- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Ex vivo Assessment of Mitochondrial Function in Human Peripheral Blood Mononuclear Cells Using XF Analyzer
Published: Vol 11, Iss 7, Apr 5, 2021 DOI: 10.21769/BioProtoc.3980 Views: 5194
Reviewed by: Luis Alberto Sánchez VargasAlexandros C KokotosAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
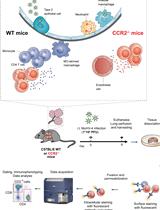
Functional Phenotyping of Lung Mouse CD4+ T Cells Using Multiparametric Flow Cytometry Analysis
Céline M. Maquet [...] Bénédicte D. Machiels
Sep 20, 2023 4230 Views
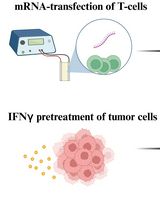
T-Cell-Based Platform for Functional Screening of T-Cell Receptors Identified in Single-Cell RNA Sequencing Data Sets of Tumor-Infiltrating T-Cells
Aaron Rodriguez Ehrenfried [...] Rienk Offringa
Apr 20, 2024 6423 Views

Novel Experimental Approach to Investigate Immune Control of Vascular Function: Co-culture of Murine Aortas With T Lymphocytes or Macrophages
Taylor C. Kress [...] Eric J. Belin de Chantemèle
Sep 5, 2025 3576 Views
Abstract
Cellular health and function, as we know today, depend on a large extent on mitochondrial function. The essential function of mitochondria is the energy production, more precisely ATP production, via oxidative phosphorylation. Mitochondrial energy production parameters therefore represent important biomarkers. Studies on human cells have mainly been performed on in vitro cell cultures. However, peripheral blood mononuclear cells (PBMCs) are particularly suitable for such examinations. That’s why this protocol describes a method to measure key parameters of mitochondrial function in freshly isolated PBMCs with the latest technology, the XF Analyzer. For this ex vivo approach PBMCs are first isolated out of human anticoagulated blood. Next, they are attached to the surface of special microplates pre-coated with Poly-D-Lysine. During the subsequent measurement of oxygen consumption rate (OCR) as well as extracellular acidification rate (ECAR) the stress reagents oligomycin, carbonyl cyanide 4-(trifluoromethoxy)phenylhydrazone (FCCP), rotenone and antimycin A are injected. Several mitochondrial parameters can be calculated from the results obtained. The application of this protocol allows the analysis of various influences, such as pharmaceuticals or environmental factors, on human cells.
Keywords: Human peripheral blood mononuclear cells (PBMCs)Background
Mitochondria play a critical role in maintaining normal cellular function. It is now common knowledge that they not only produce ATP via oxidative phosphorylation but, for example, are also involved in the metabolism of amino acids, lipids and nucleotides, diverse signaling and redox processes as well as quality control and degradation processes including mitophagy and apoptosis (Pfanner et al., 2019). However, mitochondria represent the major site of ATP synthesis in normal cells (Akbari et al., 2019). For this purpose, an electrochemical proton gradient is generated across the mitochondrial inner membrane through the multi-subunit enzyme complexes I–IV. This proton gradient is used by the ATP synthase, also known as complex V, to turn ADP into ATP (Chaban et al., 2014).
The process of oxidative phosphorylation is associated with the reduction of oxygen to water. Accordingly, the oxygen consumption rate of cells can be used for assessing mitochondrial function (Smolina et al., 2017). This principle is the basis of Seahorse XF Analyzers (Agilent Technologies). They provide the possibility to measure not only oxygen consumption rate (OCR), but also the rate of extracellular acidification (ECAR), which is a key indicator of glycolysis. The realtime measurements are carried out in multi-well plates which are provided with solid-state sensors consisting of two fluorophores. One is quenched by oxygen (O2) and the other one is sensitive to pH-value changes. The fluorophores are excited via light-emitting fiber optic bundles, which subsequently detect the fluorescence changes as a result of oxygen consumption or extracellular acidification (Plitzko and Loesgen, 2018). Furthermore, XF Analyzers enable up to four different injections per well during the measurement. All the properties mentioned constitute a significant advantage over the conventionally used clark-type oxygen electrodes for determining oxygen consumption.
Peripheral blood mononuclear cells (PBMCs) as sample in a XF Analyzer implies that the cells have to be attached to the surface of the microplates. Most commonly, this immobilization is done by means of protein solutions such as Cell-TakTM (Jones et al., 2015; Traba et al., 2016; Lee et al., 2019) or Poly-D-Lysine (Hartman et al., 2014; Nicholas et al., 2017; Thaventhiran et al., 2019). We examined both Cell-TakTM and Poly-D-Lysine and considered Poly-D-Lysine as most suitable coating method. Since we conducted the Agilent Seahorse XF Cell Mito Stress Test, optimal concentrations of the injected compounds oligomycin, FCCP, antimycin A and rotenone had to be tested as well. A typical curve of a Mito Stress Test is shown in Figure 1. Since oligomycin is an inhibitor of ATP synthase, OCR decreases after its injection. In contrast, OCR increases sharply after FCCP injection, which is an uncoupler of oxidative phosphorylation. The last injection of antimycin A and rotenone, again leads to a decline of OCR, as these two compounds inhibit complex III respectively I of the electron transport chain. The resulting curve is used to calculate various parameters of mitochondrial function (see Figure 1).
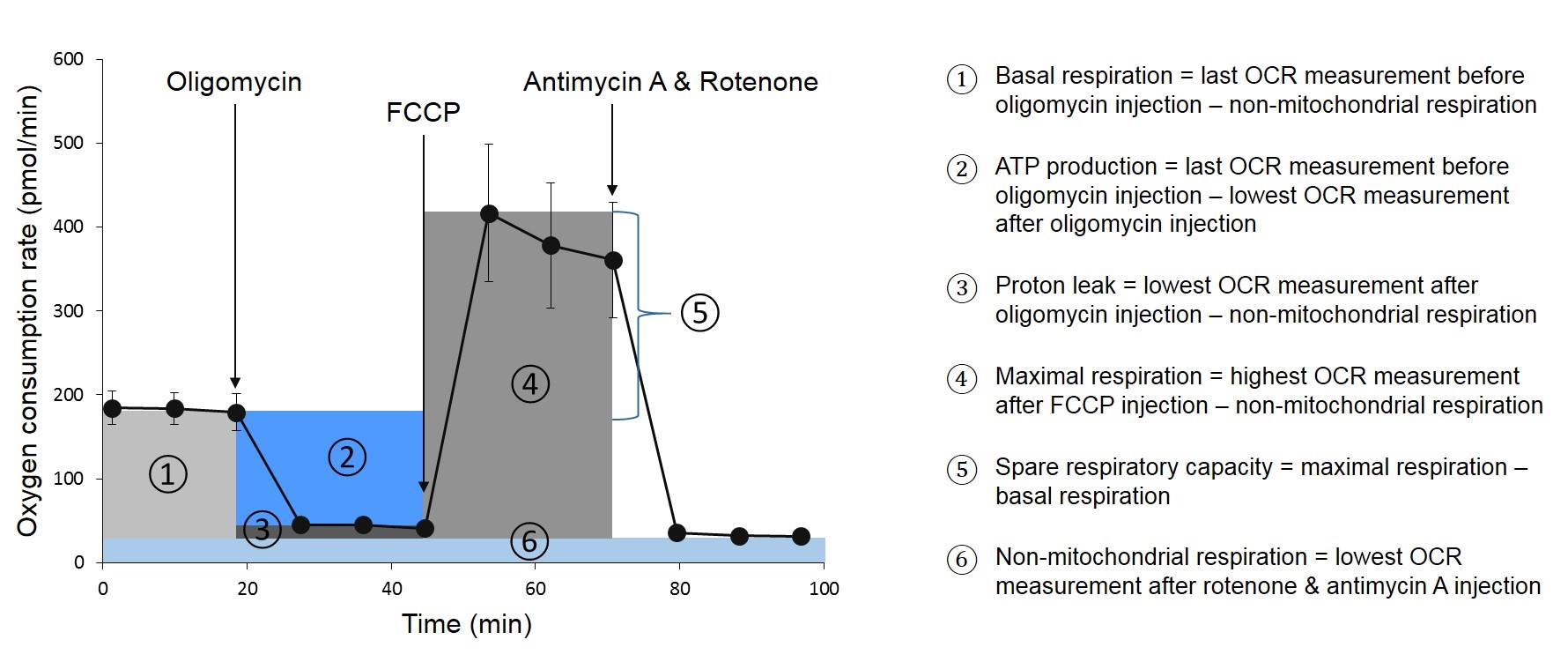
Figure 1. Assessment of mitochondrial respiration parameters by means of Agilent Seahorse XF Cell Mito Stress Test. On the left side a typical course of an OCR measurement with injections of oligomycin after the third, FCCP after the sixth and antimycin A together with rotenone after the ninth measuring point is shown. From this curve, various parameters can be calculated, which are marked in different colors. The calculations of the parameters are shown on the right side.
The protocol described in detail hereafter has been applied to examine the influence of 12 days of in vivo caloric reduction in humans. A significant increase of mitochondrial respiratory parameters could be detected in PBMCs of a subgroup of the test persons (Schöller-Mann et al., 2020). Besides such ex vivo studies the protocol can be used to screen any soluble substance, including pharmaceuticals, dietary supplements or contaminants, due to their in vitro effects on PBMCs.
Materials and Reagents
15 ml, 50 ml screw cap tubes (SARSTEDT, catalog numbers: 62.554.502 ; 62.547.254 )
1.5 ml, 2 ml reaction tubes (SARSTEDT, catalog numbers: 72.706 ; 72.695.500 )
20 µl, 200 µl, 1,000 µl pipette tips (SARSTEDT, catalog numbers: 70.1116 ; 70.760.002 ; 70.762 )
5 ml, 10 ml, 25 ml serological pipettes (SARSTEDT, catalog numbers: 86.1253.001 ; 86.1254.001 ; 86.1685.001 )
50 ml LeucosepTM tubes (Greiner Bio-One GmbH, catalog number: 227290 )
Seahorse XF24 FluxPak containing XF24 sensor cartridges, XF24 cell culture microplates and XF Calibrant Solution (Agilent Technologies, catalog number: 100850-001 )
Human venous blood, EDTA-anticoagulated – S-Monovette® 7.5 ml K3E (SARSTEDT, catalog number: 01.1605.001 )
Ficoll-PaqueTM PLUS (GE Healthcare, catalog number: 17-1440-03 )
Poly-D-Lysine solution, 1.0 mg/ml (Merck Millipore, catalog number: A-003-E , storage temp. -20 °C)
Trypan Blue solution 0.4% (Sigma-Aldrich, catalog number: 93595 )
Dulbecco’s Modified Eagle’s Medium (DMEM), high glucose (Sigma-Aldrich, catalog number: D7777 , storage temp. 4 °C)
Dimethyl sulfoxide (DMSO) (Carl Roth, catalog number: A994 )
Oligomycin from Streptomyces diastatochromogenes (Sigma-Aldrich, catalog number: O4876 , storage temp. -20 °C; mixture of isomers A, B, and C), 2.5 mM in DMSO
Carbonyl cyanide 4-(trifluoromethoxy) phenylhydrazone (FCCP; Sigma-Aldrich, catalog number: C2920 , storage temp. 4 °C), 2.5 mM in DMSO
Rotenone (Sigma-Aldrich, catalog number: R8875 ), 2.5 mM in DMSO
Antimycin A from Streptomyces sp. (Sigma-Aldrich, catalog number: A8674 , storage temp. -20 °C), 2.5 mM in DMSO
NaCl (Carl Roth, catalog number: 3957 )
NaOH (Carl Roth, catalog number: 6771 )
KCl (Carl Roth, catalog number: 6781 )
Na2HPO4·12H2O (Carl Roth, catalog number: N350 )
KH2PO4 (Carl Roth, catalog number: 3904 )
1× PBS (see Recipes)
0.9% NaCl solution (see Recipes)
10 M NaOH solution (see Recipes)
Poly-D-Lysine working solution (50 µg/ml) (see Recipes)
2.5 mM oligomycin solution (see Recipes)
2.5 mM FCCP solution (see Recipes)
0.1 M rotenone solution (see Recipes)
2.5 mM rotenone solution (see Recipes)
2.5 mM antimycin A solution (see Recipes)
Assay medium (see Recipes)
Equipment
Pipettes: Eppendorf Research® Plus 10 μl, 20 µl, 200 µl, 1,000 µl (Eppendorf, catalog numbers: 3123000020 ; 3123000039 ; 3123000055 ; 3123000063 )
Pipetting aid: PipetBoy acu 2 (Integra Biosciences, catalog number: 155 000)
Neubauer counting chamber improved (Carl Roth, catalog number: PC72.1 )
Inverted microscope: Primovert (Carl Zeiss, catalog number: 415510-1100-000 )
Water bath (GFL, catalog number: 1003 )
Incubator without CO2 (GFL, catalog number: 4010 )
Swing out rotor centrifuge: 5804 R (Eppendorf, catalog numbers: 5805000010 ; 5804709004 )
Water purification system: ELGA® PURELAB flex 3 (Veolia, catalog number: PF3XXXXM1 )
pH meter: FE20 FiveEasyTM (Mettler Toledo, cataog number: 30266626 )
Seahorse XF24 Extracellular Flux Analyzer (Agilent Technologies, catalog number: 100737-100 )
Software
Wave Controller Software (Agilent Technologies, version 1.8.1.1)
Optional: Wave Desktop Software (Agilent Technologies)
Procedure
The experimental procedure for the detection of mitochondrial function in human PBMCs using a XF Analyzer is shown schematically in Figure 2. This is followed by a detailed description of the individual work steps.
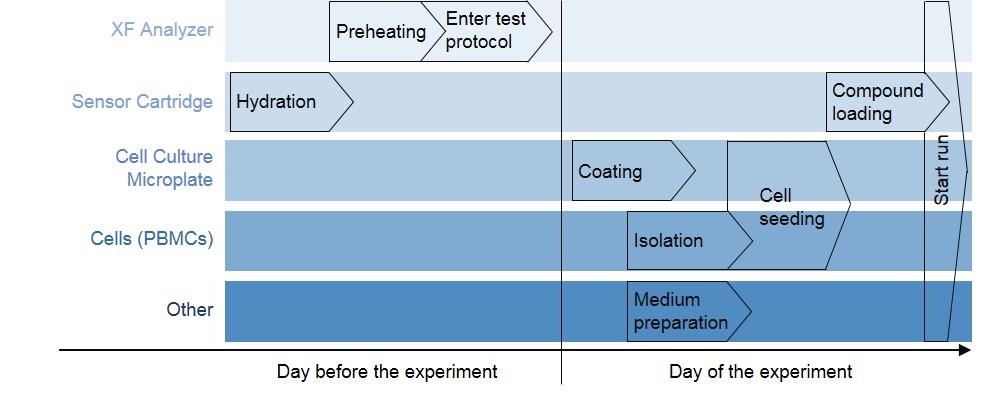
Figure 2. Workflow diagram of mitochondrial function measurement in human PBMCs using a XF Analyzer
Day before the experiment
Hydration of XF24 sensor cartridge (consisting of the cartridge lid, the sensor cartridge itself and the utility plate; see Figure 3B and 3C):
Add 1 ml XF Calibrant Solution into each well of the utility plate.
Remove air bubbles that might arise at the sensors with a pipette tip.
Incubate the assembled plate in a non-CO2 37 °C incubator overnight.
Switch on the XF24 Analyzer (see Figure 3A) and let it warm up to 37 °C overnight.

Figure 3. XF24 Analyzer (A) and XF24 sensor cartridge (B and C). The sensor cartridge, including markings of the sensors, is shown from above (B) as well as from below (C). In addition, the injection ports are marked (B) and the utility plate is depicted (C).Programming of measurement protocol on the XF Analyzer:
Equilibration: 30 min
Measurement period 1:
Mixing: 4 min
Waiting: 2 min
Measuring: 3 min
→ 3 repetitions
Injection of port A (50 µl of 7.5 µM oligomycin; final concentration 0.75 µM)
Measurement period 2:
Mixing: 4 min
Waiting: 2 min
Measuring: 3 min
→ 3 repetitions
Injection of port B (55 µl of 10 µM FCCP; final concentration 1 µM)
Measurement period 3:
Mixing: 4 min
Waiting: 2 min
Measuring: 3 min
→ 3 repetitions
Injection of port C (60 µl of 16.7 µM rotenone/antimycin A; final concentration 1.67 µM)
Measurement period 4:
Mixing: 4 min
Waiting: 2 min
Measuring: 3 min
→ 3 repetitions
Day of the experiment
Coating of XF24 cell culture microplate:
Add 30 µl of Poly-D-Lysine working solution (50 µg/ml) into each well and incubate the plate for 1 h at RT.
Discard the supernatant, wash each well with 300 µl ddH2O and let the plate air dry under sterile conditions (approximately 30 min).
Isolation of PBMCs (continue with this step during microplate coating; see Figure 4):
Add 15 ml of Ficoll-PaqueTM PLUS (RT) into a 50 ml LeucosepTM tube and centrifugate at 1,000 × g for 30 s at RT.
Dilute an appropriate volume of anticoagulated human blood at a ratio of 1:2 with 0.9% NaCl solution (at least 7.5 ml and at most 15 ml).
Transfer the resulting volume (at least 15 ml and at most 30 ml) into the prepared LeucosepTM tube and centrifuge at 1,000 × g for 10 min at RT in a swing out rotor with brakes switched off.
Harvest the PBMC fraction which appears as white layer between the plasma and the Ficoll above the porous barrier of the LeucosepTM tube [for detailed information see manufacturer's instructions or refer to Matt and Bergemann, (2019)]. To do this use a 1,000 µl Eppendorf pipette and collect the PBMCs in a 50 ml screw cap tube.
Wash the cells once with 10 ml PBS and centrifuge at 250 × g for 10 min.
Repeat the washing step twice with 5 ml PBS each.
Determine the number of cells using a Neubauer counting chamber improved.

Figure 4. Isolation of PBMCs with a LeucosepTM tube. On the left side, the tube is filled with Ficoll-PaqueTM below and anticoagulated and diluted blood above the porous barrier. On the right side, after density gradient centrifugation, the tube shows the following layers from top to bottom: plasma – PBMCs – Ficoll-PaqueTM – porous barrier – Ficoll-PaqueTM – erythrocytes and granulocytes.Prepare the assay medium and preheat at 37 °C before use.
Seeding PBMCs onto pre-coated XF24 cell culture microplate:
Note: 4 to 5 wells per condition are recommended. The wells A1, B4, C3 and D6 are normally used as background wells (without cells). An exemplary plate layout as well as an image of PBMCs seeded on Poly-D-Lysine is shown in Figure 5.
Take 5 × 105 of the isolated PBMCs per well and centrifuge at 250 × g for 5 min.
Resuspend the cell pellet in 100 µl assay medium per well.
Add 100 µl of cell suspension per well.
Fill the background wells with 450 µl assay medium.
Centrifuge the plate at 250 × g for 1 min in a swing out rotor with brakes switched off.
Add 350 µl assay medium to the wells with cells (final volume in each well: 450 µl).
Incubate the plate at 37 °C without CO2 until measurement (on average about 1 h).
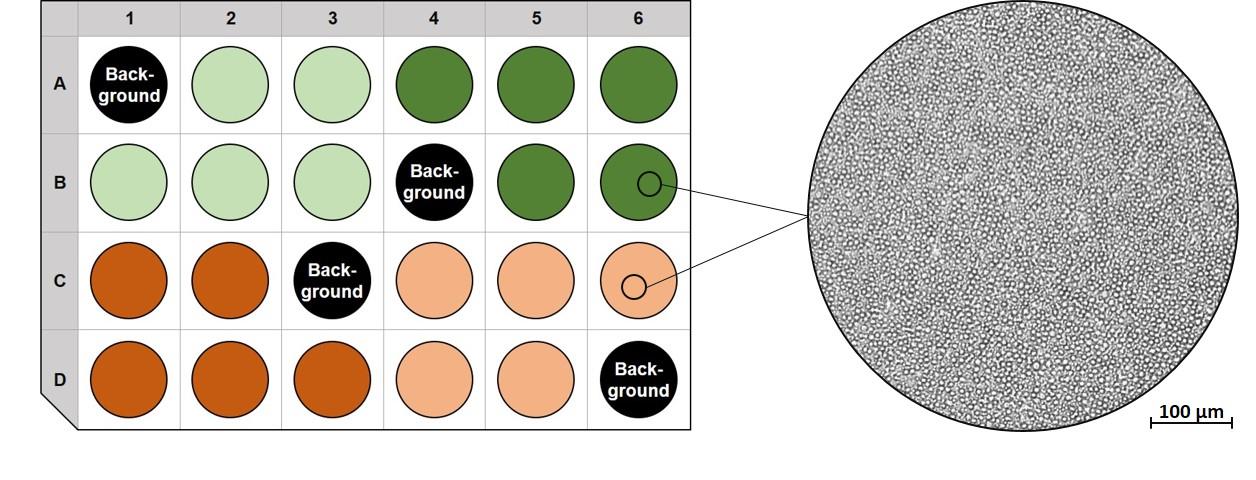
Figure 5. Exemplary plate layout of an XF24 cell culture microplate (left side) and a bright field microscopy image of PBMCs seeded on a Poly-D-Lysine coated XF24 cell culture microplate (right side). Left side: The wells A1, B4, C3 and D6 are used as background wells (without cells). One color (light green, dark green, dark orange, light orange) represents one condition each. There are 5 wells per condition. Right side: The image shows PBMCs in 100× magnification.
Loading of XF24 sensor cartridge with compounds:
Prepare compounds as follows:
Dilute 4.86 µl of 2.5 mM oligomycin solution in 1,615 µl assay medium (results in 7.5 µM).
Dilute 6.48 µl of 2.5 mM FCCP solution in 1,614 µl assay medium (results in 10 µM).
Dilute 10.82 µl of 2.5 mM rotenone solution and 10.82 µl of 2.5 mM antimycin A solution in 1,598 µl assay medium (results in 16.7 µM each).
Load the injection ports of the XF24 sensor cartridge as follows:
Note: Avoid the formation of air bubbles in the injection ports and handle the XF24 sensor cartridge with care after loading in order to prevent dripping down of the compounds.
Fill all ports A with 50 µl of 7.5 µM oligomycin.
Fill all ports B with 55 µl of 10 µM FCCP.
Fill all ports C with 60 µl of 16.7 µM rotenone/antimycin A.
Start the run in the XF Analyzer (follow the instructions of the software):
Place the XF24 sensor cartridge (without lid) in the XF Analyzer and start the calibration.
After completion of the calibration, replace the utility plate by the XF24 cell culture microplate and start measurement.
Normalization of measurement results to the number of cells:
Resuspend PBMCs in the final volume of each well (615 µl) by repeatedly pipetting up and down.
Determine the number of cells per well using a Neubauer counting chamber improved.
Data analysis
Divide the counted number of cells per well by 10,000 and enter the results in the Wave software under ‘Normalize’ in order to normalize the data (see Figure 6).
Figure 6. Screenshot of the Wave software with markings for normalization and data exportDefine outliers based on comparing the course of the OCR of single wells of one condition and exclude them from analysis.
Note: At least three replicates should be used for analysis.
Export the results to the Seahorse XF Cell Mito Stress Test Report Generator, which is linked to the Wave software (see Figure 6). This software tool automatically calculates and reports the parameters, which are analyzed by the Mito Stress Test.
Take the calculated values from the Report Generator and analyze them with an appropriate statistical software.
Representative Data
A typical result of an oxygen consumption rate measurement using a XF Analyzer and the stress reagents oligomycin, FCCP, antimycin A and rotenone (Mito Stress Test) is shown in Figure 7.

Figure 7. Oxygen consumption rate (OCR) of a Mito Stress Test of PBMCs measured with a XF Analyzer. The results of two different subjects are depicted not normalized (A) and normalized (B).
Notes
Since human blood samples are often available only once per condition, the measurement of a standard PBMC sample, which is stored deep-frozen, is recommended. This can be used for an additional normalization step.
Port D of the XF24 sensor cartridge is not used in the described experiment and therefore all ports D can be left blank. Of the ports A, B and C all 24 wells have to be filled in order to ensure an equal injection of the compounds.
Recipes
1× PBS
8 g NaCl
0.20 g KCl
2.88 g Na2HPO4·12H2O
1.24 g KH2PO4
ddH2O to 1 L, pH 7.4
0.9% NaCl solution
9 g NaCl
ddH2O to 1 L
10 M NaOH solution
40 g NaOH
ddH2O to 100 ml
Poly-D-Lysine working solution (50 µg/ml)
50 µl Poly-D-Lysine solution, 1.0 mg/ml
950 µl ddH2O
2.5 mM oligomycin solution
5 mg oligomycin
2.528 ml DMSO
2.5 mM FCCP solution
10 mg FCCP
15.737 ml DMSO
0.1 M rotenone solution
1 g rotenone
25.353 ml DMSO
2.5 mM rotenone solution
25 µl of 0.1 M rotenone solution
975 µl DMSO
2.5 mM antimycin A solution
25 mg antimycin A
18.529 ml DMSO
Assay medium
0.675 g Dulbecco’s Modified Eagle’s Medium, high glucose
ddH2O to 50 ml, pH 7.4
Acknowledgments
This work was funded by the Baden-Württemberg Ministry of Science, Research and Art via the “Cooperative Graduate School InViTe”. We like to thank Dr. med. Adrian Schulte and his team of the F. X. Mayr Bodensee Centre in Überlingen-Hödingen for providing the blood samples.
Competing interests
The authors declare no competing interests.
Ethics
The experiments were conducted in accordance with the declaration of Helsinki and approved by the ethics committee of the Landesärztekammer Baden-Württemberg, Germany. Furthermore, all volunteers were informed in advance and gave their written consent to the use of their blood samples.
References
- Akbari, M., Kirkwood, T. B. L. and Bohr, V. A. (2019). Mitochondria in the signaling pathways that control longevity and health span. Ageing Res Rev 54: 100940.
- Chaban, Y., Boekema, E. J. and Dudkina, N. V. (2014). Structures of mitochondrial oxidative phosphorylation supercomplexes and mechanisms for their stabilisation. Biochim Biophys Acta 1837(4): 418-426.
- Hartman, M. L., Shirihai, O. S., Holbrook, M., Xu, G., Kocherla, M., Shah, A., Fetterman, J. L., Kluge, M. A., Frame, A. A., Hamburg, N. M. and Vita, J. A. (2014). Relation of mitochondrial oxygen consumption in peripheral blood mononuclear cells to vascular function in type 2 diabetes mellitus. Vasc Med 19(1): 67-74.
- Jones, N., Piasecka, J., Bryant, A. H., Jones, R. H., Skibinski, D. O., Francis, N. J. and Thornton, C. A. (2015). Bioenergetic analysis of human peripheral blood mononuclear cells. Clin Exp Immunol 182(1): 69-80.
- Lee, H. T., Lin, C. S., Pan, S. C., Wu, T. H., Lee, C. S., Chang, D. M., Tsai, C. Y. and Wei, Y. H. (2019). Alterations of oxygen consumption and extracellular acidification rates by glutamine in PBMCs of SLE patients. Mitochondrion 44: 65-74.
- Matt, K. and Bergemann, J. (2019). Ex vivo Analysis of DNA Repair Capacity of Human Peripheral Blood Mononuclear Cells by a Modified Host Cell Reactivation Assay. Bio-protocol 9 (15): e3325.
- Nicholas, D., Proctor, E. A., Raval, F. M., Ip, B. C., Habib, C., Ritou, E., Grammatopoulos, T. N., Steenkamp, D., Dooms, H., Apovian, C. M., Lauffenburger, D. A. and Nikolajczyk, B. S. (2017). Advances in the quantification of mitochondrial function in primary human immune cells through extracellular flux analysis. PLoS One 12(2): e0170975.
- Pfanner, N., Warscheid, B. and Wiedemann, N. (2019). Mitochondrial proteins: from biogenesis to functional networks. Nat Rev Mol Cell Biol 20(5): 267-284.
- Plitzko, B. and Loesgen, S. (2018). Measurement of Oxygen Consumption Rate (OCR) and Extracellular Acidification Rate (ECAR) in Culture Cells for Assessment of the Energy Metabolism. Bio-protocol 8(10): e2850.
- Schöller-Mann, A., Matt, K., Schniertshauer, D., Hochecker, B. and Bergemann, J. (2020). 12 days of in vivo caloric reduction can improve important parameters of aging in humans. Mech Ageing Dev 188: 111238.
- Smolina, N., Bruton, J., Kostareva, A. and Sejersen, T. (2017). Assaying Mitochondrial Respiration as an Indicator of Cellular Metabolism and Fitness. Methods Mol Biol 1601: 79-87.
- Thaventhiran, T., Wong, W., Alghanem, A. F., Alhumeed, N., Aljasir, M. A., Ramsey, S., Sethu, S., Yeang, H. X. A., Chadwick, A. E., Cross, M., Webb, S. D., Djouhri, L., Ball, C., Stebbings, R. and Sathish, J. G. (2019). CD28 Superagonistic Activation of T Cells Induces a Tumor Cell-Like Metabolic Program. Monoclon Antib Immunodiagn Immunother 38(2): 60-69.
- Traba, J., Miozzo, P., Akkaya, B., Pierce, S. K. and Akkaya, M. (2016). An Optimized Protocol to Analyze Glycolysis and Mitochondrial Respiration in Lymphocytes. J Vis Exp(117). DOI: 10.3791/54918.
Article Information
Copyright
© 2021 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Meßmer, A. L., Matt, K. C., Hochecker, B. and Bergemann, J. (2021). Ex vivo Assessment of Mitochondrial Function in Human Peripheral Blood Mononuclear Cells Using XF Analyzer. Bio-protocol 11(7): e3980. DOI: 10.21769/BioProtoc.3980.
Category
Immunology > Immune cell function > Lymphocyte
Cell Biology > Cell-based analysis > Mitochondrial respiration
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









