- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Imaging Microtubules in vitro at High Resolution while Preserving their Structure
Published: Vol 11, Iss 7, Apr 5, 2021 DOI: 10.21769/BioProtoc.3968 Views: 3610
Reviewed by: Thirupugal GovindarajanOneil Girish BhalalaAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Sensitive and Adaptable Turn-On Maturation (ATOM) Fluorescent Biosensors for Detecting Subcellular Localization of Protein Targets in Cells
Harsimranjit Sekhon [...] Stewart N. Loh
Mar 20, 2025 2304 Views
Assay for Site-Specific Homologous Recombination Activity in Adherent Cells, Suspension Cells, and Tumor Tissues
Yuki Yoshino [...] Natsuko Chiba
Apr 5, 2025 2494 Views
SiMPull-POP: Quantification of Membrane Protein Assembly via Single Molecule Photobleaching
Ryan J. Schuck [...] Rajan Lamichhane
Jan 5, 2026 351 Views
Abstract
Microtubules (MT) are the most rigid component of the cytoskeleton. Nevertheless, they often appear highly curved in the cellular context and the mechanisms governing their overall shape are poorly understood. Currently, in vitro microtubule analysis relies primarily on electron microscopy for its high resolution and Total Internal Reflection Fluorescence (TIRF) microscopy for its ability to image live fluorescently-labelled microtubules and associated proteins. For three-dimensional analyses of microtubules with micrometer curvatures, we have developed an assay in which MTs are polymerized in vitro from MT seeds adhered to a glass slide in a manner similar to conventional TIRF microscopy protocols. Free fluorescent molecules are removed and the MTs are fixed by perfusion. The MTs can then be observed using a confocal microscope with an Airyscan module for higher resolution. This protocol allows the imaging of microtubules that have retained their original three-dimensional shape and is compatible with high-resolution immunofluorescence detection.
Keywords: MicrotubulesBackground
Microtubules (MT) are polymers made by the combination of the heterodimers α- and β-tubulins and are a major component of the cell cytoskeleton. They are involved in fundamental mechanisms of cell function such as mitosis, intracellular transport, cytokinesis and maintenance of cell shape (Akhmanova and Steinmetz, 2015). Although inherently very rigid, MTs often appear curved in cells and few proteins have been described to bend microtubules (Brangwynne et al., 2006; Bechstedt et al., 2014; Leung et al., 2020; Cuveillier et al., 2020). Since the early 1970s (Weisenberg, 1972), the study of MTs in vitro has led to a better understanding of the molecular mechanism involved in the formation and dynamics of MTs. However, a detailed analysis of the shape of the microtubules remains technically difficult. Two main approaches are currently used: electron microscopy for the very detailed images obtained (up to a separation limit of a few Angstrom) (Alushin et al., 2014; Harris, 2015), and TIRF microscopy which allows the live observation of dynamic microtubules using fluorescently-labelled molecules (separation limit of about 200 nm) (Al-Bassam, 2014). However, these techniques are not suitable for the complete observation of microtubules with large three-dimensional curvatures such as the helical shape they adopt in the presence of MAP6 (Cuveillier et al., 2020). In order to obtain more clues on the structure of the MTs, we developed an assay that combines the use of fluorescent proteins and high-resolution imaging by Airyscan confocal microscopy (achievable resolution down to 120 nm in XY), while preserving the original shape of the MTs which is very sensitive to manipulation. This protocol has the advantage of avoiding unusual equipment and material. Moreover, it can be adapted to different super-resolution techniques such as expansion microscopy or stimulated-emission-depletion (STED) microscopy which would allow to increase the resolution up to 10 times (Blom and Brismar, 2014) and reach a separation limit of 50-20 nm.
Materials and Reagents
Tubulin, Atto-565 labelled tubulin, and biotinylated tubulin are prepared as already fully described in Ramirez-Rios et al. (2017). Store up to 1 year in liquid nitrogen
Cover glasses 26 × 76 mm #1 (VWR, catalog number: 630-2910 )
Double-face precut tape 70 µm thick, 3 mm wide (LIMA Company, catalog number: 0000P70PC3003 )
Siligum wax plate (VWR, MODU140013 )
1.5 ml Eppendorf tubes (Fisher Scientific, catalog number: 11558232 )
0.5 ml Eppendorf tubes (Fisher Scientific, catalog number: 10318661 )
Petri dishes (Greiner Bio-One, catalog number: 663102 )
Polycarbonate centrifuge tubes (Beckman, catalog number: 343775 )
0.22 µm filters (Merck Millipore, catalog number: SLGP033RS )
Silane-PEG (MW 30k) (Creative PEG-Works, catalog number: PSB-2014 )
Silane-PEG-biotin (MW 3,400) (LaysanBio, catalog number: BIOTIN-PEF-SIL , MW 3,400)
NeutrAvidin (ThermoScientific, catalog number: 31000 )
PLL20K-G35-PEG2K (PLL-g-PEG) (Jenkem, catalog number: 13022755 )
Pluronic F-127 (Sigma-Aldrich, catalog number: P2443 )
Bovine Serum Albumin (BSA) (Sigma-Aldrich, catalog number: A7030 )
Acetone 100% (VWR, catalog number: 20066.321 )
Ethanol 96% (VWR, catalog number: 20823362 )
Hellmanex III (Sigma-Aldrich, catalog number: Z805939-1EA )
Phosphate buffer saline (PBS) (Sigma-Aldrich, catalog number: P4417 )
1,4-Piperazinediethanesulfonic acid (PIPES) (Sigma-Aldrich, catalog number: P6757 )
Potassium hydroxide (KOH) (Sigma-Aldrich, catalog number: 484016 )
Potassium chloride (KCl) (Sigma-Aldrich, catalog number: P9541 )
Ethylene glycol-bis(2-aminoethylether)-N,N,N′,N′-tetraacetic acid (EGTA) (Sigma-Aldrich, catalog number: E3889 )
Magnesium chloride (MgCl2) (Sigma-Aldrich, catalog number: 2670 )
Sodium hydroxide (NaOH) (Carlo Erba, catalog number: 480507 )
DL-Dithiothreitol (DTT) (Sigma-Aldrich, catalog number: D0632 )
Guanosine 5′-triphosphate (GTP) (Sigma-Aldrich, catalog number: G8877 )
Methyl cellulose 1,500 cP (Sigma-Aldrich, catalog number: M0387 )
GMPCPP (Euromedex, JE-NU-405S )
HEPES (Sigma-Aldrich, catalog number: H3375 )
Glucose (Sigma-Aldrich, catalog number: G8270 )
Glucose oxidase (Sigma-Aldrich, catalog number: G6766-10KU )
Catalase (Sigma-Aldrich, catalog number: C9322-1G )
Diamond pencil (Agar Scientific, catalog number: AGT5347 )
Glutaraldehyde solution 25% (Sigma-Aldrich, catalog number: G5882 )
Clean nitrogen air flow (Air Liquide)
Liquid nitrogen (Air Liquide)
5× BRB80 (see Recipes)
BSA 10% (see Recipes)
Neutravidine (see Recipes)
DTT, 200 mM (see Recipes)
KCl, 500 mM (see Recipes)
PLL-PEG (see Recipes)
Silane-PEG or Silane-PEG-biotin (see Recipes)
GTP, 20 mM (see Recipes)
Glucose, 450 mg/ml (see Recipes)
Deoxymix (catalase and glucose oxydase) (see Recipes)
NaOH, 1 M (see Recipes)
BSA 1%/BRB80 (see Recipes)
Methyl cellulose 1,500 cP (see Recipes)
HEPES, 10 mM (see Recipes)
Pluronic F27, 10% (see Recipes)
PBS (see Recipes)
Neutravidin stock solution (see Recipes)
Red tubulin mix (see Recipes)
Imaging buffer (see Recipes)
Equipment
Inverted Eclipse Ti microscope (Nikon) with PSF focus
Apochromat 100×/1.49 N.A. oil immersion objective (Nikon) with heated objective (Okolab)
Temperature chamber (Technico Plast)
Ilas2 TIRF system (Roper Scientific)
Cooled Charged-coupled device camera ( Evolve 512 , Photometrics)
Temperature Stage Controller (Linkam Scientific)
LSM 710 confocal (Zeiss) Airyscan
Plan Apochromat 100×/1.4 N.A. oil immersion objective (Zeiss)
Ultracentrifuge (Beckman Coulter Optima, Model Max-XP, catalog number: 393315 )
Rotor (Beckman, model: TLA100-1, catalog number: 343837)
Plasma cleaner FEMTO Diener Electric (Germany) coupled to a vacuum pump Trivac D2.5E Oerliken (Germany)
Sonicator ( Elmasonic S30 , Elma, Germany)
Glass staining dishes and tray (VWR, catalog number: MARI4200004 )
Plastic box for glass microscopy (any brand)
Diamond Pencil (Oxford Instruments, catalog number: T5347 )
Tweezers (Dutscher, catalog numbers: 076100 and 005093 )
Gloves powder-free (any brand)
Water-bath (any brand)
Software
MetaMorph 7.8.5 software (Molecular Devices, https://www.moleculardevices.com)
Zen Black 2.1 (Zeiss)
Procedure
Flow chamber preparation
Note: This procedure is adapted from Portran et al. (2013) and Leslie et al. (2013). Cover glasses are used to make both sides of the perfusion chambers. Cover glasses are handled with tweezers or by hand with non-powdered gloves on the edges when drying with airflow so as not to damage their surface by more than 1 cm.
Glass cleaning
Immerge the cover glasses in a clean glass staining dish (105 × 85 × 70 mm) filled with 200-250 ml acetone and sonicate for 30 min in sweep mode.
Note: 6 cover glasses are required to make 10 flow chambers, but some glass breakage may occur during the procedure, so it is advisable to start with a little more cover glasses.
Replace acetone and incubate for an additional 30 min with orbital shaking (80 rpm).
Note: Acetone from the second bath can be recycled for the first bath of the next cleaning procedure.
Incubate 15 min with ethanol and rinse 10 times with deionized filtered water.
Incubate 2 h in prewarmed (60 °C) 2% (v/v) Hellmanex on an orbital shaker (80 rpm), then rinse 10 times with deionized filtered water.
Sonicate 15 min in 1 M NaOH solution.
Rinse 10 times with deionized filtered water.
Sonicate 15 min in ethanol and rinse each coverslip 10 times in a large volume (2 × 2 L) of deionized filtered water.
Dry the cover glasses with a clean air flow (nitrogen gas).
Dispense the glasses into two clean, dry glass staining tray for activation and silanization (see the next step below).
Note: 5 silane-PEG-biotin-coated glasses and 1 silane-PEG-coated glass are used to prepare 10 perfusion chambers (see below paragraph 3).
Glasses activation and silanization
Plasma activate the glasses for 3 min at 80% of maximum power at a pressure of 0.7 mbar.
Rapidly immerge the glasses in either silane-PEG-biotin or silane-PEG solutions and incubate overnight on an orbital shaker.
Note: Identify the glass staining dishes containing either silane-PEG-biotin or silane-PEG to avoid mistakes.
Recover the coating solutions (they can be reused for several months as long as they remain clean) and replace them with ethanol (Figure 1A).
Wash each coating slide 10 times in a beaker containing ethanol (Figure 1B) and 10 times in a large volume (2 × 2 L) of water (Figure 1C, 1D).
Note: We recommend to wash first the silane-PEG slides then the silane-PEG-biotin ones to avoid potential silane-PEG-biotin binding to silane-PEG slides.
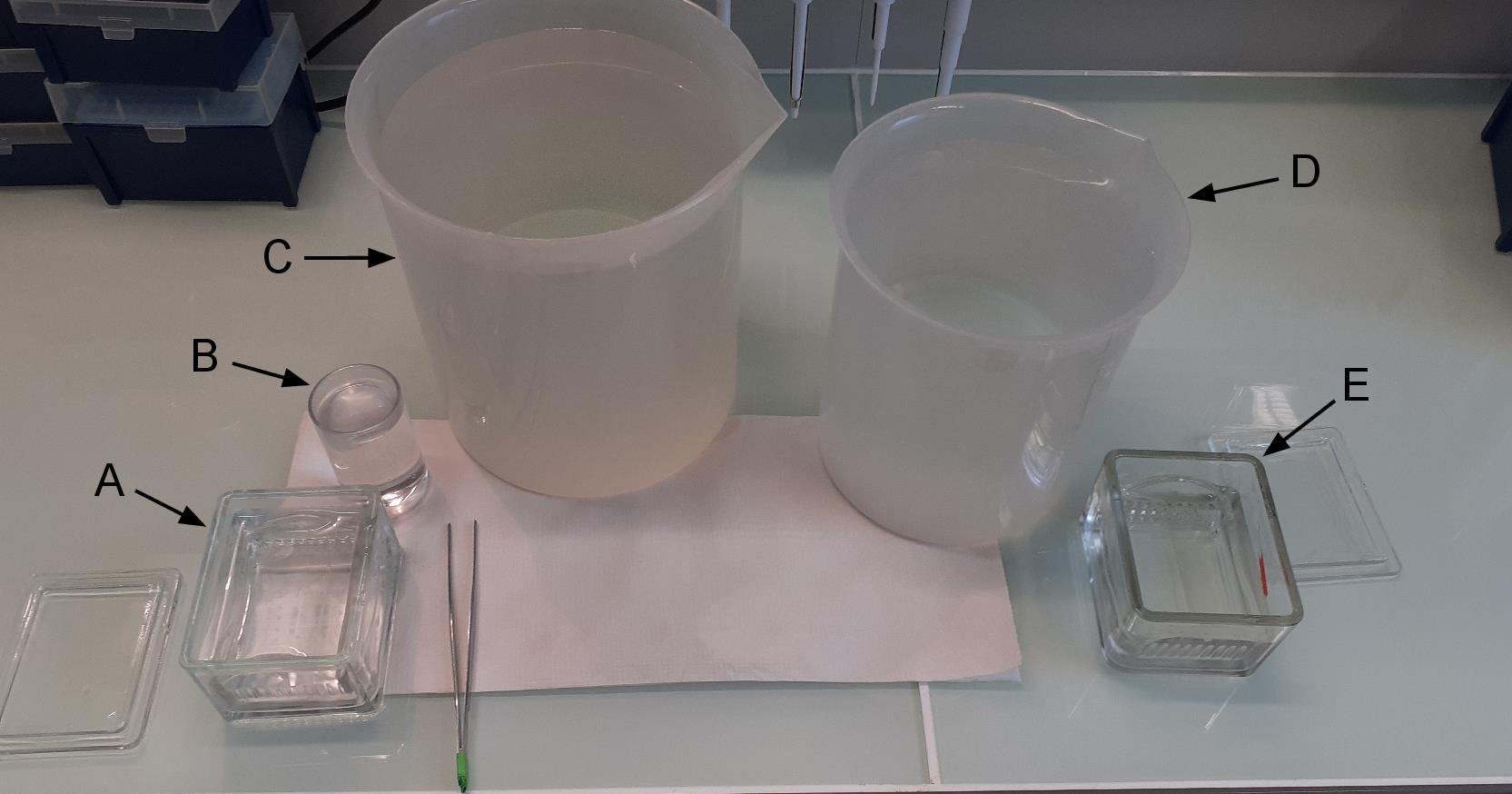

Figure 1. Washing of coated slides. In A we have coated slides in ethanol. B contains ethanol. C and D contain deionized filtered water. E containing deionized filtered water is used to keep rinsed slides before drying.Dry the coverglasses with a stream of nitrogen.
Store the silanized lamellae at 4 °C in a plastic box sealed with plastic film for a maximum of one week.
Construction of the flow chamber
Note: When handling the silanized cover glasses, make sure that the face inside the chamber remains clean and un-touched.
Transfer 1 silane-PEG-biotin-coated slide to a clean Petri dish and, using the diamond pencil, cut the glass into four parts (approx. 19/26 mm) (Figures 2A, 2B).
Note: When cutting with the diamond pencil, the Petri dish bottom bends leading to slide breaking. To avoid this, we use a thin plastic that we put under the Petri dish (Figure 2A).
Similarly, transfer the silane-PEG-coated slide to another Petri dish and cut them into 7 parts of about 11 mm width. Discard the two end parts and cut the others in half to make fourteen 11 × 13 mm pieces (Figures 2C, 2D).
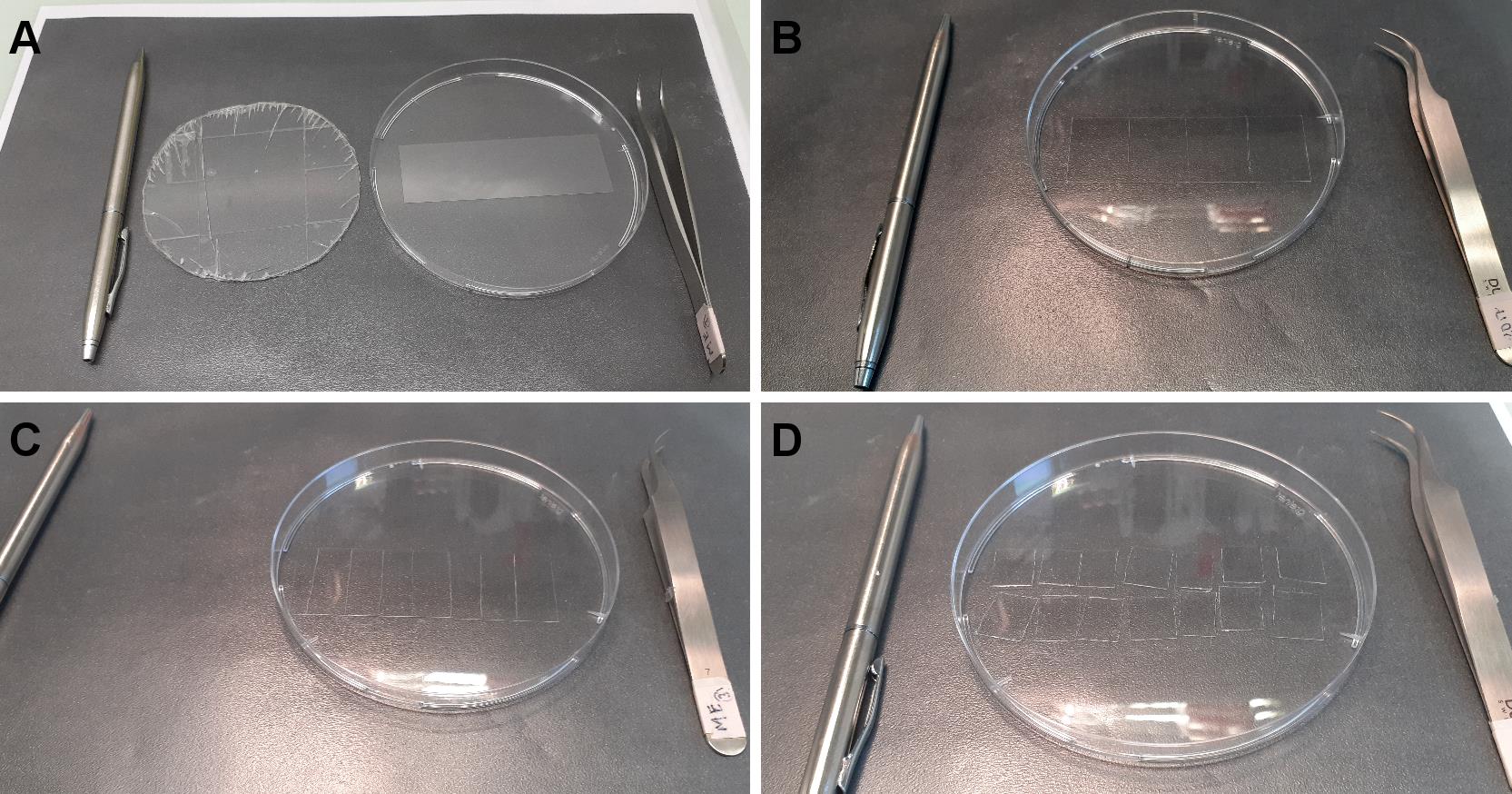

Figure 2. Cutting coated slides. A. From left to right, diamond pencil, plastic to avoid breaking, Petri dish containing silane-PEG-biotin slide, tweezers. B. The slide is cut in four equal parts. C. A silane-PEG slide is cut in 7 parts. D. Each part is cut in half.Place two pieces of double-sided adhesive tape, 5 mm apart, in the center of a piece of silane-PEG-biotin coated coverslips (Figure 3A).
Tape one piece of PEG-silane-coated strips above the two pieces of tape (the volume of the chamber is about 5 µl) (Figure 3B).
Note: Carefully press the silane-PEG-coated strip on the tape to ensure correct sealing. You might gently remove excess tape (Figure 3D).
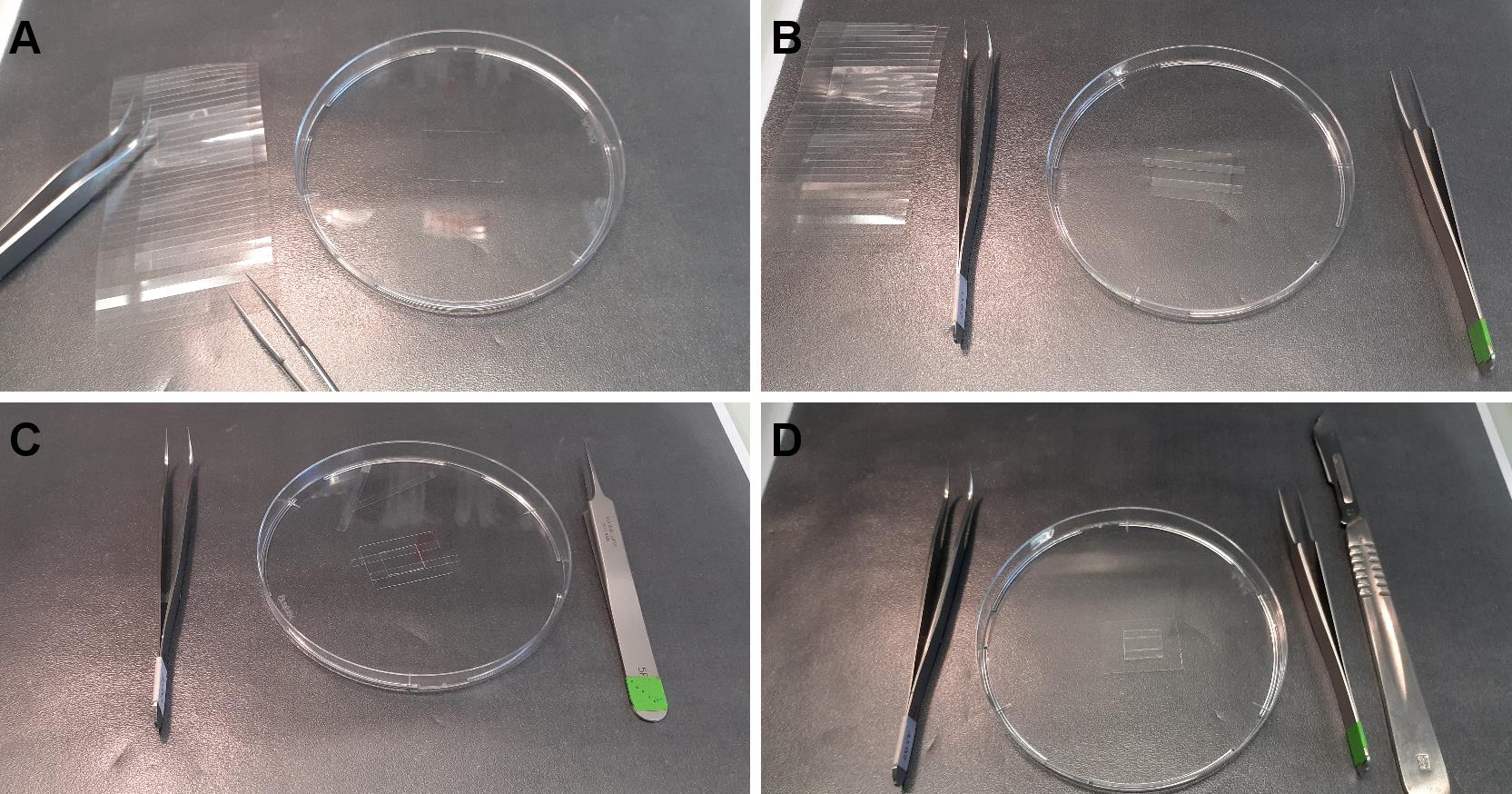

Figure 3. Construction of the flow chamber. A. One piece of silane-PEG-biotin is put in a Petri dish. B. Adhesive tape is stuck to the slide. C. Protective plastic above the tape is removed and one piece of silane-PEG slide is placed onto the tape. D. Excessive tape is cut and removed.
Preparation of GMPCCP seeds
Note: GMPCPP seeds tend to depolymerize at low temperatures, try to keep them warm all along the procedure.
Prepare a mix of freshly thawed ATTO-565 tubulin and biotinylated tubulin at a 1:1 molar ratio for a final concentration of 10 µM tubulin in BRB80 supplemented with 0.5 mM GMPCPP.
Note: Usually, 300 µl of seeds are necessary for at least 500 experiments.
The mix is placed at 35 °C in a water-bath during 1 h to allow polymerization.
Spin down the GMPCPP seeds at 130,000 × g using a TLA 100 rotor during 5 min at 35 °C.
Discard supernatant and gently rinse with 2 × 100 µl of prewarmed BRB80.
Note: After the centrifugation, the red seeds are visible in the pellet. Avoid resuspending the pellet during washing steps.
Gently resuspend the pellet in prewarmed BRB80 containing 1 mM GMPCPP.
Note: Resuspending might take some time. For 300 µl of polymerization solution, resuspend the pellet in 500 µl.
Aliquot by 1-2 µl in 0.5 ml Eppendorf and quickly freeze in liquid nitrogen.
Store in liquid nitrogen up to one year.
Microtubule polymerization and fixation
Note: The perfusion is performed by loading the solution into one side of the chamber while wiping the other side with a piece of paper (Figure 4).
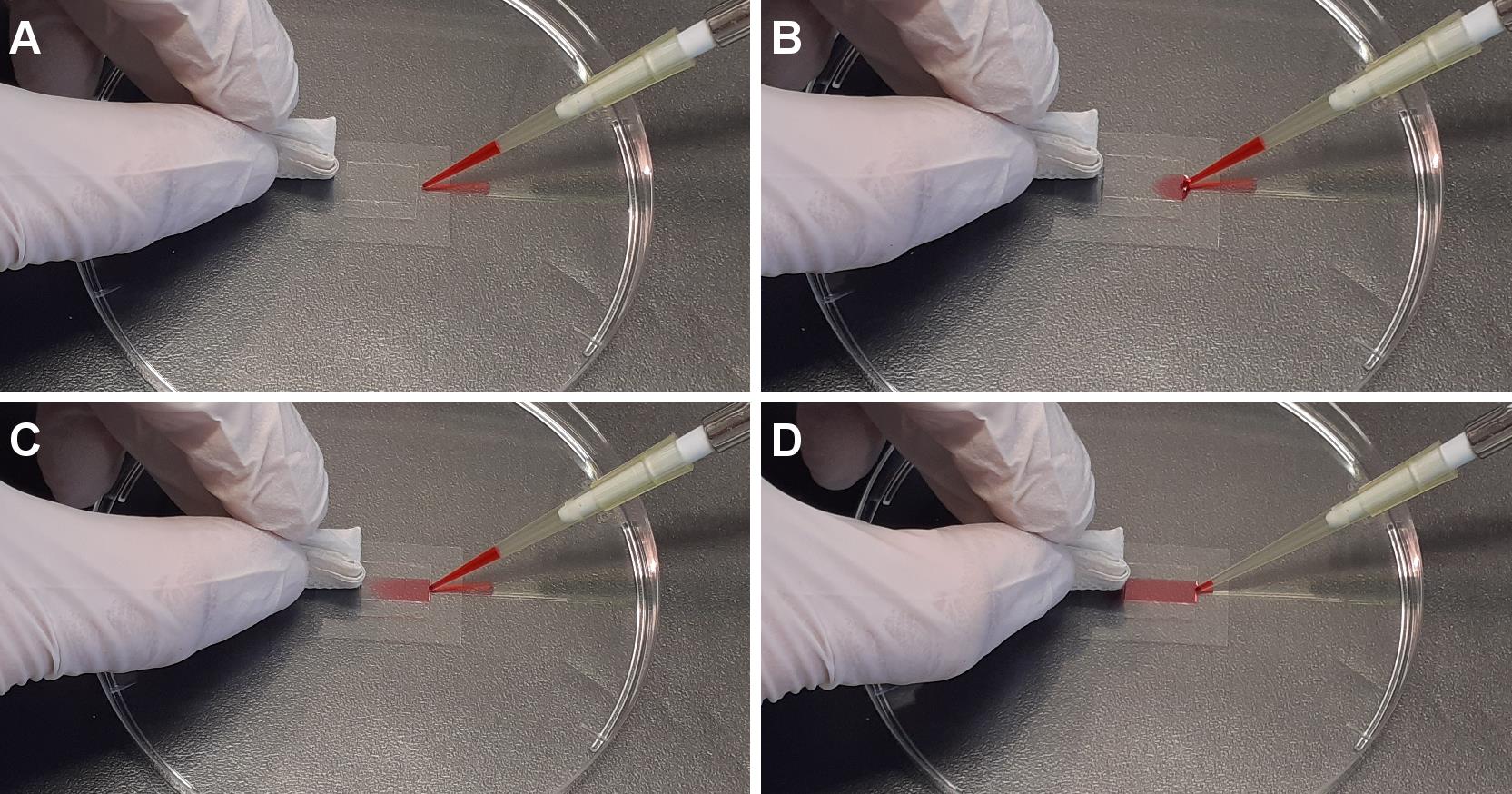
Figure 4. Perfusion in flow chamber. From A to D are sequential images of perfusion of the desired solution into the flow chamber.GMPCPP seeds immobilization in the flow chamber
Perfuse 10 µl of Neutravidin (25 µg/ml in 1% (w/v) BSA/BRB80) in the flow chamber and wait for 2 min.
Perfuse 30 µl of PLL-PEG (0.1 mg/ml in 10 mM HEPES, pH 7.4) and wait for 30 s.
Wash by perfusing 60 µl of 1% (w/v) pluronic in 1% (w/v) BSA/BRB80.
Wash by perfusing 2 times with 70 µl 1% (w/v) BSA/BRB80.
Quickly thaw GMPCPP seeds and dilute in 1% (w/v) BSA/BRB80.
Note: Adjust seeds dilution if needed, usually start with a 1/100 to 1/500 dilution.
Perfuse 30 µl of diluted GMPCPP seeds and let them adhere for 5 min at RT.
Wash unbound seeds 3 times with 70 µl 1% (w/v) BSA/BRB80.
Microtubule polymerization and fixation
Perfuse freshly prepared polymerization mix (Recipe 19).
Note: Adjust tubulin/associated proteins concentration if needed. It depends on the affinity and effects of the associated proteins of interest. For tubulin, a concentration of 5 to 20 µM is a good start and for associated proteins from 5 nM to 500 nM should be informative.
Put the flow chamber in a warm (32-35 °C) place with a water-saturated atmosphere to avoid desiccation for MT polymerization (usually at least 30 min).
After polymerization, fix MTs by gently perfusing approximately 30 µl of 0.1% Methyl cellulose 1,500 cP, 0.5% (v/v) Glutaraldehyde/BRB80.
Note: Critical step, fast perfusion will alter MT structure. Put the chamber inside a Petri dish (you might use tape to prevent it from moving). Then, slightly tilt the Petri dish and add drop by drop the fixing buffer at the top of the chamber. The solution will slowly flow down the chamber. You can use blotting paper to clean excess liquid. The fixation buffer should be at least 3 min inside the chamber for proper fixation. Even if the fixation occurs rapidly, dilution might induce MT depolymerization. To overcome this issue, use MT stabilizing compounds such as taxol (taxol affects MT persistence length) or grow MT longer.
Wash fixation buffer with 50 µl of 0.1% (v/v) Methyl cellulose 1,500 cP/BRB80.
Perfuse imaging buffer (BRB80 supplemented with 0.1% Methyl cellulose 1,500 cP, 2 mg/ml glucose, 1mg/ml glucose oxidase and 150 µg/ml catalase).
Seal the flow chamber with the siligum wax.
Note: Imaging of the chambers should be done within one week.
Imaging of the microtubules
Control fixation: TIRF Microscopy
Note: To estimate the effect of the fixation process on the sample, it is necessary to compare the sample before/after fixation.
Place the flow chamber on the inverted microscope.
Note: For imaging before fixation, set the temperature of the stage at 35 °C to avoid MT depolymerization.
Adjust laser intensity, exposure and focus in order to observe the microtubules.
Acquire images both before (Figure 5A) and after fixation (Figure 5B) of the sample.
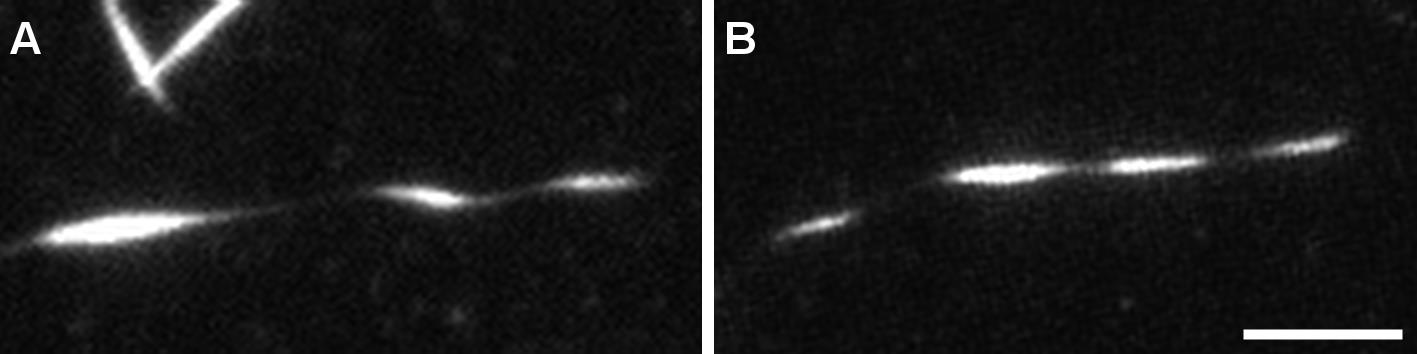
Figure 5. Structure of helical microtubules is not altered by the fixation process. TIRFM image of a helical MT before (A) and after (B) fixation. MTs were grown from GMPCPP seeds with 12 µM tubulin (9:1 non-labeled tubulin:ATTO-565 labelled tubulin) with 200 nM MAP6-N-GFP. At this concentration, MAP6-N-GFP induces microtubule coiling. Scale bar: 5 µm.Confocal imaging with Airyscan processing
Place the chamber above the objective.
Adjust focus and laser intensity in order to observe MTs.
Perform acquisition (z-stack series using 220 nm step-size).
Use Zen built-in Airyscan processing. See Figure 6 for TIRF versus confocal imaging with Airyscan processing of helical MT. Figure 7 shows the advantage of performing z-stacks on MTs with a particular shape.
Analyse images using Zen Blue.
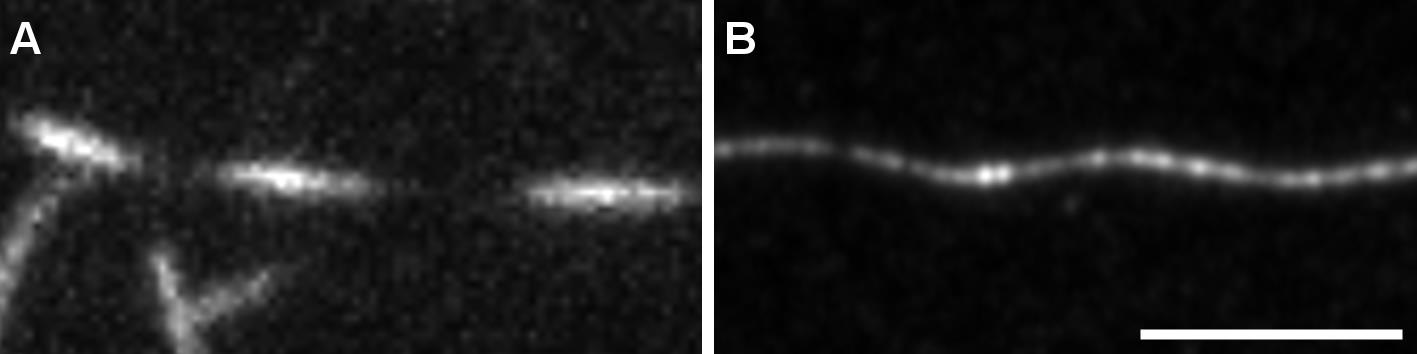
Figure 6. Observation of helical MTs with TIRFM or confocal Airyscan. MTs were grown as in Figure 5 and fixed using the described protocol. A. Fixed helical MTs observed using TIRF microscopy. B. z-projection of a fixed helical MT observed using confocal microscope followed by Airyscan processing. Scale bar: 5 µm.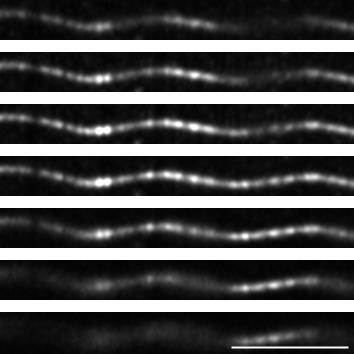
Figure 7. z-stack of a helical MT observed using confocal imaging with Airyscan processing. Images of the z-stack of the same MT as in Figure 6B, with a step of 220 nm between each plane. Scale bar: 5 µm.
Recipes
All solutions were conserved at indicated temperature without any observable deterioration along time unless stated otherwise. Most solutions were aliquoted to avoid freezing-thawing cycles.
5× BRB80
36.28 g PIPES
1.5 ml of MgCl2 (1 M)
570.6 mg EGTA
Dissolve in 250 ml of deionized water and adjust pH to 6.85 with KOH
Add deionized water to 300 ml, filtrate, aliquot (1 ml and 50 ml) and store at -20 °C
Note: The 5× BRB80 is used to make the BRB80 solution and to adjust the final concentration of the reaction to 1× BRB80. Upon 5× dilution the pH drops to 6.75.
BSA 10%
Dissolve 2 g of BSA in 20 ml PBS
Filter and store at -20 °C up to 1 year
Neutravidine
Dissolve 10 mg Neutravidine in 10 ml H2O, aliquot (5 and 100 µl) and store at -20 °C
Before the experiment, predilute 5 µl with 195 µl of 1% BSA/BRB80 (final concentration 25 µg/ml)
DTT, 200 mM
Dissolve 617 mg DTT powder in 4 ml deionized H2O, filtrate, aliquot and store at -20 °C
The day of use, dilute 1/5 in H2O or BRB80. Discard aliquot after each day of use
KCl, 500 mM
Dissolve 1 g KCl into 50 ml deionized H2O (-20 °C) filtrate, aliquot and store at -20 °C
PLL-PEG
Make aliquot of the powder of known weight (20-30 mg) under argon gas and store at -20 °C.
To make stock solution, dissolve the powder in 10 mM HEPES (pH 7.4) at 1 mg/ml, make 50 µl aliquots and store at -20 °C.
The day of use, dilute one 50 µl aliquot with 450 µl of 10 mM HEPES (pH 7.4). Diluted PLL-PEG can be stored at 4 °C for one week.
Silane-PEG or Silane-PEG-biotin
Notes:
Silane-PEG or Silane-PEG-biotin powders are weighted and aliquoted in 200 mg per eppendorf tube under argon gas. The tubes are tightly sealed with parafilm and stored at -20 °C.
Silane-PEG and Silane-PEG-biotin solutions must be kept anhydrous and in the dark.
Dissolve 200 mg powder in 200 ml of 96% ethanol plus 0.4 ml of 37% HCl.
To solubilize the Silane-PEG solution, heat to 50 °C in a water-bath. Keep the solution in a glass bottle.
Silane-PEG and silane-PEG-biotin solution are stored at room temperature in the dark for up to 4 months or around 15 coating. If experiments look dirty in the background, use freshly prepared coating solutions.
GTP 20 mM
Dissolve 250 mg GTP in 5 ml H2O
Make aliquots and store at -20 °C
Before use, dilute the stock solution to 20 mM in H2O or BRB80
Glucose 450 mg/ml
Dissolve 2.250 g of glucose in 5 ml of BRB80
Filtrate, aliquot and store at -80 °C
The day of use pre-dilute 1/10 in BRB80 and keep on ice
Deoxymix (catalase and glucose oxydase)
Dissolve 35 mg of catalase plus 250 mg of glucose-oxydase in 10 ml of BRB80
Filtrate, make 25 µl aliquots, freeze in liquid nitrogen and store at -80 °C
Discard thawed aliquot after each day of use
NaOH, 1 M
Dissolve 8 g of NaOH in 200 ml deionized water
Discard after each use
BSA 1%/BRB80
Dilute 500 µl of 10% BSA with 1 ml of BRB80 5× in 3.5 ml of filtrated deionized water
Methyl cellulose 1,500 cP
Dissolve 100 mg of methyl cellulose 1,500 cP in 10 ml of prewarm (60 °C) deionized water
Gently shake on rotating wheel for 30 min
Store at 4 °C for two weeks
HEPES, 10 mM
Dissolve 477 mg in 200 ml deionized water
Adjust pH to 7.4 with KOH
Store in aliquots at -20 °C
Pluronic F27, 10%
Dissolve 100 mg in 1 ml of deionized water
Store at 4 °C up to 2 months
PBS
Dissolve 1 tablet in 200 ml of deionized water
Store at -20 °C
Neutravidin stock solution
Dissolve Neutravidin in deionized water at a final concentration of 1 mg/ml
Store at -20 °C
Red tubulin mix
Dilute purified non-labelled and ATTO-565 tubulin at a 9:1 molar ratio in BRB80
Spin down aggregates at 100,000 × g, 4 °C using TLA100 rotor
Estimate tubulin concentration by measuring the OD280nm (1OD280nm = 1 mg/ml = 10 µM of tubulin)
Aliquot, quickly freeze and store in liquid nitrogen for up to 1 year
Polymerization mix
BRB80 supplemented with:
50 mM KCl
1% BSA
4 mM DTT
1 mM GTP
1 mM glucose
0.05% Methyl cellulose 1,500 cP
1/50 deoxy mix (70 µg/ml catalase, 500 µg/ml glucose oxidase, 1 mg/ml glucose)
12 µM red tubulin mix
200 nM MAP6-N-GFP
Acknowledgments
Fundings: This work was supported by INSERM (Institut National de la Santé Et de la Recherche Médicale), CEA (Commissariat à l’Energie Atomique), CNRS (Centre National de la Recherche Scientifique), Université Grenoble Alpes, by grants from the Agence Nationale de la Recherche ANR MAMAs 2017-CE11-0026, ANR-15-IDEX-02 NeuroCoG in the framework of the “Investissements d’avenir” program and by fundings from the Ministère de l’enseignement supérieur et de la recherche. GIN is a member of the Grenoble Center of Excellence in Neurodegeneration (GREEN). The Photonic Imaging Center of Grenoble Institute Neuroscience (Univ. Grenoble Alpes–Inserm U1216) is part of the ISdV core facility and certified by the IBiSA label.
This protocol is derived from Cuveillier et al. (2020).
Competing interests
The authors declare no competing interests.
References
- Akhmanova, A. and Steinmetz, M. O. (2015). Control of microtubule organization and dynamics: two ends in the limelight. Nat Rev Mol Cell Biol 16(12): 711-726.
- Al-Bassam, J. (2014). Reconstituting Dynamic Microtubule Polymerization Regulation by TOG Domain Proteins.In: Methods in Enzymology (Vol. 540, p. 131-148). Elsevier.
- Alushin, G. M., Lander, G. C., Kellogg, E. H., Zhang, R., Baker, D. and Nogales, E. (2014). High-Resolution Microtubule Structures Reveal the Structural Transitions in αβ-Tubulin upon GTP Hydrolysis.Cell 157(5): 1117-1129.
- Bechstedt, S., Lu, K. and Brouhard, G. J. (2014). Doublecortin Recognizes the Longitudinal Curvature of the Microtubule End and Lattice. Curr Biol 24(20): 2366-2375.
- Blom, H. and Brismar, H. (2014). STED microscopy : Increased resolution for medical research? J Intern Med 276(6): 560-578.
- Brangwynne, C. P., MacKintosh, F. C., Kumar, S., Geisse, N. A., Talbot, J., Mahadevan, L., Parker, K. K., Ingber, D. E. and Weitz, D. A. (2006). Microtubules can bear enhanced compressive loads in living cells because of lateral reinforcement. J Cell Biol 173(5): 733-741.
- Cuveillier, C., Delaroche, J., Seggio, M., Gory-Fauré, S., Bosc, C., Denarier, E., Bacia, M., Schoehn, G., Mohrbach, H., Kulić, I., Andrieux, A., Arnal, I. and Delphin, C. (2020). MAP6 is an intraluminal protein that induces neuronal microtubules to coil. Sci Adv 6(14): eaaz4344.
- Harris, J. R. (2015). Transmission electron microscopy in molecular structural biology : A historical survey.Arch Biochem Biophys 581: 3-18.
- Leslie, K. and Galjart, N. (2013). Going Solo: measuring the motions of microtubules with an in vitro assay for TIRF microscopy. In: Methods in Cell Biology (Vol. 115, p. 109-124). Elsevier.
- Leung, J. M., Nagayasu, E., Hwang, Y.-C., Liu, J., Pierce, P. G., Phan, I. Q., Prentice, R. A., Murray, J. M. and Hu, K. (2020). A doublecortin-domain protein of Toxoplasma and its orthologues bind to and modify the structure and organization of tubulin polymers. BMC Mol Cell Biol 21(1): 8.
- Portran, D., Gaillard, J., Vantard, M. and Thery, M. (2013). Quantification of MAP and molecular motor activities on geometrically controlled microtubule networks.Cytoskeleton 70(1): 12-23.
- Ramirez-Rios, S., Serre, L., Stoppin-Mellet, V., Prezel, E., Vinit, A., Courriol, E., Fourest-Lieuvin, A., Delaroche, J., Denarier, E. and Arnal, I. (2017). A TIRF microscopy assay to decode how tau regulates EB’s tracking at microtubule ends. In: Methods in Cell Biology (Vol. 141, p. 179-197). Elsevier.
- Weisenberg, R. C. (1972). Microtubule Formation in vitro in Solutions Containing Low Calcium Concentrations. Science 177(4054): 1104-1105.
Article Information
Copyright
© 2021 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Cuveillier, C., Saoudi, Y., Arnal, I. and Delphin, C. (2021). Imaging Microtubules in vitro at High Resolution while Preserving their Structure. Bio-protocol 11(7): e3968. DOI: 10.21769/BioProtoc.3968.
- Cuveillier, C., Delaroche, J., Seggio, M., Gory-Fauré, S., Bosc, C., Denarier, E., Bacia, M., Schoehn, G., Mohrbach, H., Kulić, I., Andrieux, A., Arnal, I. and Delphin, C. (2020). MAP6 is an intraluminal protein that induces neuronal microtubules to coil. Sci Adv 6(14): eaaz4344.
Category
Neuroscience > Cellular mechanisms
Biochemistry > Protein > Imaging
Biochemistry > Protein > Fluorescence
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










