- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
AIMTOR, a BRET Biosensor for Live Recording of mTOR Activity in Cell Populations and Single Cells
(*contributed equally to this work) Published: Vol 11, Iss 8, Apr 20, 2021 DOI: 10.21769/BioProtoc.3989 Views: 4460
Reviewed by: Joachim GoedhartSalma MerchantAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
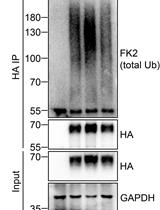
Analysis of the Ubiquitination and Phosphorylation of Vangl Proteins
Di Feng [...] Bo Gao
Oct 20, 2022 3466 Views
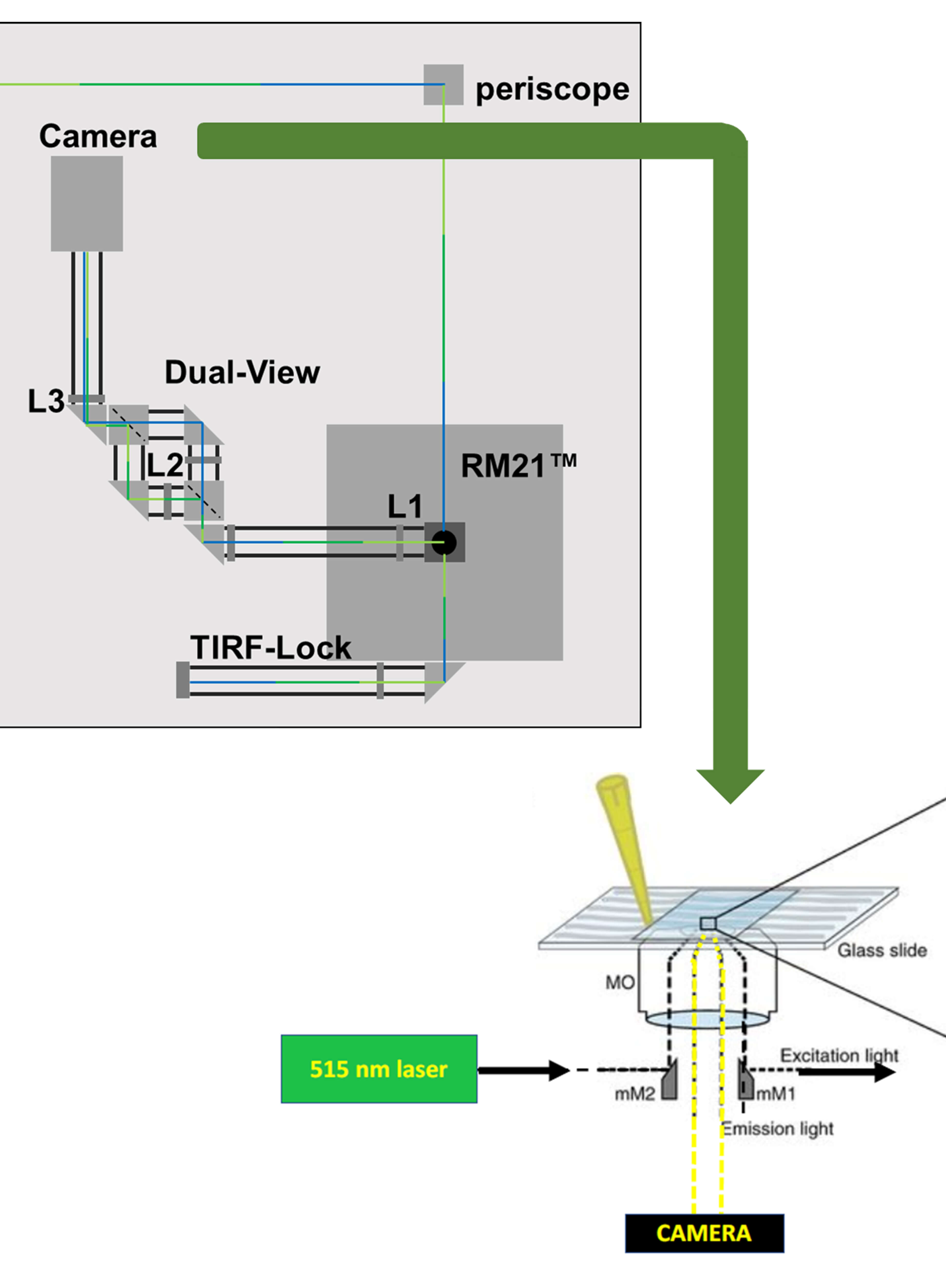
Imaging Membrane Proteins Using Total Internal Reflection Fluorescence Microscopy (TIRFM) in Mammalian Cells
Kirin D. Gada [...] Leigh D. Plant
Feb 20, 2023 2013 Views
Abstract
Mammalian target of rapamycin (mTOR) controls many crucial cellular functions, including protein synthesis, cell size, energy metabolism, lysosome and mitochondria biogenesis, and autophagy. Consequently, deregulation of mTOR signaling plays a role in numerous pathological conditions such as cancer, metabolic disorders and neurological diseases. Developing new tools to monitor mTOR spatiotemporal activation is crucial to better understand its roles in physiological and pathological conditions. However, the most widely used method to report mTOR activity relies on the quantification of specific mTOR-phosphorylated substrates by western blot. This approach requires cellular lysate preparation, which restricts the quantification to a single time point. Here, we present a simple protocol to study mTOR activity in living cells in real time using AIMTOR, an intramolecular BRET-based (bioluminescence resonance energy transfer) biosensor that we recently designed (Bouquier et al., 2020). We describe transfection of AIMTOR in the C2C12 cell line and procedures to monitor BRET in a cell population using a plate reader and in single cells by microscopy. Importantly, this protocol is transposable to any cell line and primary cells. In addition, several subcellular compartment-specific versions of AIMTOR have been developed, enabling compartmentalized assessment of mTOR activity. This protocol describes how to use the sensitive AIMTOR biosensor to investigate mTOR signaling dynamics in living cells.
Graphic abstract:
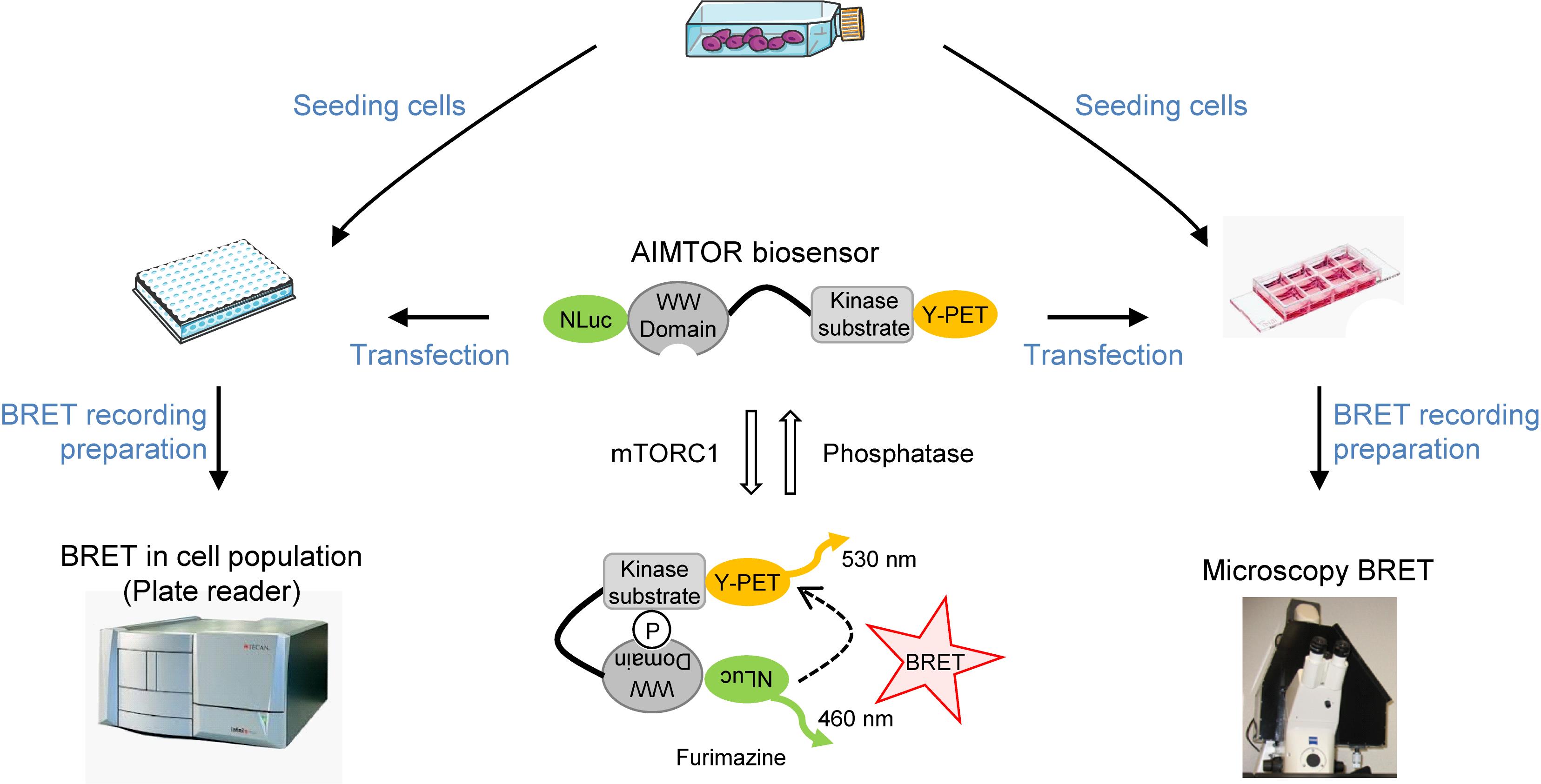
AIMTOR protocol overview from seeding cells to live BRET recording
Background
The mTOR signaling pathway is conserved within the eukaryotic domain and controls major metabolic and anabolic pathways involved in cellular homeostasis, including proteostasis, lipogenesis, and glucose and mitochondrial metabolism. The mTOR kinase occupies the central stage of this signaling pathway and its activation/inhibition depends on a myriad of environmental cues, including nutrient availability, growth factors, energy status and various stresses. mTOR signaling is regulated by complex and sometimes redundant mechanisms depending on cellular type and context. As expected given its central role in cellular homeostasis, many if not all human physiopathological conditions involve mTOR signaling to some degree or will at least interfere with its activation/regulation at some stage of their progression. The mTOR kinase belongs to several multiprotein complexes, including mTORC1 and mTORC2, which exhibit distinct cellular functions and substrate specificities. The mTORC1 complex primarily controls cell growth and anabolism pathways, whereas the mTORC2 complex is involved in regulation of cellular metabolism (Saxton and Sabatini, 2017). Efforts are underway to better investigate mTOR signaling and activation modes in several cell types. To this end, specific tools are required to accurately monitor mTOR kinase activity in living cells in real time.
However, current methods mostly rely on endpoint approaches requiring cell lysis, precluding dynamic studies of live cells and kinetic investigations. For example, western blot analysis requires quantification of the abundance of several specific endogenous substrates phosphorylated by mTOR in cell or tissue protein lysates. To circumvent this drawback, genetically encoded biosensors have been designed that allow kinase activity to be monitored in living cells. Indeed, fluorescence resonance energy transfer (FRET) biosensors have been developed to allow imaging of the activity of several important signaling kinases in living cells, including ERK, S6K, AKT, and mTOR. These intramolecular FRET biosensors monitor conformational change-induced shifts in the proximity between donor and acceptor fluorophores upon phosphorylation of a specific kinase substrate. However, drawbacks related to the use of excitation light, such as cell auto-fluorescence, spectral overlaps between donor and acceptor excitation and fluorophore bleaching, complicate the analysis and interpretation of FRET data. Because it does not require light excitation, bioluminescence resonance energy transfer (BRET) circumvents the major technical limitations of FRET. The donor for BRET is a luciferase entity that emits light only after chemical activation. BRET has been shown to be an efficient approach not only for protein/protein interaction studies, but also for the design of biosensors to monitor enzymatic activity.
Here, we describe a protocol in which the genetically encoded BRET biosensor AIMTOR (Bouquier et al., 2020) is used to monitor mTOR cell signaling by BRET imaging in individual cells under a microscope or in cell populations with a plate reader. This detailed protocol was recently validated to investigate mTORC1 signaling and dysfunction in muscle cells and primary neurons (Bouquier et al., 2020). However, AIMTOR can be used in virtually all types of cultured cells, cell lines or other primary cells. AIMTOR enables delineation of mTOR signaling, identification of activators and inhibitory conditions, and screening of drugs/compounds that interfere with this important cellular hub. In addition, several versions of AIMTOR targeted to various subcellular compartments have been designed to better characterize the existence of discrete compartmentalized pools of mTOR activity. Future versions of AIMTOR utilizing new vectors containing tissue-specific promoters or other subcellular tags will expand the range of methods available for monitoring mTOR signaling directly in live cells and organs.
Materials and Reagents
100 mm TC-treated Culture Dish (Corning, catalog number: 430167 ) and TTP 6-well plates (Sigma, catalog number: Z707759 )
8-well IbiTreat µ-Slide (ibidi, catalog number: 80826) or 8-well Polylysine-coated Slides (ibidi, catalog number: 80824)
96-well clear bottom white microplate (Greiner, catalog number: 655098 )
1.5 ml and 15 ml tubes
C2C12 mouse myoblast cell line (ATCC, catalog number: CRL-1772 )
AIMTOR plasmid DNA T757 (Addgene ID 140828) and nonphosphorylatable A757 mutant (Addgene ID 140829), store at -20 °C
Dulbecco’s Modified Eagle’s Medium (Sigma, catalog number: D5796-500ml ), store at 4 °C
Dulbecco’s Modified Eagle’s Medium (Gibco, without phenol red, glutamine and glucose A1443001-500 ml), store at 4 °C
Glucose 20% (1.11 M), filtered and sterilized; store at 4 °C. Glucose 20% is used to complement phenol red-free and glucose-free DMEM with 25 mM glucose.
Fetal bovine serum (Dominique Dutscher, catalog number: S1810-500), store at -20 °C
Glutamine 200 mM (Sigma, catalog number: G7513-100ml ), store at -20 °C
Na pyruvate 100 mM (Sigma, catalog number: S8636-100ml), store at 4 °C
Antibiotics penicillin and streptomycin, 10,000 U/ml (Sigma, catalog number: P4333 ), store at -20 °C
HEPES buffer, 1 M, filtered and sterilized (Sigma, catalog number: H3662 ), store at 4 °C
PBS (Sigma, catalog number: D8537-500 ml), store at room temperature
Trypsin/EDTA (Sigma, catalog number: T4049-100ml ), store at 4 °C
Furimazine (Promega, Nano-Glo luciferase assay, catalog number: N1120 ), store at -20 °C
JETPEI (Polyplus Transfection Reagent, catalog number: 101-10N ), store at 4 °C
Helix-IN DNA transfection reagent (OZBiosciences, catalog number: HX10100, store at -20 °C) or any agent to efficiently transfect your cells of interest without affecting cell phenotype and viability
DMEM regular growth medium (contains phenol red) (see Recipes)
DMEM growth medium and reading medium (see Recipes)
Equipment
MicroPipettes
Multicanal pipette (10-200 µl) and/or repipetor with 500 µl-10 µl tip (Eppendorff)
Water-jacketed CO2 incubator
Class II biological safety cabinet
Thoma counting chamber or other cell counting device
Centrifuge (SIGMA, model: 3-16KL ) with a rotor (SIGMA, catalog number: 11222) suitable for 6-well plates (for JETPEI transfection)
For BRET population measurement
SynergyTM 2 Biotek Microplate reader (or an equivalent microplate reader) allowing sensitive measurement of bioluminescence filtered emission light around 460 nm (emission peak of nanoluciferase induced by furimazine) and 530 nm (emission peak of YPET)
Emission filters: 485/20 nm (Biotek, catalog number: 7082221) and 530/25 nm (Biotek, catalog number: 7082223)
Gen5TM Reader Control and Data Analysis Software to program the plate reader and generate raw data files (in Excel, Microsoft OfficeTM)
For BRET imaging
Homemade dark box with sliding trap (Figure 1A)
Computer with Metamorph software (Figure 1B)
Bioluminescence-dedicated microscope device customized from an inverted fluorescence microscope (Carl Zeiss, Axiovert 200M) (Coulon et al., 2008; Perroy, 2010) (Figure 1C). Please note that due to the strong brightness of NanoLuc, it is now possible to use a classic inverted microscope (Kim and Grailhe, 2016)
Evolve camera equipped with an EMCCD detector, back-illuminated, with On-chip Multiplication Gain (Photometrics) (Figure 1D)
Multiposition stage control (Marzhauser Wetzlar, Tango 2 Desktop, catalog number: 00-76-150-1802) (Figure 1E)
Illumination source: Lumencor (Gataca, Spectra X light engine) (Figure 1F)
Plan-Neofluar 40×/1.3 oil objective M27 (Carl Zeiss)
Emission filters: 450/70 nm (Semrock, catalog number: FF01-450/70-25) and HQ535/50nm (Chroma, catalog number: 63944)
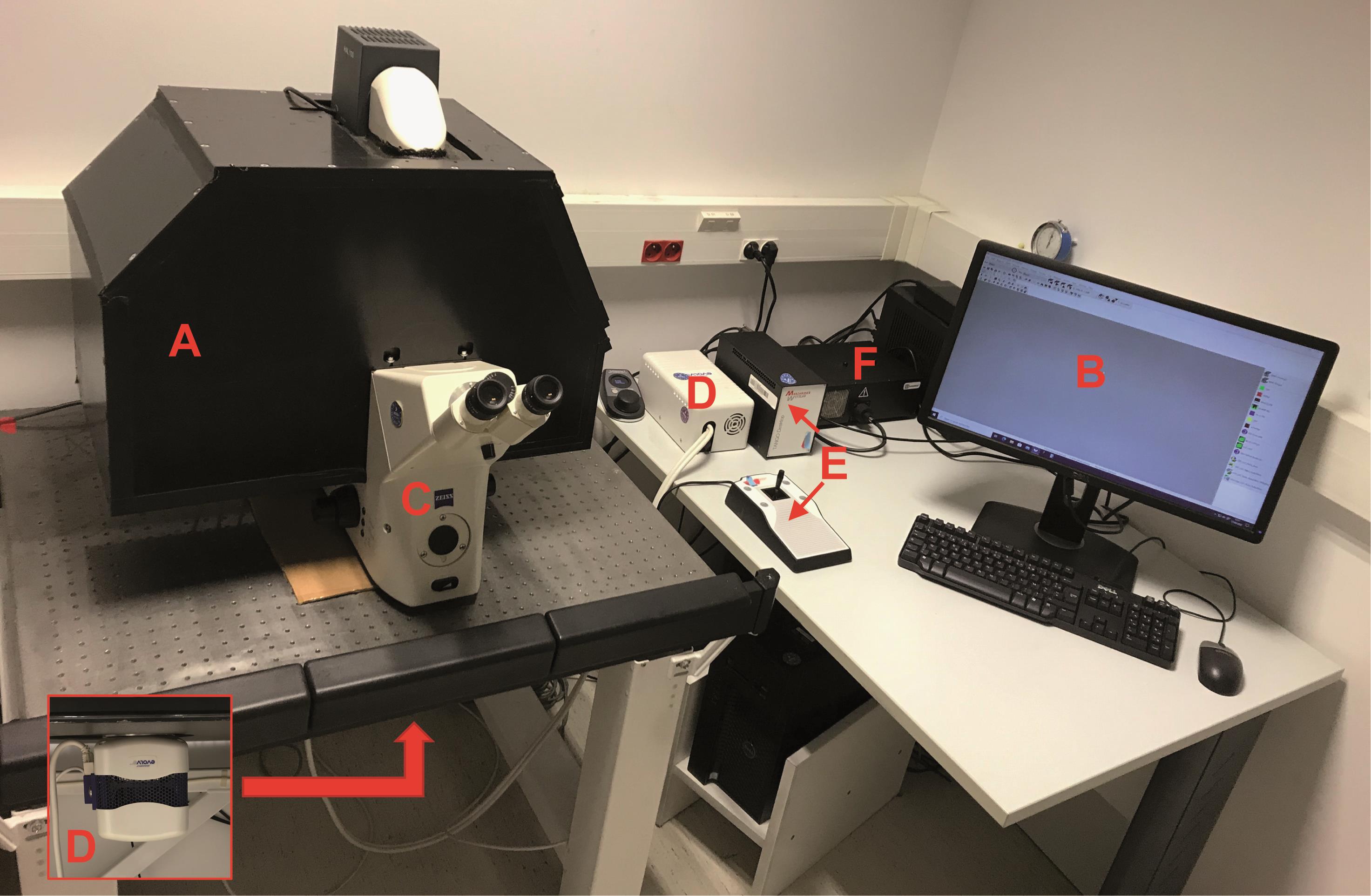

Figure 1. Microscopy BRET imaging setup. A. Dark box. B. Computer. C. Microscope. D. Camera. E. Multiposition stage control. F. Light source.
Software
Gen5TM Reader Control and Data Analysis Software for Biotek microplate reader
Metamorph image acquisition software (Molecular Devices)
FIJI (Schindelin et al., 2012)
BRET analyzer FIJI plugin (Chastagnier et al., 2018) https://github.com/ychastagnier/BRET-Analyzer
Graphpad Prism 7 software (GraphPad Software Inc.)
Procedure
C2C12 cell line maintenance and seeding
Maintain cultured C2C12 cells in 100 mm culture plates in a 37 °C incubator with 5% CO2 and split cells when confluence reaches 80–90%. Use regular culture medium containing phenol red (see Recipes section).
Two to three days before BRET measurement, trypsinize, collect and seed cells according to the plate format (see below). Typically, 150 × 103 cells/well in a 6-well plate before dispatching in white 96-well plates for BRET in cell populations, and 10 × 103 to 15 × 104 in an IbiTreat µ-slide for BRET imaging.
C2C12 transfection
For BRET in cell populations, we routinely transfect with JETPEI, but any other efficient approaches can be used.
Day 1, seed 150 × 103 cells/well in 6-well plates.
Day 2, the following amount and volumes are given to transfect one well of a 6-well plate: In 1.5 ml microcentrifuge tube A: mix 50 ng of AIMTOR DNA with 100 µl of sterile NaCl (150 mM). Use a non-coding plasmid (pBluescript or other) to normalize the DNA amount to 1 µg.
In 1.5-ml microcentrifuge tube B, combine 4 µl of JETPEI transfection reagent (4 µl/μg of DNA) with 100 µl of sterile NaCl (150 mM). Vortex the tube for 10 s.
Add the JETPEI reagent mixture in microcentrifuge tube B dropwise into microcentrifuge A (containing the DNA mixture). Vortex the tube for 10 s and centrifuge briefly.
Incubate for 20 min at room temperature.
Add the content of the combined JETPEI-DNA mixture dropwise into the wells.
Centrifuge the 6-well plates for 5 min at 280 × g.
Incubate cells at 37 °C in 5% CO2 for 24 h.
Day 3:
Wash each well with 1 ml PBS.
Add 300 µl Trypsin/EDTA.
Incubate 5–10 min at 37 °C.
Add 1 ml complete medium without phenol red.
Resuspend cells with a 1-ml micropipette and collect them in a 15-ml tube.
Centrifuge cells for 5 min (500 × g).
Aspirate supernatant and resuspend cells in 2.5 ml of culture medium
Dispatch the cells in a 96-well plate: 100 µl/well in triplicate wells × the number of conditions for each transfection.
Incubate for 24–48 h before proceeding to plate BRET reading.
For single-cell BRET imaging via microscopy
Note: We use the HelixIN transfection protocol, but any other efficient transfection protocol can be used.
Day 1: Seed 10 × 103 to 15 × 103 cells in each well of an 8-well untreated or polylysine-treated ibidi slide.
Thaw HelixIN and HB100X stock reagents and vortex them briefly.
Dilute HelixIN reagent 5× in Milli-Q sterile water (1 µl of HelixIN + 4 µl of H2O).
For a single well of the 8-well ibidi slide, combine in a 1.5-ml microcentrifuge tube: 1 µl (50 ng) of AIMTOR DNA (stock 50 ng/µl), 1.5 µl (250 ng) of non-coding plasmid (stock 200 ng/µl), and 30 µl of serum-free DMEM.
Add 3.5 µl of diluted HelixIN.
Mix gently, but do not vortex!
Wait 30 min at room temperature.
Add the total 36-µl dropwise into one well of an 8-well ibidi slide.
Add 3.3 µl of HB100×.
Incubate the cells for 24–48 h in a 37 °C incubator with 5% CO2.
Proceed to BRET imaging via microscopy (see below).
BRET monitoring in cell populations
Turn on the Synergy 2 (or equivalent) plate reader and open the Gen5TM Reader software.
Program the protocol for BRET plate reading:
Select wells to be read in the “plate read mode” allowing several readings of the plate (lasting 20 min to 1 h max).
For each plate reading, program two successive 1-s readings at wavelengths of 485 nm and 530 nm, allowing the calculation of BRET intensity based on the ratio of the light unit emission at 530 nm to the light unit emission at 485 nm.
Set up the dynamic range and other parameters according to the plate reader.
Prepare 10× furimazine [200 µM] in a 1.5-ml tube by diluting furimazine stock in BRET reading medium.
Remove the 96-well plate containing cells expressing AIMTOR from the CO2 incubator.
Aspirate culture medium and replace with 90 µl of fresh BRET reading medium (phenol red free) immediately before plate reading.
Add 10 µl of furimazine (200 µM) diluted in BRET reading medium (20 µM final concentration) to each well containing cells using a multichannel pipette or a repipetor (Eppendorf).
Perform plate reading for a maximum of 1 h (allowing successive readings of each well depending on the number of wells to be read and the speed of the plate reader).
Export an Excel file with the raw experimental data showing the light unit emission at 530 nm and 485 nm for each well over time.
BRET imaging via microscopy
Turn on the microscope system and open the Metamorph image acquisition software (Figure 1). Wait approximately 1 h until the camera temperature has reached -80 °C.
During that time, take one 8-well µ-slide out of the incubator and remove the culture medium from the wells you want to image without drying the cells. Wash the cells once with 300 µl of PBS and fill each well with 200 µl of BRET experiment culture medium (see Recipes section). Change the medium at least 30 min before imaging because this procedure can affect mTOR signaling.
Place the dish on the microscope stage and secure it so it cannot move.
Collect a background dark image using the same experimental conditions that you will use for imaging. For your first acquisition, you will have to perform this step after BRET recording, and without the dish.
Focus the image on a field of cells using epifluorescence illumination to detect Y-PET and collect an image to capture the localization of the AIMTOR biosensor (Figure 2A).
If your setup is equipped with a multi-position stage, save the position and repeat this step for the number of positions you wish to use. Note that the more positions you save, the longer the delay will be between consecutive time points for each field.
Prepare a fresh mixture consisting of 3 µl of furimazine with 57 µl of BRET experiment culture medium. Carefully add 50 µl of this mixture on top of the cells to achieve a final concentration of 50 µM furimazine. Wait for 1 min to allow furimazine to diffuse in the well.
Make sure to turn off all microscope illumination and room lighting. Close the dark box.
Image channels at wavelengths of 460 nm and 530 nm (determine the appropriate exposure time so that none of the channels are over-exposed).
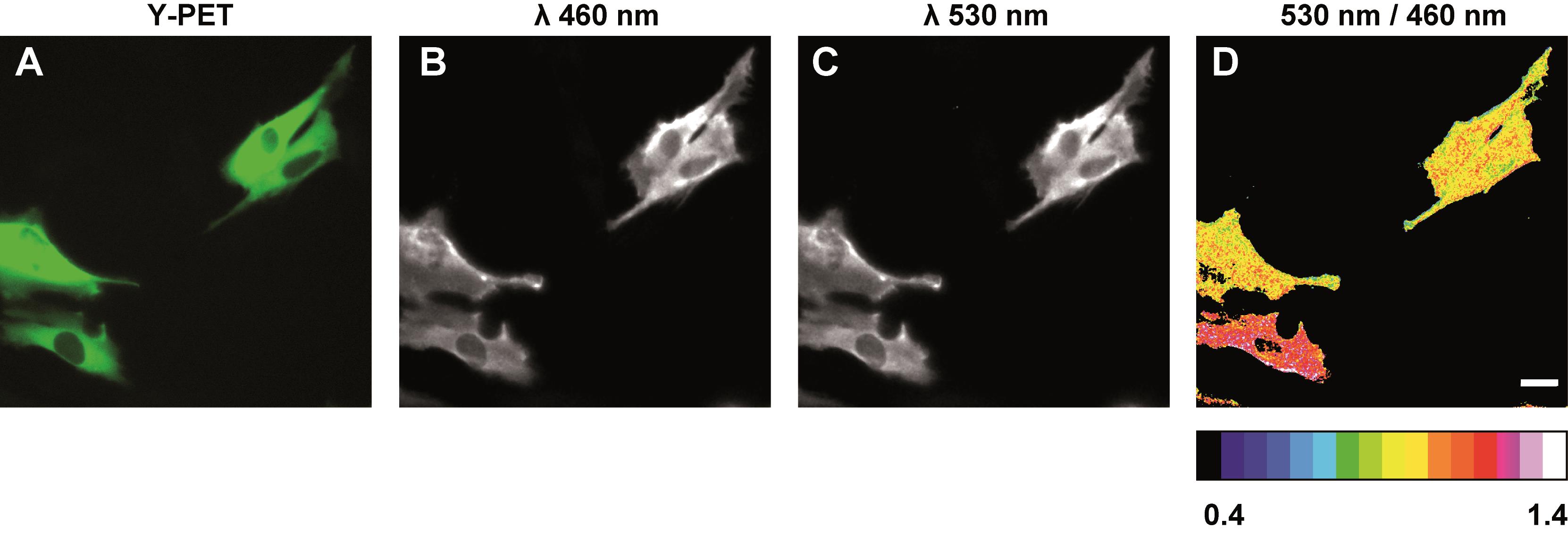
Figure 2. BRET imaging of mTORC1 activity. A. Example of reporter expression detected in AIMTOR-transfected C2C12 cells by Y-PET expression. NLuc emission (B, Em 460) and Y-PET emission (C, Em 530) initiated by the non-radiative transfer of energy from NLuc after furimazine addition. D. Pixel-by-pixel division of the Em 530 and Em 460 images gives rise to the pseudo-colored BRET image (Em 530/Em 460). Scale bar: 20 µm.
Data analysis
Data analysis of BRET in cell populations
Export an Excel file with raw data for each well showing the light unit emission at wavelengths of 530 nm and 485 nm over 20 to 60 min (several successive readings of the plate) (Figure 3, Table A).
Exclude BRET measurements when the light emitted by the donor is out of the linear range of the fluorimeter or when NanoLuc is saturated or emits less than 6000 light units at Em485nm. This minimum value was determined using a Biotek Synergy 2 reader by serial dilution of cells expressing NanoLuc to verify the linearity/accuracy of the reader and to determine the lowest threshold value allowing accurate BRET measurement. This threshold value will depend on the characteristics of each plate reader (reader setting sensitivity and range).
Calculate the BRET intensity for each well and each time point (BRET ratio = Em530/Em485).
Calculate the mean BRET intensity ratio for triplicate wells at each reading time point (Figure 3; Table A).
Calculate the BRET intensity integrated during 20 or 60 min and average the BRET intensity of a given condition for a minimum of 3 independent transfections (each loaded in triplicate) (Figure 3, Table B). Determine the AIMTOR BRET intensity over time (Figure 3C).
Pool the mean BRET intensity data in GraphPad Software and perform the appropriate statistical analysis.
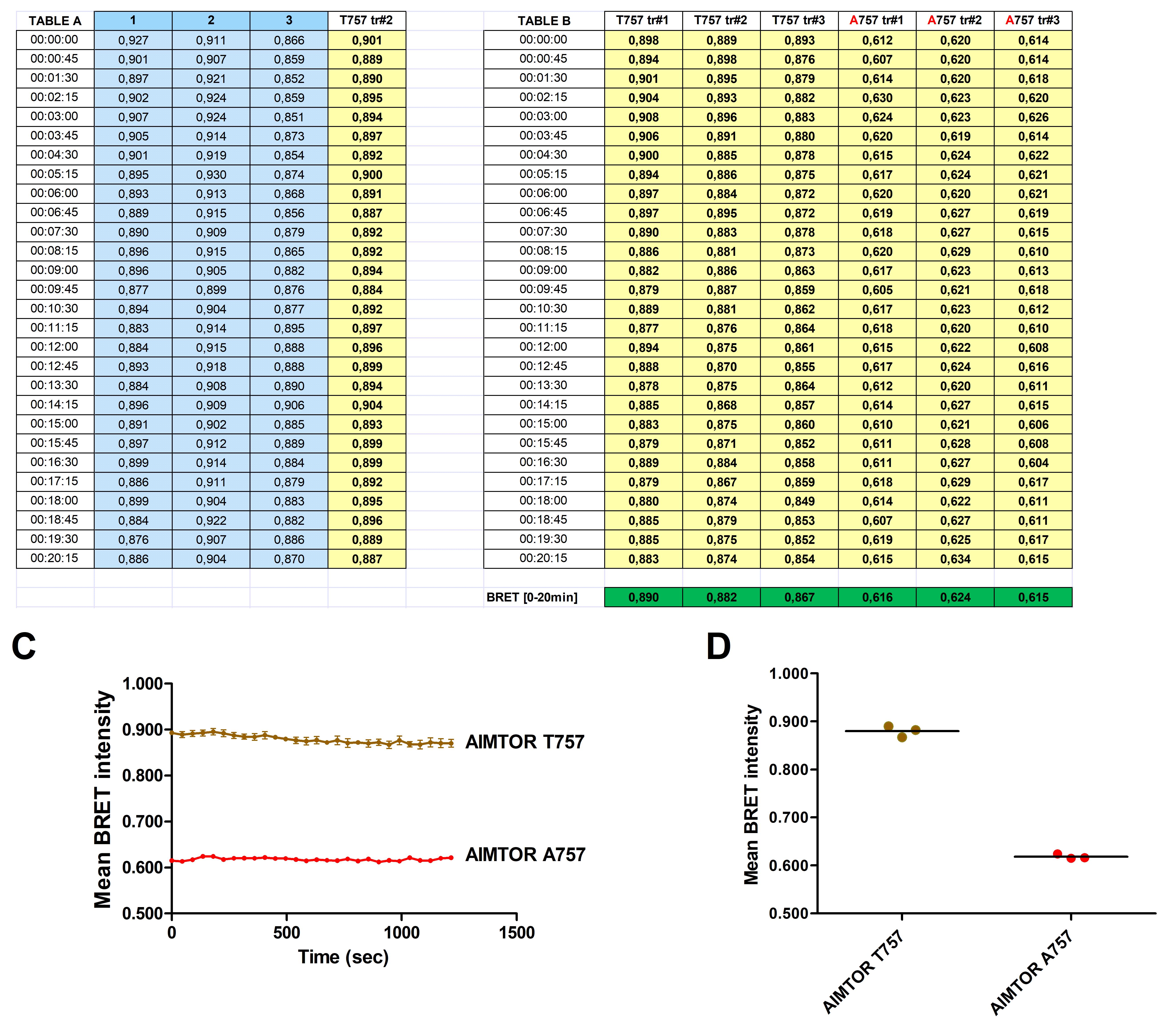
Figure 3. Example of a typical AIMTOR BRET analysis in a population of C2C12 cells using a plate reader. Table A: BRET intensity (BRET Ratio Em530/Em485) over time in triplicate wells of one single transfection with AIMTOR (blue colored columns) and the resulting mean BRET ratio of triplicate wells (yellow-colored columns). Table B: BRET intensity over time of 3 independent transfections for AIMTOR (T757) and 3 independent transfections for its nonphosphorylatable control mutant (A757). Each yellow column represents the mean BRET intensity over time for a single transfection calculated from triplicate wells (not shown) and the bottom green lane shows the mean BRET intensity of each transfection integrated over a 20-min reading. Graphs C and D show the BRET intensity of 3 independent C2C12 transfections of AIMTOR T757 and its nonphosphorylatable A757 mutant; BRET intensity during the 20-min monitoring period (C) and mean BRET intensity integrated over 20 min of plate reading (D).
Data analysis of BRET imaging via microscopy
Exclude cells in which the NanoLuc signal saturates the camera.
Repeat the experiment in at least three different batches of cells and collect at least ten fields per condition. Increase these parameters depending on the variability of your experiment.
If you are evaluating kinetics, save images as a stack for each wavelength and position acquisition to facilitate further analysis (step 4). If not, ignore this step.
Use our previously published open source plugin for Fiji (Chastagnier et al., 2018) to perform analysis on each cell/subcellular compartment (https://github.com/ychastagnier/BRET-Analyzer). Briefly, it performs 4 key steps:
Image background subtraction.
Image alignment over time.
Composite thresholding of the image.
Pixel by pixel division of the image and distribution of the ratio intensity on a pseudocolor scale.
Pool the mean BRET intensity data in GraphPad Software and perform the appropriate statistical analysis.
Notes
Facts and tips to consider to select the best emission filters for your experiment:
For optimal light detection, the emission filter should be centered on the dye’s emission peak. However, depending on the bandwidths of the emission filter, sufficient light can be collected for BRET experiments even if the emission filter is not perfectly centered on the emission peak. For example, in this study, we used [485/20nm] and [450/70nm] emission filters to monitor nanoluciferase emission using the microplate reader and microscope, respectively. Both filters are not precisely centered at 460 nm, the peak of emission wavelength of nanoluciferase, but they allowed us to collect a sufficient signal. However, for these reasons, the absolute values of the BRET ratios calculated based on these two approaches cannot be formally compared.
Facts and tips to consider when selecting an AIMTOR BRET reading medium:
Comparing BRET intensity data can be meaningless when BRET recordings are not performed with the same reading medium and conditions, as mTOR activity (and AIMTOR) is potentially sensitive to many parameters and monitoring conditions, such as cell confluency and shaking of the plate. It is, therefore, crucial to use a unique reading buffer and protocol when comparing various experimental conditions. As mTOR kinase activation depends notably on amino acids, serum and other compounds, the presence/lack of one or more of these compounds can alter BRET intensity and its stability during monitoring periods lasting 20 to 60 min. For AIMTOR BRET intensity recording, plate reading can be performed in several types of medium, providing that the selected medium is devoid of phenol red to eliminate any detrimental interference with BRET monitoring. However, a simple PBS buffer containing 0.1% glucose can also be used for AIMTOR BRET cell population reading. Interestingly, we noticed that adherent cells put in suspension (after washing in PBS, trypsinization, cell collection by centrifugation and resuspension in BRET reading buffer) can also be used to measure the BRET intensity of AIMTOR. However, in this case, the BRET intensity of the resuspended cells is lower than that of the same cell type if it is maintained in an adherent state during BRET reading. Cell confluency is another parameter that can alter mTOR activity and should therefore be considered carefully when comparing conditions. Finally, shaking of the plate just after furimazine addition may also have an impact on AIMTOR BRET intensity, and we do not recommend the use of a shaking step in the plate reading protocol.
Successive acquisitions can be performed over time to monitor mTOR dynamics (Bouquier et al., 2020).
Recipes
For C2C12 cell growth and maintenance
DMEM regular growth medium (contains phenol red)
DMEM (10% FBS + 1 mM Na pyruvate + 1/100 penicillin and streptomycin)
DMEM growth medium and reading medium
DMEM (without phenol red)
25 mM glucose
2 mM glutamine
10% FBS
1 mM Na pyruvate
1/100 penicillin and streptomycin
The culture medium is replaced by the BRET reading medium immediately before performing a plate reading or a BRET imaging experiment.
To control the possible pH drop that could occur during imaging, HEPES (15 mM, pH 7.4) is added to the BRET reading medium based on a DMEM formula without phenol red. We did not detect any significant differences when reading BRET in a cell population with a DMEM-based medium formula based on whether it did or did not contain HEPES.
PBS containing 0.1% glucose can also be used for cell population BRET reading (adherent or resuspended cells).
Acknowledgments
This work was supported by INRAE (VO) and by the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation programme (JP, grant agreement No. 646788), ANR Lanthslider (JP, ANR-17-CE11-0046).
Competing interests
The authors declare no competing interests.
References
- Bouquier, N., Moutin, E., Tintignac, L. A., Reverbel, A., Jublanc, E., Sinnreich, M., Chastagnier, Y., Averous, J., Fafournoux, P., Verpelli, C., Boeckers, T., Carnac, G., Perroy, J. and Ollendorff, V. (2020). AIMTOR, a BRET biosensor for live imaging, reveals subcellular mTOR signaling and dysfunctions. BMC Biol 18(1): 81.
- Chastagnier, Y., Moutin, E., Hemonnot, A. L. and Perroy, J. (2018). Image processing for bioluminescence resonance energy transfer measurement-BRET-analyzer.Front Comput Neurosci 11: 118.
- Coulon, V., Audet, M., Homburger, V., Bockaert, J., Fagni, L., Bouvier, M. and Perroy, J. (2008). Subcellular imaging of dynamic protein interactions by bioluminescence resonance energy transfer. Biophys J 94(3): 1001-1009.
- Kim, J. and Grailhe, R. (2016). Nanoluciferase signal brightness using furimazine substrates opens bioluminescence resonance energy transfer to widefield microscopy. Cytometry A 89(8): 742-746.
- Perroy J. (2010) Subcellular dynamic imaging of protein–protein interactions in live cells by bioluminescence resonance energy transfer. In: Papkovsky, D. (Ed.). Live Cell Imaging. Methods in Molecular Biology (Methods and Protocols), vol 591. Humana Press.
- Saxton, R. A. and Sabatini, D. M. (2017). mTOR Signaling in Growth, Metabolism, and Disease. Cell 168(6): 960-976.
- Schindelin, J., Arganda-Carreras, I., Frise, E., Kaynig, V., Longair, M., Pietzsch, T., Preibisch, S., Rueden, C., Saalfeld, S., Schmid, B., Tinevez, J. Y., White, D. J., Hartenstein, V., Eliceiri, K., Tomancak, P. and Cardona, A. (2012). Fiji: an open-source platform for biological-image analysis. Nat Methods 9(7): 676-682.
Article Information
Copyright
© 2021 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Bouquier, N., Moutin, E., Perroy, J. and Ollendorff, V. (2021). AIMTOR, a BRET Biosensor for Live Recording of mTOR Activity in Cell Populations and Single Cells. Bio-protocol 11(8): e3989. DOI: 10.21769/BioProtoc.3989.
Category
Cell Biology > Cell signaling > Phosphorylation
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link