to plantlet pots containing HF medium with appropriate antibiotics at 28°C under 10 h days light (Figure 5D). Several days later, remove the surgical tape from the top of the pots and then remove the pot cover (Figure 5E) (see Note 7).
Transplant transformant rice plants to 6.5 cm pots containing rice seedling raising soil, with one plantlet per pot, in a growth chamber at 28°C (Figure 5F).
When the transformants are 15 cm tall, transplant them to 23-cm pots containing paddy field soil, with one plantlet per pot, in a greenhouse at 28°C until harvest (Figure 5G-5H).
Conduct PCR with DNA from the transformed rice plants to confirm T-DNA integration into rice plants (Figure 6).
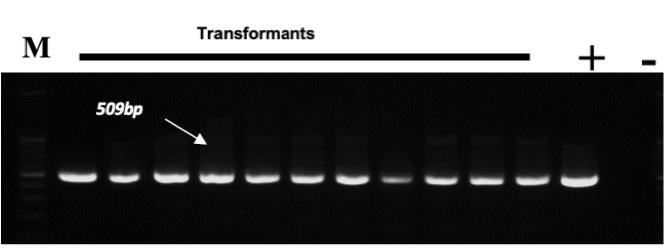
Figure 6. Agarose gel profile for verifying the integration of T-DNA insertion into the plant genome. A 509bp LUC gene amplicon size. M, marker; +, construct DNA; -, Taichung 65 gDNA.
Notes
Bacto agar dissolves well with autoclaving. Add the hygromycin after cooling to 50-45°C because hot culture media will degrade the antibiotic. For A. tumefaciens strain EHA105, add 0.2 ml of 50 mg/ml hygromycin per 200 ml LB medium. For strain EHA 101, add 0.2 ml of 50 mg/ml hygromycin and 0.2 ml 500 mg/ml carbenicillin per 200 ml LB medium.
Meropenem (25 mg/L) in the selection medium completely suppresses the overgrowth of Agrobacterium, which results in high transformation efficiency (Ogawa and Mii, 2007).
Seeds one year after harvesting – rather than seeds taken immediately after harvesting – are preferable for callus induction. Healthy mature seeds are critical as starting material for transformation and high-frequency callus formation. Remove lemma and palea (outer coat) from 40-50 seeds (enough for one transformation). Remove starchy endosperm from the seed to reduce contamination.
Callus selection is a key point for efficient transformation. We often check the culture and remove any contaminated seed parts immediately with a sterilized spoon. In such cases, uncontaminated seed parts are transferred to new plates.
Select four single Agrobacterium colonies. Perform PCR using marker-specific primers for the reporter gene to confirm its incorporation into the plant genome. Bacterial suspension density should be OD600 = 0.2 (Ozawa, 2012) or lower (Nishimura et al., 2006), and OD600 = 0.05-0.1 is recommended. This process is important to prevent excess Agrobacterium growth, which can result in damage to calli. Approximately 150 calli can be placed on a single plate (Hiei and Komari, 2008). Seal plates with Parafilm to prevent evaporation.
Wash calli after co-cultivation. Resistant calli contain small nodular embryos on the surface. A critical step is to remove Agrobacterium from calli by washing without causing much physical damage to the nodular embryo cells, as this would prevent the regeneration of the transgene. Prior the co-cultivation, it is very important to ensure that the selectable-antibiotic is removed appropriately so that it does not affect calli growth. Place the calli on the filter paper to allow the extra water to evaporate.
Hardening: The hardening of transgenic plantlets is a crucial step prior to transplanting them into soil. The hardening can be done slowly from low to high light intensity conditions as well as from high to low humidity. The agar medium can be gently removed from the root by rinsing with water.
Recipes
Sterilizing solution
Solution A
Add 50 ml sodium hypochlorite 10% into 50 ml dH2O
Store at RT
Solution B
Add 50 ml Sodium hypochlorite, 50 ml dH2O, and 50 μl Tween® 20
Store at RT
Antibiotics
Carbenicillin (500 mg/ml)
Dissolve 5 g carbenicillin powder completely in 10 ml dH2O.
Sterilize the solution through 0.22 μm syringe filter.
Store at -20°C in 1 ml aliquots.
Hygromycin (50 mg/ml)
Dissolve 500 mg hygromycin B in 10 ml dH2O.
Sterilize the solution through 0.22-μm syringe filter.
Store at -20°C in 1.5 ml portions.
Kanamycin (50 mg/ml)
Dissolve 500 mg Kanamycin to 10 ml.
Sterilize the solution through 0.22-μm syringe filter.
Store at -20°C in 1.5 ml portions.
Acetosyringone (100 mg/ml)
Dissolve 1 g acetosyringon to 1 ml DMSO.
Dilute with 10 ml dH2O.
Sterilize the solution through 0.22-μm syringe filter.
Store at -20°C in 1.5-2 ml aliquots.
Acetosyringone (10 mg/ml)
Dissolve 100 mg acetosyringon to 1 ml DMSO.
Dilute with 10 ml dH2O.
Sterilize the solution through 0.22-μm syringe filter.
Store at -20°C in 1.5-2 ml aliquots.
Meropenem (12.5 mg/ml)
Dissolved 12.5 g meropenem in 10 ml dH2O.
Sterilize the solution through 0.22-μm syringe filter.
Aliquot in 2 ml portions and store at -20°C.
2,4-D (0.2 mg/ml)
Dissolve 20 mg of 2,4-D powder in 0.5 ml DMSO.
Adjust to 100 ml dH2O.
Store at -20°C.
NAA (0.2 mg/ml)
Dissolve 20 mg of NAA completely in 1 ml 0.1 N sodium hydroxide solution.
Adjust volume to 100 ml dH2O.
Store at -20°C.
Kinetin (1 mg/ml)
Dissolve 10 mg kinetin in 0.2 ml 1 M sodium hydroxide.
Adjust the volume to 10 ml with dH2O.
Store at -20°C.
Prepare cultivation medium (Tanle 1)
Culture nutritional components: All culture media are prepared fresh in 1,000 ml dH2O, autoclaved at 120°C for 20 min and cooled to 50-40°C; finally, it is asepticaly distributed to approximately 12 (90 mm × 20 mm) Petri dishes in a laminar flow hood. The poured medium plates containing antibiotics can be stored at 4°C for up to one month.
Table 1. List of callus induction and regeneration medium components
Medium Components N6D 2N6-AS 2N6D-AS solution N6D-HC REIII HF Sucrose
Glucose30 g
-30 g
10 g30 g
10 g30 g
-30 g
-30 g
-Sorbitol - - - 30 g Casamino acids 0.30 g 0.30 g 0.30 g 0.30 g 1g - Proline 2.878 g - - 2.878 g - - Murashige and Skoog - - - - 4.4 g 4.4 g CHU (N6) 3.981 g 3.981g 3.981g 3.981 g - - CHU N6- Vitamin solution 1 ml 1 ml 1 ml 1 ml - - 2,4-D (0.2 mg/ml) 10 ml 10 ml 10 ml 10 ml - Acetosyringon (10 mg/ml) - 1 ml 1 ml - - - NAA (x 5000) - - - 10 μl Kinetin (x 500) - - - 1 ml Gelrite (0.4%) 4 g 4 g - 4 g 4 g 4 g Carbenicillin 500 (mg/ml) - - - 1 ml - Hygromycin 500 (mg/ml) - - - 1 ml 1 ml Meropenem 12.5 (mg/ml) - - - 2 ml LB solid medium (200 ml)
LB BROTH 5 g
Bacto Agar 1.5 g
Autoclave (120°C for 20 min)
LB liquid medium (200 ml)
LB BROTH 5 g
Autoclave (120°C for 20 min)
YEP medium (200 ml)
Yeast Extract 2 g
Bacto Peptone 2 g
NaCl 1 g
Autoclave (120°C for 20 min)
Acknowledgments
This work was supported in part by a Grant-in-Aid for Scientific Research (KAKENHI) from the Japan Society for the Promotion of Science (Grant number: 19P19394). We are grateful to professor Calvin O. Qualset, University of California Davis, for reading the manuscript and providing comments and suggestions. This protocol is derived from previous publications (Satoh-Cruz et al., 2010; Fukuda et al., 2011 and 2013; Elakhdar et al., 2019).
Competing interests
The authors declare that they have no conflict of interests.
References
- Chan, M. T., Chang, H. H., Ho, S. L., Tong, W. F. and Yu, S. M. (1993). Agrobacterium-mediated production of transgenic rice plants expressing a chimeric alpha-amylase promoter/beta-glucuronidase gene. Plant Mol Biol 22(3): 491-506.
- Dai, S., Zheng, P., Marmey, P., Zhang, S., Tian, W., Chen, S., et al. (2001). Comparative analysis of transgenic rice plants obtained by Agrobacterium-mediated transformation and particle bombardment. Mol Breed 7: 23-25.
- Elakhdar, A., Ushijima, T., Fukuda, M., Yamashiro, N., Kawagoe, Y. and Kumamaru, T. (2019). Eukaryotic peptide chain release factor 1 participates in translation termination of specific cysteine-poor prolamines in rice endosperm. Plant Sci 281: 223-231.
- Fukuda, M., Satoh-Cruz, M., Wen, L., Crofts, A. J., Sugino, A., Washida, H., et al. (2011). The small GTPase Rab5a is essential for intracellular transport of proglutelin from the golgi apparatus to the protein storage vacuole and endosomal membrane organization in developing rice endosperm. Plant Physiol 157(2): 632-644.
- Fukuda, M., Wen, L., Satoh-Cruz, M., Kawagoe, Y., Nagamura, Y., Okita, T. W., et al. (2013). A guanine nucleotide exchange factor for Rab5 proteins is essential for intracellular transport of the proglutelin from the golgi apparatus to the protein storage vacuole in rice endosperm. Plant Physiol 162(2): 663-674.
- Furtado, A., Henry, R. J. and Takaiwa, F. (2008). Comparison of promoters in transgenic rice. Plant Biotechnol J 6(7): 679-693.
- Gelvin, S. B. (2010). Plant proteins involved in Agrobacterium-mediated genetic transformation. Annu Rev Phytopathol 48: 45-68.
- Gupta, P., Raghuvanshi, S., and Tyagi, A. K. (2001). Assessment of the Efficiency of Various Gene Promoters via Biolistics in Leaf and Regenerating Seed Callus of Millets, Eleusine coracana and Echinochloa crusgalli. Plant Biotechnol 18(4): 275-282.
- Hakata, M., Hidemitsu, N., Keiko, I., Akio, M., Mariko, K., Naoko, I., Jinhuan, P., Kou, A., Akihiko, H., Tomoko, T., Jianyu, S., Motoko, I., Hiroko, K., Takanari, I., Minami, M., Shoshi, K., Yoshiaki, N., Hirohiko, H. and Hiroaki, I. (2010). Production and characterization of a large population of cDNA-overexpressing transgenic rice plants using Gateway-based full-length cDNA expression libraries. Breeding Science 60(5): 575-585.
- Hiei, Y. and Komari, T. (2008). Agrobacterium-mediated transformation of rice using immature embryos or calli induced from mature seed. Nat Protoc 3(5): 824-834.
- Hiei, Y., Komari, T. and Kubo, T. (1997). Transformation of rice mediated by Agrobacterium tumefaciens. Plant Mol Biol 35(1-2): 205-218.
- Hiei, Y., Ohta, S., Komari, T. and Kumashiro, T. (1994). Efficient transformation of rice (Oryza sativa L.) mediated by Agrobacterium and sequence analysis of the boundaries of the T-DNA. Plant J 6(2): 271-282.
- Hood, E. E., Gelvin, S. B., Melchers, L. S., and Hoekema, A. (1993). New Agrobacterium helper plasmids for gene transfer to plants. Transgenic Res 2(4): 208-218.
- Iglesias, V. A., Moscone, E. A., Papp, I., Neuhuber, F., Michalowski, S., Phelan, T., Spiker, S., Matzke, M. and Matzke, A. J. (1997). Molecular and cytogenetic analyses of stably and unstably expressed transgene loci in tobacco. Plant Cell 9(8): 1251-1264.
- Iso, E. (1957). Lecture on Rice Culture. Department of Economics, Ryukyu District Government. 491pp (in Japanese).
- Kumamaru, T., Uemura, Y., Inoue, Y., Takemoto, Y., Siddiqui, S. U., Ogawa, M., Hara-Nishimura, I. and Satoh, H. (2010). Vacuolar processing enzyme plays an essential role in the crystalline structure of glutelin in rice seed. Plant Cell Physiol 51(1): 38-46.
- Nishimura, A., Aichi, I. and Matsuoka, M. (2006). A protocol for Agrobacterium-mediated transformation in rice. Nat Protoc 1(6): 2796-2802.
- Ogawa, Y., and Mii, M. (2007). Meropenem and moxalactam: Novel β-lactam antibiotics for efficient Agrobacterium-mediated transformation. Plant Sci 172(3): 564-572.
- Ozawa, K. (2012). A high-efficiency Agrobacterium-mediated transformation system of rice (Oryza sativa L.). Methods Mol Biol 847: 51-57.
- Ozawa, K., and Takaiwa, F. (2010). Highly efficient Agrobacterium-mediated transformation of suspension-cultured cell clusters of rice (Oryza sativa L.). Plant Sci 179(4): 333-337.
- Park, S. H., Pinson, S. R. M., and Smith, R. H. (1996). T-DNA integration into genomic DNA of rice following Agrobacterium inoculation of isolated shoot apices. Plant Mol Biol 32(6): 1135-1148.
- Park, S. H., Yi, N., Kim, Y. S., Jeong, M. H., Bang, S. W., Choi, Y. D. and Kim, J. K. (2010). Analysis of five novel putative constitutive gene promoters in transgenic rice plants. J Exp Bot 61(9): 2459-2467.
- Pawlowski, W. P. and Somers, D. A. (1996). Transgene inheritance in plants genetically engineered by microprojectile bombardment. Mol Biotechnol 6(1): 17-30.
- Saika, H. and Toki, S. (2010). Mature seed-derived callus of the model indica rice variety Kasalath is highly competent in Agrobacterium-mediated transformation. Plant Cell Rep 29(12): 1351-1364.
- Satoh-Cruz, M., Crofts, A. J., Takemoto-Kuno, Y., Sugino, A., Washida, H., Crofts, N., et al. (2010). Protein disulfide isomerase like 1-1 participates in the maturation of proglutelin within the endoplasmic reticulum in rice Endosperm. Plant Cell Physiol 51(9): 1581-1593.
- Toki, S. (1997). Rapid and efficient Agrobacterium-mediated transformation in rice. Plant Mol Biol Report. 15: 16-21.
- Zhou, J., Yang, Y., Wang, X., Yu, F., Yu, C., Chen, J., Cheng, Y., Yan, C. and Chen, J. (2013). Enhanced transgene expression in rice following selection controlled by weak promoters. BMC Biotechnol 13: 29.
Article Information
Copyright
© 2021 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Elakhdar, A., Fukuda, M. and Kubo, T. (2021). Agrobacterium-mediated Transformation of Japonica Rice Using Mature Embryos and Regenerated Transgenic Plants. Bio-protocol 11(18): e4143. DOI: 10.21769/BioProtoc.4143.
Category
Plant Science > Plant transformation > Agrobacterium
Environmental science > Plant
Molecular Biology > DNA > DNA cloning
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link








