- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
High-throughput Method for Detecting Siderophore Production by Rhizosphere Bacteria
Published: Vol 11, Iss 9, May 5, 2021 DOI: 10.21769/BioProtoc.4001 Views: 5810
Reviewed by: Yufang LuAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Detecting Photoactivatable Cre-mediated Gene Deletion Efficiency in Escherichia coli
Yuta Koganezawa [...] Miki Umetani
Jun 5, 2023 1970 Views
Co-culture Wood Block Decay Test with Bacteria and Wood Rotting Fungi to Analyse Synergism/Antagonism during Wood Degradation
Julia Embacher [...] Martin Kirchmair
Oct 5, 2023 1900 Views

Quantifying Bacterial Chemotaxis in Controlled and Stationary Chemical Gradients With a Microfluidic Device
Adam Gargasson [...] Harold Auradou
Feb 20, 2025 1752 Views
Abstract
Siderophores, a key substance that microorganisms produce to obtain iron under iron-limited conditions, play an important role in regulating interactions between beneficial bacteria and pathogenic bacteria. A large number of bacteria were isolated from the rhizosphere, and we used the method presented here to assay the siderophore production by these rhizosphere bacteria. This method is a modified version of the universal chrome azurol S (CAS) assay that uses a 96-channel manual pipetting workstation. By combining the liquid CAS assay with the multi-channel pipette workstation, high-throughput and rapid detection of siderophore production can be achieved. In summary, this method can be used to gain a general understanding of siderophore production by rhizosphere bacteria.
Keywords: Rhizosphere bacteriaBackground
Siderophores have an extremely strong affinity for ferric iron and are extremely important to bacterial survival. Almost all known bacterial species can produce siderophores (Miethke and Marahiel, 2007), which affect their interaction with microorganisms in close proximity. Siderophores produced by neighboring organisms can act as growth factors that promote the growth of unculturable microorganisms (Kaeberlein et al., 2002), and they also play a significant role in the biological control mechanism against certain phyto-pathogens. Over forty years ago, Kloepper et al. (1980) were the first to illustrate the importance of siderophore production as a mechanism of biological control of Erwinia carotovora by several plant growth-promoting Pseudomonas fluorescens strains such as A1, BK1, TL3B1, and B10. Siderophores play an important role in the composition and function of the rhizosphere microbiome and plant health.
The chrome azurol S (CAS) assay, the most common method for detecting siderophore production (Schwyn and Neilands, 1987), is based on a competition for Fe3+ between the ferric complex of the dye CAS and the siderophore. Increasing the knowledge of siderophore production by rhizosphere bacteria will contribute to a better understanding of the interactions among rhizosphere bacteria. High-throughput detection of siderophore production can be achieved by combining the CAS assay with a 96-channel manual pipetting workstation. In this method, siderophore production was assayed by a modified version of the universal chemical assay developed by Schwyn and Neilands (1987) in 2,150 representative bacterial members, which will help to better understand the abilities of rhizobacteria to produce siderophores.
Materials and Reagents
96-well plate (96-well clear; Costar, catalog number: 3599)
Disposable Petri dish (90 mm; Jiangsu Kangjian Medical Products, catalog number: 161-0901)
96-well plate with 0.22-μm filter membranes (MultiScreenHTS GV Filter Plate, 0.22 µm, clear, sterile; Millipore®, catalog number: MSGVS2210)
Pseudomonas aeruginosa PAO1 (Ghysels et al., 2005)
Burkholderia cepacia H111 (Sathe et al., 2019)
Pseudomonas aeruginosa PAO1ΔpvdDΔpchEF (Ghysels et al., 2005)
Pseudomonas aeruginosa H111ΔorbJΔpchAB (Sathe et al., 2019)
Tryptone (OXOID, catalog number: LP0042)
Soy peptone (Chemical Reagent Co., Ltd. of Sinopharm Group, Shanghai Test, catalog number: 69047737)
NaCl (Nanjing Chemical Reagent Co., Ltd. CAS: 7647-14-5)
Agar (Fujian Jinyan Marine Biotechnology Co., Ltd., Chengfeng. CAS: 9002-18-0)
K2PHO4 (Nanjing Chemical Reagent Co., Ltd. CAS: 7778-77-0)
MgSO4·7H2O (Nanjing Chemical Reagent Co., Ltd. CAS: 10034-99-8)
Glycerol (Chemical Reagent Co., Ltd. of Sinopharm Group, Shanghai Test, catalog number: 10010618)
Casamino acids (DSLAB, catalog number: 18A0050)
FeCl3 (Chemical Reagent Co., Ltd. of Sinopharm Group, Shanghai Test, catalog number: 10011918)
Tris-HCl (1 M Tris-HCl, pH = 6.8, BL514A, 100 ml)
Gelatin (Chemical Reagent Co., Ltd. of Sinopharm Group, Shanghai Test, catalog number: 10010328)
Chrome azurol S (Chromeazurol S; Fluka, catalog number: 72687-25G)
HTDMA (VETEC, catalog number: V900413-100G)
Anhydrous piperazine (Piperazine, Reagentplus 99%, Sigma-Aldrich, CAS: 110-85-0)
Glucose (Nanjing Chemical Reagent Co., Ltd. CAS: 50-99-7)
Yeast extract (YEAST EXTRACT; OXOID, catalog number: LP0021)
Beef extract (Chemical Reagent Co., Ltd. of Sinopharm Group, Wokai, catalog number: 69004461)
Glycerin (Nanjing Chemical Reagent Co., Ltd. CAS: 56-81-5)
1/10 tryptone soya agar (1/10 TSA) (see Recipes)
1/10 tryptone soya broth (1/10 TSB) (see Recipes)
MKB medium (see Recipes)
Iron-rich medium (see Recipes)
MS buffer solution (see Recipes)
CAS assay solution (see Recipes)
Equipment
96-channel manual pipetting workstation (Suzhou SinoAnalysis Instrument Co., Ltd., SinoAnalysis, SC9000)
Microplate centrifuge (Hunan Hirsch Instruments, Benchtop Low Speed Centrifuge, TD5A)
Constant-temperature shaker (MIN QUAN, MQD-BIR)
Microplate reader (SpectraMax M5, Sunnyvale, CA, USA)
-80°C freezer (Haier, vertical ultra-low temperature storage box, DW-86L62, 2013 model)
Centrifuge (Eppendorf AG, 22331 Hamburg, 5424EH062551)
Pipette gun (Eppendorf Research plus)
Autoclave (Tega SANYO Industry Co., Ltd, MLS-3780, Tottori, Japan)
Constant-temperature and -humidity incubator (new seedlings, waterproof electric heating constant-temperature incubator, GNP-9080BS-III)
Balance (Sartorius, model: BSA2202S)
Vortex (SCIENTIFIC INDUSTRIES, USA)
Software
R 3.1.2 program (https://github.com/shaohuagu/sidanalysis.git)
Excel 2016
SPSS 20.0
Adobe Illustrator CS5
Procedure
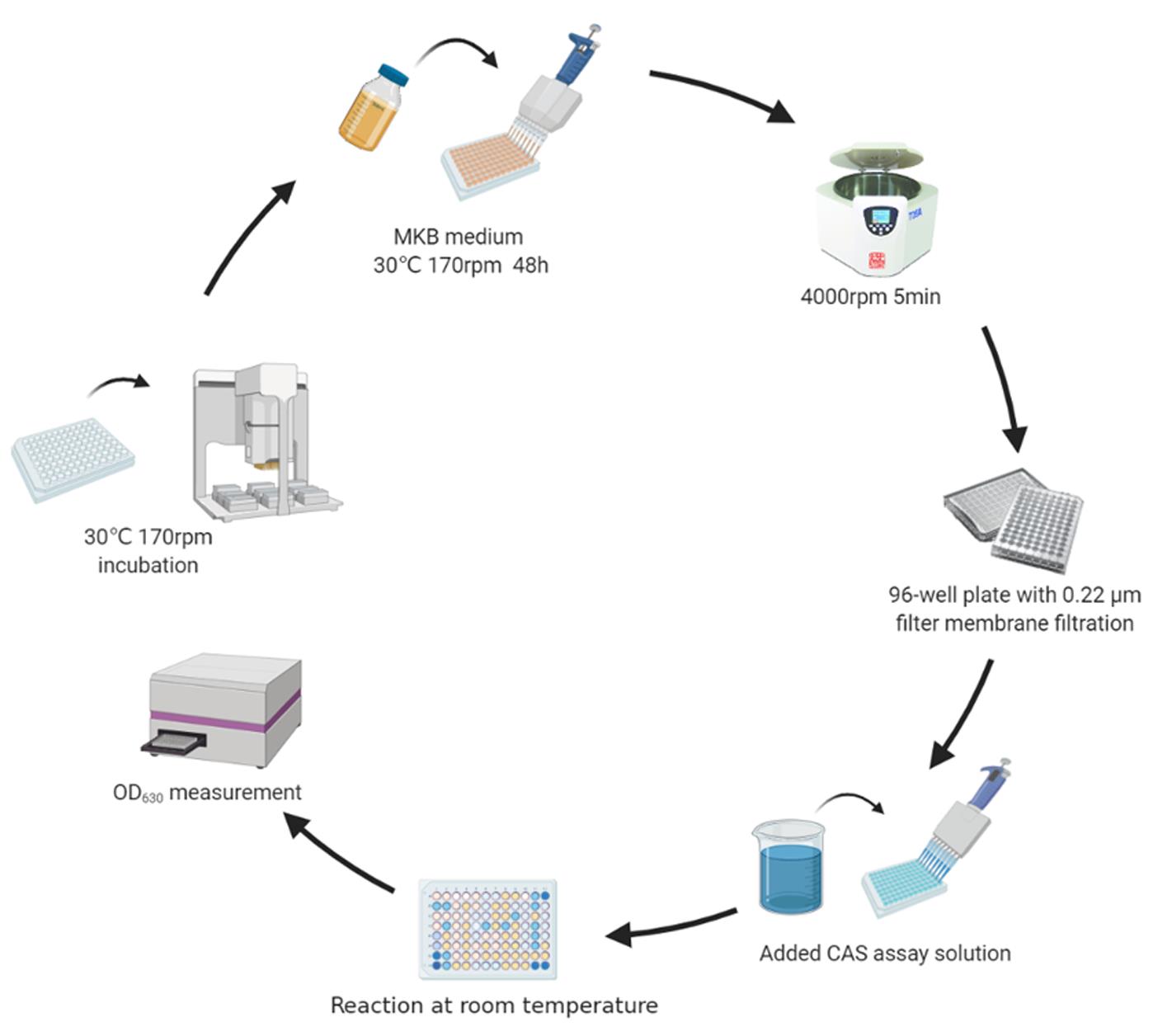
Figure 1. Determination of siderophore production
Rhizosphere soil sampling
Rhizosphere soil samples were collected from tomato plants located in four different fields. The excess soil should be gently shaken off and discarded; the remaining soil attached to the roots is considered the rhizosphere soil (Hu et al., 2016) and should be collected for use.
Isolation of rhizobacteria
Mix 1 g rhizosphere soil with 9 ml MS buffer solution in a rotary shaker (170 rpm) at 30°C for 30 min.
Dilute the soil suspension to a concentration of 10-5-10-6 g/ml with sterile water. Spread 100 μl diluted soil suspensions on 1/10 tryptone soya agar (TSA).
After a 48-h incubation at 30°C in the dark, randomly pick 32 isolates per rhizosphere soil sample and restreak on TSA plates for colony purification.
Culture all purified isolates in 100 μl tryptone soya broth (TSB) in 96-well microtiter plates at 30°C with shaking (rotary shaker at 170 rpm) for 18 h.
Add 100 μl 30% (v/v) glycerin to the fermentation broth and mix well. Store the rhizobacteria at -80°C.
Measuring siderophore production of rhizobacteria (Figure 1)
Revive the isolates by transferring 5 μl respective freezer stocks into a clean 96-well plate containing 195 μl TSB per well. Culture the bacteria overnight at 30°C with shaking (rotary shaker set at 170 rpm).
Transfer 10 μl overnight cultures into clean 96-well plates containing 190 μl MKB iron-limited medium and MKB iron-rich medium. Incubate for 48 h at 30°C with shaking (rotary shaker set at 170 rpm).
Harvest the cell-free supernatant from the bacterial cultures by centrifugation (4,000 rpm, 5 min at 4°C) and filtration (using a 0.22-µm filter).
Use the liquid version of the chrome azurol S (CAS) assay, in which 100 µl cell-free supernatant (three biological replicates for each of the 2,150 soil isolates) is added to 100 μl CAS assay solution in a 96-well plate using a 96-channel manual pipetting workstation. Add 100 μl MKB iron-limited meduim or MKB iron-rich medium to 100 μl CAS assay solution as a control group.
Incubate the reaction mixture without agitation for 2 h at room temperature.
The OD630 of the reaction mixture (A) and the control group (Ar) was measured using a plate reader (SpectraMax M5) at room temperature. Siderophores induce a color change in the CAS medium, which lowers the OD630 measurements.
Siderophore production was quantitated using the following formula: 1 − A ÷ Ar.
Organic acids in media components and other secreted compounds can also bind iron; therefore, it is essential to estimate the CAS signal background that is not due to siderophores. We assessed this signal background using defined siderophore-deficient mutants from two species (P. aeruginosa PAO1ΔpvdDΔpchEF and B. cenocepacia H111ΔorbJΔpchAB) (Ghysels et al., 2005; Sathe et al., 2019) and their corresponding wild types using the same protocol as described above. We then averaged the CAS background signals of the two siderophore-deficient mutants, which was used as a cut-off to distinguish siderophore producers from non-producers among our 2,150 rhizobacteria(Figure 2A).
Data analysis
Isolation of rhizobacteria
Almost 16% of the single colonies picked at random could not grow individually on TSA plates. Finally, 2,150 purified strains were isolated from 80 rhizosphere soil samples.
Measuring siderophore production by rhizobacteria
The calculation for the relative production of siderophores was obtained using this formula: siderophore unit = 1 − A ÷ Ar, which was used to estimate the levels of secreted siderophores in the supernatant of all 2,150 bacterial isolates. This assay serves as a proxy for siderophore production and revealed that up to 95% of the isolates produced siderophores, given that their CAS values exceeded those of siderophore-deficient control strains(Figure 2B). Estimating background CAS values is important since this assay also measures the binding activities of other organic iron-binding compounds. When the CAS assay was repeated under iron-rich conditions, we found that up to 99% of the siderophore producers upregulated siderophore production under iron-limited conditions as compared with under iron-rich conditions. Under iron limitation, siderophore production followed a bimodal distribution, with isolates producing either high or low quantities of siderophores(Figure 2B). These results suggest that siderophore production is a widespread trait across the taxa and sampling sites examined here under iron-limited conditions.
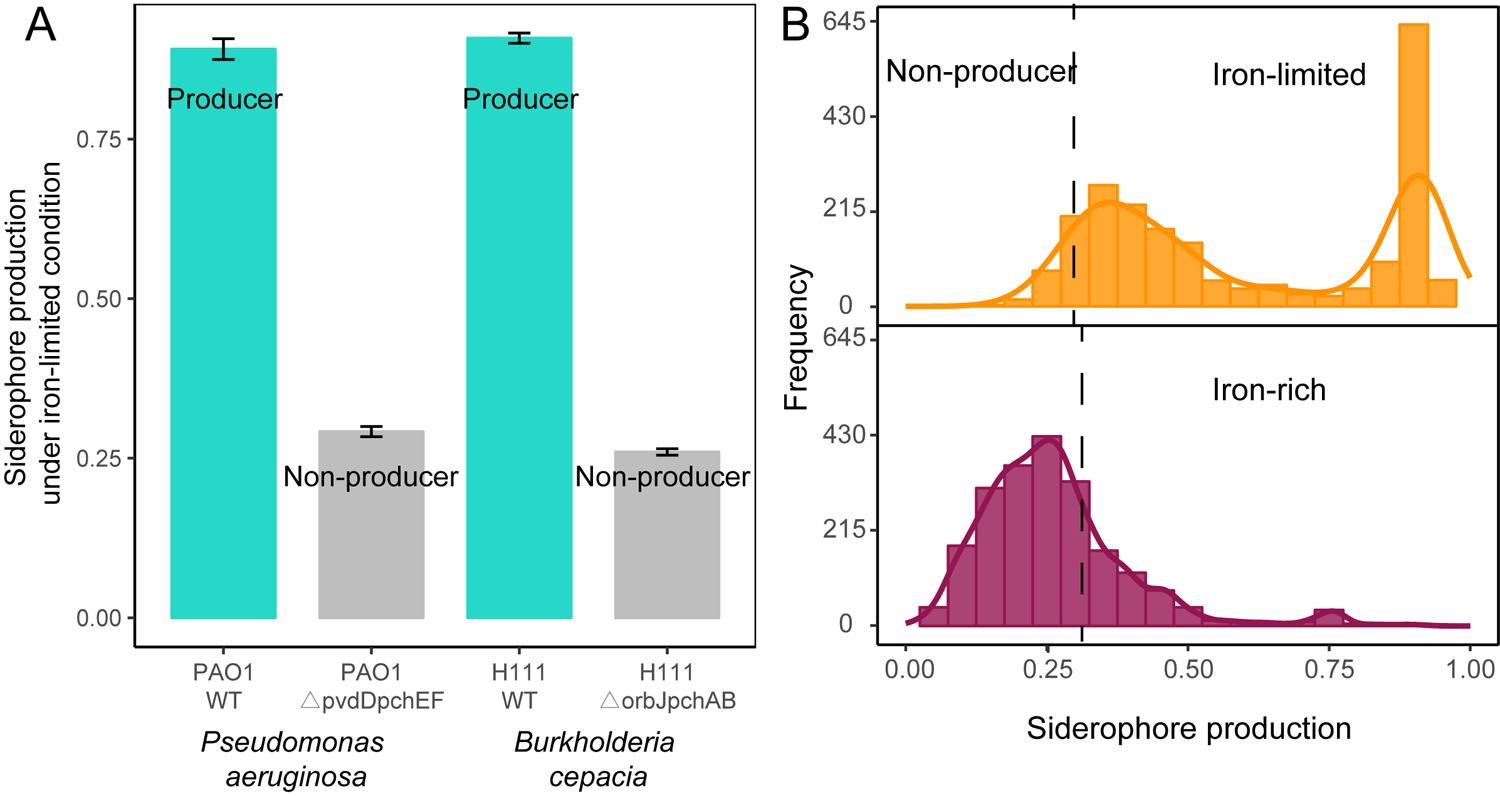
Figure 2. Siderophore production of rhizobacteria under iron-limited and iron-rich conditions
Notes
During the preparation of MKB medium, all supplies, including glass bottles, measuring cylinders, glass rods, and weighing spoons, must be iron-free. Glassware must be soaked in 6 M hydrochloric acid overnight and rinsed with distilled water several times to remove iron.
Both MKB medium solutions need to be separately prepared and autoclaved at 115°C for 30 min.
Recipes
1/10 tryptone soya agar (1/10 TSA)
1.5 g/L tryptone
0.5 g/L soytone
0.5 g/L sodium chloride
15 g/L agar
pH 7.0
1/10 tryptone soya broth (1/10 TSB)
1.5 g/L tryptone
0.5 g/L soytone
0.5 g/L sodium chloride
pH 7.0
MKB medium
2.5 g/L K2HPO4
2.5 g/L MgSO4·7H2O
15 ml/L glycerin
5.0 g/L casamino acids
pH 7.2
Iron-rich medium
Add FeCl3 to MKB medium to a final concentration of 50 μM
MS buffer solution
50 mM Tris-HCl, pH 7.5
100 mM NaCl
10 mM MgSO4
0.01% gelatin
CAS assay solution
Add 1.5 ml 1 mM FeCl3 to 7.5 ml 1 mM Chrome azurol S and mix well. Add 50 ml 4 mmol/L HTDMA while stirring, followed by 30 ml 1 M piperazine solution. Finally, add deionized water to 100 ml
Acknowledgments
This research was financially supported by the National Natural Science Foundation of China (grant nos. 41922053 to Z.W., 41807045 to T.Y. and 31972504 to Y.X.) and the Natural Science Foundation of Jiangsu Province (grant nos. BK20180527 to T.Y. and BK20170085 to Z.W.). The Laboratory of Rhizosphere Micro-ecology (LorMe), College of Resources and Environmental Sciences, Nanjing Agricultural University, guaranteed the successful completion of this experiment.
Competing interests
The authors declare no competing interests.
Ethics
The experimental procedures involved in this article are in line with ethical and moral requirements, and have not caused adverse consequences to society or the environment.
References
- Ghysels, B., Ochsner, U., Mollman, U., Heinisch, L., Vasil, M., Cornelis, P. and Matthijs, S. (2005). The Pseudomonas aeruginosa pirA gene encodes a second receptor for ferrienterobactin and synthetic catecholate analogues. FEMS Microbiol Lett 246(2): 167-174.
- Hu, J., Wei, Z., Friman, V. P., Gu, S. H., Wang, X. F., Eisenhauer, N., Yang, T. J., Ma, J., Shen, Q. R., Xu, Y. C. and Jousset, A. (2016). Probiotic Diversity Enhances Rhizosphere Microbiome Function and Plant Disease Suppression. mBio 7(6).
- Kaeberlein, T., Lewis, K., Epstein, S. S. (2002). Isolating "uncultivable" microorganisms in pure culture in a simulated natural environment. Science 296(5570): 1127-1129.
- Kloepper, J. W., Leong, J., Teintze, M. and Schroth, M. N. J. C. M. (1980). Pseudomonas siderophores: A mechanism explaining disease-suppressive soils. 4(5): 317-320.
- Miethke, M. and Marahiel, M. A. (2007). Siderophore-based iron acquisition and pathogen control. Microbiol Mol Biol Rev 71(3): 413-451.
- Sathe, S., Mathew, A., Agnoli, K., Eberl, L. and Kummerli, R. (2019). Genetic architecture constrains exploitation of siderophore cooperation in the bacterium Burkholderia cenocepacia. Evol Lett 3(6): 610-622.
- Schwyn, B. and Neilands, J. B. (1987). Universal chemical assay for the detection and determination of siderophores. Anal Biochem 160(1): 47-56.
Article Information
Copyright
© 2021 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Gu, S., Wan, W., Shao, Z. and Wei, Z. (2021). High-throughput Method for Detecting Siderophore Production by Rhizosphere Bacteria. Bio-protocol 11(9): e4001. DOI: 10.21769/BioProtoc.4001.
Category
Microbiology > Microbial physiology > Adaptation
Biological Sciences > Microbiology > Microbial communities
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










