- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Bacterial Conjugation Protocol for Ruminant Mycoplasmas
(*contributed equally to this work) Published: Vol 11, Iss 2, Jan 20, 2021 DOI: 10.21769/BioProtoc.3893 Views: 3453
Reviewed by: Alba BlesaOscar Quijada Pich Alvaro Eduardo Galvis

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Efficient Agrobacterium-mediated Transformation of the Elite–Indica Rice Variety Komboka
Van T. Luu [...] Wolf B. Frommer
Sep 5, 2020 9922 Views
A New Tool for the Flexible Genetic Manipulation of Geobacillus kaustophilus
Ryotaro Amatsu [...] Ken-ichi Yoshida
Sep 5, 2022 1918 Views
A Quick DNA Extraction Method for High Throughput Screening in Gram-positive Bacteria
Nuo Chen and Xiaoming Yuan
Apr 20, 2023 2744 Views
Abstract
In Mycoplasma agalactiae, two simultaneous processes of DNA transfer have been described that require direct cell-to-cell contact and are similar to conjugation. One involves the self-transmission of an integrative conjugative element (ICE) while the second concerns the horizontal transfer of large and small fragments of chromosomal DNA. Here, we describe an optimized conjugation protocol for the horizontal transfer of ICE or chromosomal DNA carrying antibiotic resistance markers (i.e., tetracycline, gentamicin, puromycin) from donor to recipient mycoplasma cells. Calculation of the conjugation frequencies, selection and characterization of transconjugants are detailed. This protocol has been developed with M. agalactiae but has been successfully used for M. bovis and can be adapted to other related mycoplasma species.
Keywords: MycoplasmaBackground
Conjugative, horizontal DNA transfer is a key player of microbial diversification. By promoting an active transfer of DNA through close cell-to-cell contact (Lederberg and Tatum, 1946), this phenomenon facilitates the rapid acquisition of new traits from external resources. Mycoplasmas (class Mollicutes) are wall-less bacteria with reduced genomes whose evolution has been considered to be only driven by gene loss and in which horizontal gene transfer (HGT) was long thought to be marginal. During this past decade, comparative genomic analyses have revisited this paradigm and have shown (i) the occurrence of HGTs events in between mycoplasma species sharing same hosts (Sirand-Pugnet et al., 2007) and (ii) the presence of integrative and conjugative elements (ICEs) in a large number of sequenced mycoplasma genomes (Calcutt et al., 2002; Marenda et al., 2006; Dordet-Frisoni et al., 2013; Tardy et al., 2015; Meygret et al., 2019), raising the prospect that these minimal bacteria might be able to conjugate.
In vitro conjugation-like process has been demonstrated in Mycoplasma agalactiae (Dordet-Frisoni et al., 2013, 2014 and 2019), a mycoplasma species pathogenic for ruminants. Mating assays were based on mixing in liquid culture two strains tagged with antibiotic resistant genes and resulted in a bidirectional DNA transfer process (Dordet-Frisoni et al., 2013 and 2014). The first is the conventional horizontal dissemination of a mycoplasma ICE (MICE), from ICE-positive to ICE-negative cells (Dordet-Frisoni et al., 2013). The second concerns the transfer of large and small blocks of chromosomal DNA from ICE-negative to ICE-positive cells and was further referred to as MCT for mycoplasma chromosomal transfer (Dordet-Frisoni et al., 2014 and 2019; Faucher et al., 2019). M. agalactiae conjugation has been shown to rely on the presence of an ICE in at least one parent cell, but both DNA transfers are physically independent (taking place in opposite direction) and occur at the same frequency (around 10-8 transconjugants/total CFU) (Dordet-Frisoni et al., 2014).
The detailed protocol presented here describes an optimized mating procedure developed for the conjugation of M. agalactiae and related species. For instance, as described in Dordet-Frisoni et al. (2014), this protocol has also been successfully applied to M. bovis and can most likely be adapted to other mycoplasma species with some adjustments.
Materials and Reagents
Sterile tips for micropipettes (STARLAB, catalog numbers: S1122-1830 [1,000 µl], S1120-3810 [10-20 µl] )
Sterile 1.5 ml and 2 ml microcentrifuge tubes (SORENSON, catalog number: 017040A; TREFF, catalog number: 96.09329.9.01)
Sterile plastic Petri dish plates (90 mm diameter, Clearline, catalog number: 076084BS )
0.22 µm pore filter (PVDF 33 mm, ClearLine, Dutscher, catalog number: 051733)
Donor and recipient cells, harbouring different selective markers for antibiotic selection #
#Note: M agalactiae strains* were tagged with selectable markers previously introduced by transposition using a pMT85-derived plasmid (Figure 1) (Zimmerman and Herrmann, 2005). pMT85 and derivatives bear a mini-Tn with either the aacA-aphD gentamicin resistance cassette, the tetM determinant for resistance to tetracycline or the pac gene conferring puromycin resistance. Marker genes are flanked by two Tn4001 inverted repeats (IRs). Since the mini-Tn backbone contains no transposase gene (it is located elsewhere in the pMT85), its insertion into the cell chromosome is stable (Baranowski et al., 2010).
Several Mycoplasma agalactiae strains such as 5632, PG2, 4867, 14628 and 4055, have been successfully employed in mating assays (Dordet-Frisoni et al., 2019), but conjugation experiment has been also successful with other ruminant mycoplasmas species such as M. bovis PG45 strain (Dordet-Frisoni et al., 2014) and M. mycoides subsp. capri GM12 strain (personal data). For conjugation to occur, at least one of the two partners should carry an ICE. “Concerning chromosomal transfers (MCT), only two M. agalactiae strains have been documented as recipient cells, namely 5632 and 4867, while all strains tested so far can act as donor. As for ICEA, all strains carrying an active ICE are potentially donor cell. All parental M. agalactiae strains cited above can be acquired at the Vigimyc collection (Chazel et al., 2010).
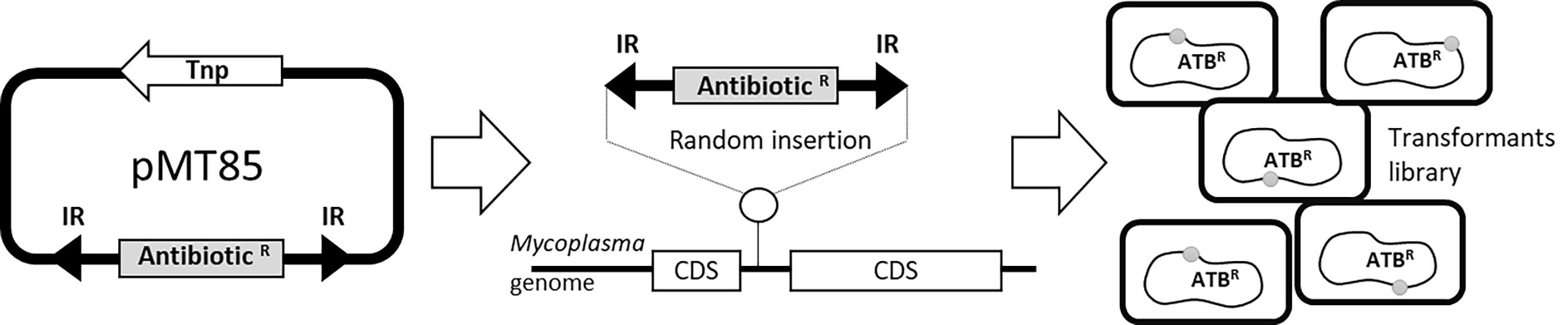
Figure 1. Schematic representing M. agalactiae antibiotic tagging procedure. Selectable markers are introduced by transposition using the mini transposon pMT85, which bear antibiotic resistance genes as selection markers (AntibioticR). The AntibioticR marker is flanked by two inverted (IR) Tn4001 repeats. During insertion, the transposase (Tnp) which is located outside the IRs is eliminated and the selection marker is integrated into the chromosome in a random (in coding (CDS) or non-coding sequences) and a stable manner. This experiment results in mycoplasmas transformants tagged with resistance markers (ATBR) at different locations on the chromosome.Appropriate selectable antibiotics:
Gentamicin (GmR) (Gibco, catalog number: 15750037 )
Tetracycline (TetR) (Sigma-Aldrich, catalog number: T3383 )
Puromycin (PurR) (Fisher, catalog number: 10054207 )
Ampicillin (Sigma-Aldrich, catalog number: A9518 )
PPLO broth (Difco, catalog number: 255420 )
Tryptone peptone (Difco, catalog number: 211705 )
Bacto peptone (Difco, catalog number: 211677 )
FBS (Fetal Bovine Serum) (Gibco, catalog number: 10270-106 )
CMRL 1066 (10x) (Gibco, catalog number: 21540-026 )
TC yeastolate (Difco, catalog number: 255772 )
Yeast extract 15% (Gibco, catalog number: 18180-059 )
D-(+)-Glucose (Sigma-Aldrich, catalog number: G6152 )
Phenol red 0.5% (Sigma-Aldrich, catalog number: P0290 )
Na Pyruvate (Sigma-Aldrich, catalog number: P2256 )
Select agar (Invitrogen, catalog number: 30391-023 )
NaOH (Sigma-Aldrich, catalog number: S8045 )
D-PBS (Gibco, catalog number: 14190169 )
Tween-20 (Sigma-Aldrich, catalog number: P9416 )
Ultra pure Tris (Invitrogen, catalog number: 15504020 )
HCl (Sigma-Aldrich, catalog number: 320331 )
Proteinase K 20 mg/ml (Qiagen catalog number: 19131 )
Taq DNA Polymerase (New England Biolabs catalog number: M0267L )
dNTP mix 10 mM (New England Biolabs catalog number: N0447S )
Taq Buffer 10x (New England Biolabs catalog number: M0267L )
Sterile SP4 liquid medium (see Recipes)
Sterile SP4 agar plates (see Recipes)
M. agalactiae Lysis buffer (see Recipes)
Equipment
Air-flow cabinet (Faster, model: SF Classic 212 )
37 °C incubator (Firlabo, model: FA Type B 407L-FA407B )
Microcentrifuge with rotor for 1.5-2 ml tubes (Eppendorf, model: 5424 )
Vortex (Scientific Industries, model: GENIE 2 )
Binocular magnifier (Olympus, model: SZ 60 )
Multiblock heater (Lab Line Instruments, model: 2052-1CCE )
Thermocycler (Mastercycler EP Eppendorf Gradient, model: 5341 )
Micropipettes (GIilson Pipetman Classic 10 μl to 1 ml range, Dutscher, catalog numbers: 066002 , 066003 , 066004 , 066005 and 066006 )
Graduated pipettes (Falcon 5 to 25 ml range pipettes, Dutscher, catalog numbers: 357543 , 357551 and 357535 )
Pipettor (Pipetboy Acu 2, INTEGRA, DUTSCHER, catalog number: 060201 )
Procedure
A schematic of the procedure is also provided bellow (Figure 2).
Culture preparation
Inoculate 1 ml of SP4 liquid medium with 10 μl of each saturated liquid growth culture of M. agalactiae (standard bacterial stock at ~109 CFU/ml) previously stored at -80 °C in the absence of glycerol or particular storage conditions and defrosted at room temperature.
Incubate the culture at 37 °C for 24 h without shaking in an incubator #.
#Note: Growth rate depends on the M. agalactiae strain and can vary from 16 h to 48 h of incubation in SP4 medium. Change from red to orange of the phenol-red present in the SP4 medium indicates optimal growth.
Mating procedure
Mix 1 ml# of each saturated mycoplasma culture (between 108 and 109 CFU/ml) in a 2 ml sterile microcentrifuge tubes.
# Note: The ratio of donor:recipient mycoplasma cells should be close to 1:1. Depending on the assays or on the concentration of the two mycoplasma cultures (e.g., due to differences in growth rates), the ratio can be changed to 1:2 (vol:vol) or 1:10 (vol:vol) in the same final volume of 2 ml.
Centrifuge the 2 ml mixed cultures for 5 min at 8,000 x g at room temperature (20-25 °C).
Carefully discard the supernatant using a 1 ml micropipette.
Resuspend the pellets with 1 ml of SP4 liquid medium at room temperature, proceed to 2 to 3 drawing/discharging cycles with a 1 ml micropipette.
Incubate over night# at 37 °C without shaking.
#Note: Conjugation events can be observed after 3 h of incubation, but the highest mating frequency is observed after at least 16 h.
Plating
A part of the mating mix (10 to 50 µl) is used to determine the total CFU#. For this purpose, prepare serial dilutions in DPBS 1x + 5% decomplemented FBS (from 10-1 to 10-7) and spot 10 µl of each dilution onto a SP4 agar plate containing no selective pressure.
#Note: If the conjugation efficiency is expressed by the ratio of transconjugants/donor cells or transconjugant/recipient cells, spot 10 µl of each dilution onto a SP4 agar plate containing the antibiotic marker corresponding to the donor/recipient strain, at the appropriate concentration.
Inoculate 300 µl of the mating mix onto selective SP4 agar plates. Spread the liquid onto the surface of the Petri dish (90 mm) by gentle circular movements so that the entire dish (except the edges) is covered#.
#Note: All the mating mix can be plated in 3 Petri dishes. For highly efficient matings, dilutions should be made before plating (1/10x, 1/100x maximum). These dilutions are made with sterile DPBS 1x complemented with 5% decomplemented FBS in microcentrifuge tubes. The mating mixture can be stored as such directly at -80 °C if necessary.
Incubate the SP4 agar plates at 37 °C for 3 to 5 days#.
#Note: To prevent desiccation during incubation the Petri dishes should be stored inside closed plastic bags, in the 37 °C incubator.
Calculation of conjugation frequencies
Using the binocular magnifier (10x-63x), count the Colony Forming Unit (CFU) on the 10µl dilution spots or the 300 µl spreads (Figure 2) of each selective and non-selective SP4 agar plates.
Calculate the mating frequency# as the CFUs grown in both selective markers (i.e., TetR and GmR) divided by the total CFU obtained in dilution spots spread on SP4 agar without selective pressure.
#Note: The conjugation frequency can also be expressed as the number of transconjugants per donor or recipient cell by dividing the CFU cultivated in the two selective markers, by the CFU cultivated with only one of the two selective marker (that of the donor, or that of the recipient, see note above).
At least three independent experiments should be performed for reliable and reproducible data.
Picking, cloning and storage of transconjugants
Pick colonies with a sterile 200 µl micropipette sterile tip and transfer it into 1 ml of SP4 liquid medium supplemented with the 2 appropriate antibiotics.
Incubate at 37 °C for 24 to 96 h#.
#Note: Transconjugants can be directly stored at -80 °C at this stage .
To ensure the clonality of transconjugants, three successive subcloning steps can be performed as previously described (Rosengarten et al., 1994), with the last broth culture being filtered through a 0.22 µm pore filter before storage at -80 °C.
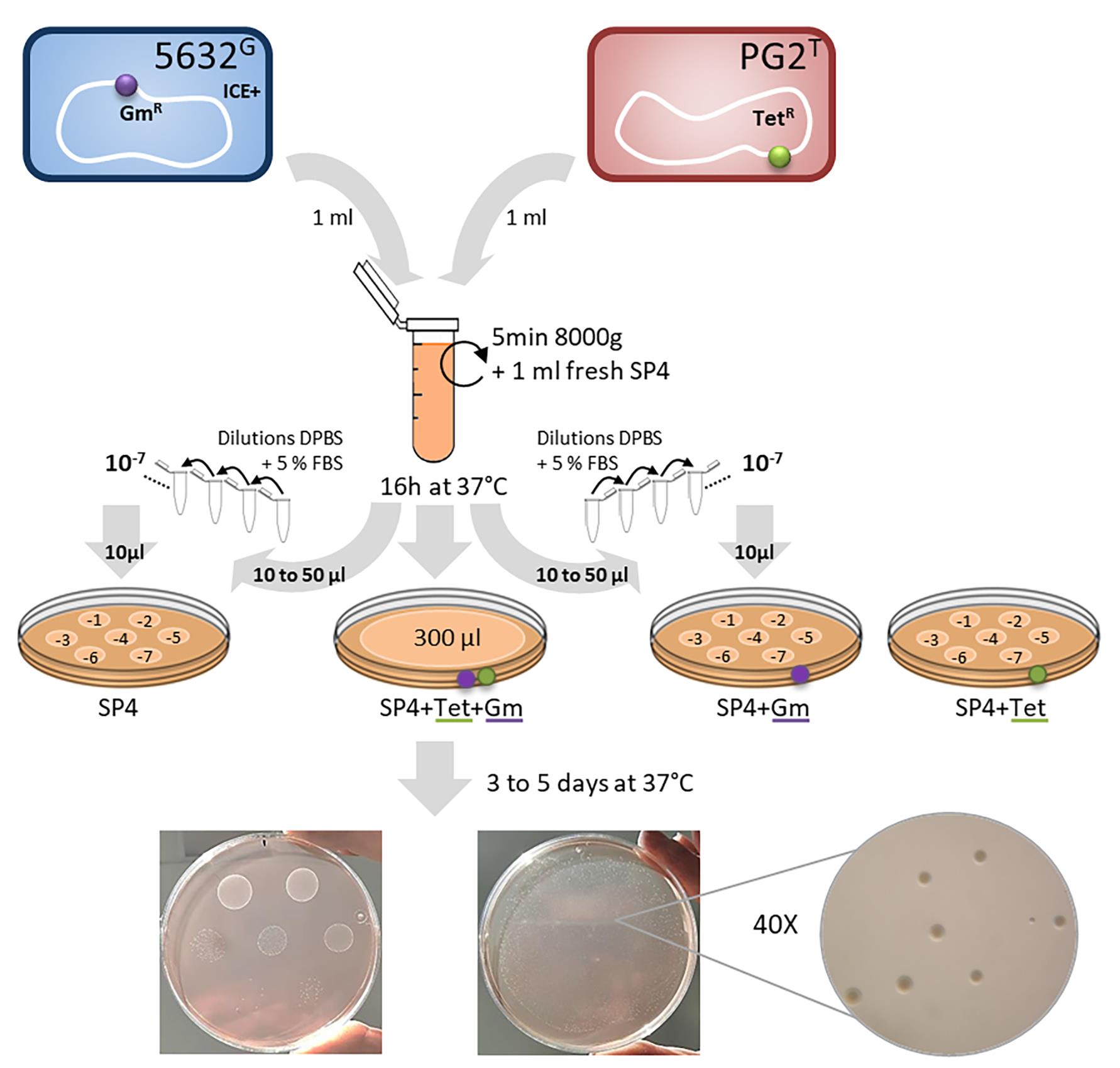
Figure 2. Schematic representing M. agalactiae mating procedure. Here, as an example, the M. agalactaiae strain 5632 (ICE containing strain, ICE+) was mated with strain PG2 (ICE-). Prior to mating, strains were independently tagged by stably inserting the gentamicin resistance gene (GmR, in purple) or the tetracycline resistance gene (TetR, in green) in their genome, using the pMT85 plamid (Baranowski et al., 2010). Saturated cultures of each of these strains (≈ 109 CFU/ml) were mixed in equal proportion (1 ml each), gently centrifuged, resuspended in fresh SP4 medium and incubated together. The mix culture was then directly plated on appropriate selective medium and dilutions were inoculated onto selective and non-selective plates to determine the conjugation frequency.
Verification of putative transconjugants
Centrifuge a 1 ml culture of cloned transconjugants at 12,000 x g for 25 min at 4 °C.
Remove the supernatant and add 50 μl of lysis buffer (see Recipes section) to the cell pellet
Incubate at 37 °C for 1 h.
Inactivate the proteinase K contained in the lysis buffer by heating at 95 °C for 10 min.
Perform PCR amplifications using specific primers targeting the resistance genes present in the parental strains used for mating (Table 1) (Dordet-Frisoni et al., 2014). To do this, take 2 µl of the lysed transconjugants culture and perform the PCR using the standard Taq DNA polymerase procedure (here, Taq DNA polymerase from NEB).
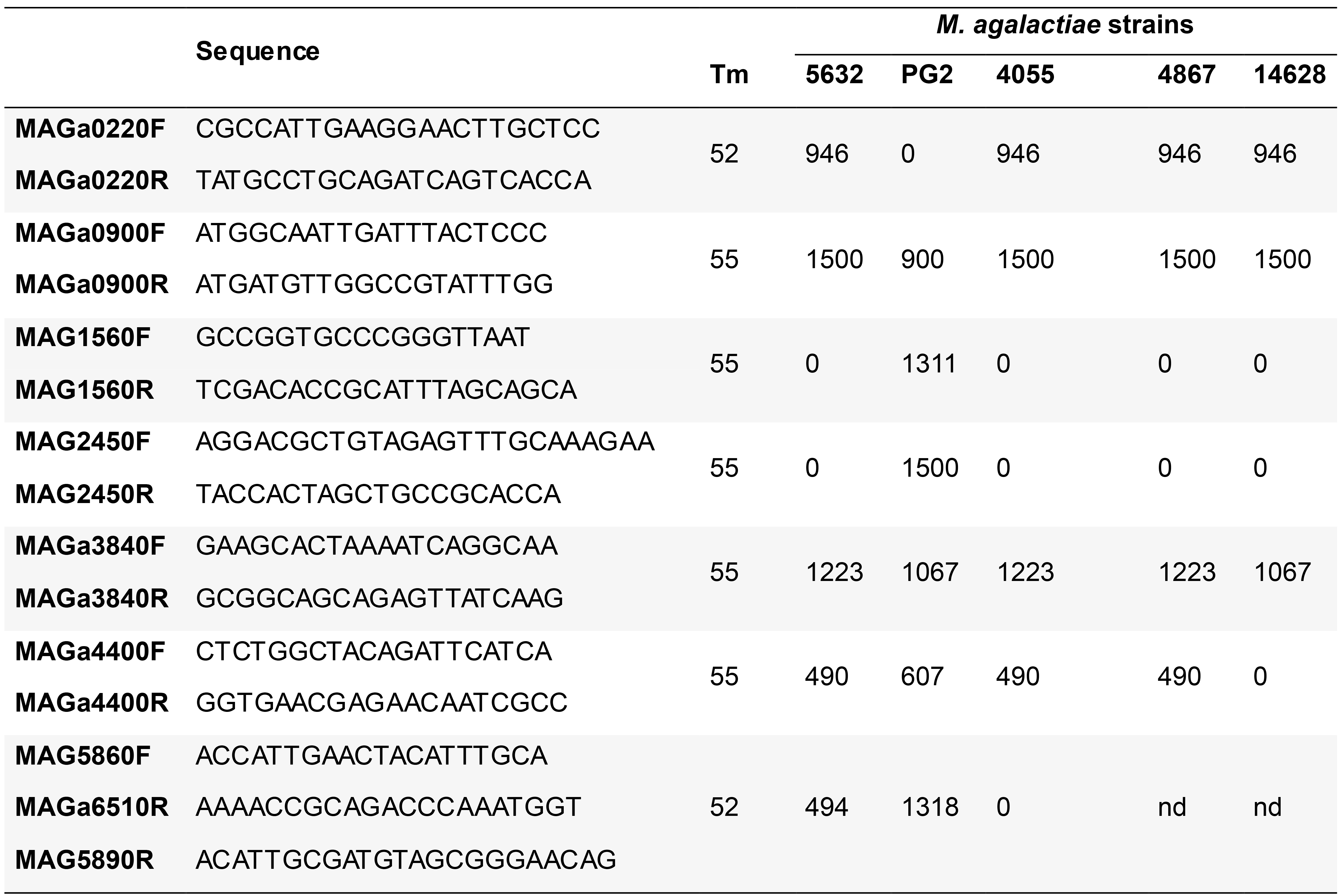
Identify transconjugant genomic backbone by PCR using a set of 7 primer pairs (Table 2) that covers clockwise the M. agalactiae genome and produces PCR fragments specific of the M. agalactiae genomes (Dordet-Frisoni et al., 2014) or by whole genome sequencing (Illumina) (Dordet-Frisoni et al., 2019).
Table 1. Oligonucleotide used for the detection of antibiotic markers: annealing temperature and expected PCR product size (bp)

Table 2. Oligonucleotides used for transconjugant typing: sequences, annealing temperature and expected PCR products (bp) according to the strains
Recipes
SP4 liquid medium (for 1 L)
3.5 g PPLO broth
10 g Tryptone peptone
5.325 g Bacto peptone
690 ml distilled water
pH 7.8 (pH may be adjusted by the addition of NaOH 5 N solution)
This mixture should be autoclaved prior to adding:
170 ml decomplemented FBS (Fetal Bovine Serum, sterile solution)
50 ml CMRL 1066 (10x, sterile solution)
50 ml TC yeastolate (sol 4% in H2O, filtered 0.22 µm)
25 ml Yeast extract (15%, sterile solution)
10 ml D-(+)-Glucose (sol 50% in H2O, filtered 0.22 µm)
1 g Ampicillin
4 ml Phenol red, 0.5%
10 ml Sodium pyruvate (sol 0.5 M in H2O, filtered 0.22 µm)
SP4 agar medium
Proceed as for SP4 liquid medium but with some modifications
Before autoclaving add:
9 g select agar
After autoclaving
Do not add phenol red and sodium pyruvate
M. agalactiae Lysis buffer
1 ml Tris-HCl (1 M, pH 8.5)
5 µl Tween-20
125 µl Proteinase K 20 mg/ml
Add sterile ultrapure water up to a final volume of 10 ml
Acknowledgments
This work was supported by Grant ANR09MIE016 (MycXgene) from the French national funding research agency (ANR), by INRA and ENVT. The protocol is derived from works published in Dordet-Frisoni et al. (2013 and 2019).
Competing interests
The authors state that there are no competing interests
References
- Baranowski, E., Guiral, S., Sagne, E., Skapski, A. and Citti, C. (2010). Critical role of dispensable genes in Mycoplasma agalactiae interaction with mammalian cells. Infect Immun 78(4): 1542-1551.
- Calcutt, M. J., Lewis, M. S. and Wise, K. S. (2002). Molecular genetic analysis of ICEF, an integrative conjugal element that is present as a repetitive sequence in the chromosome of Mycoplasma fermentans PG. J Bacteriol 184(24): 6929-6941.
- Chazel, M., Tardy, F., Le Grand, D., Calavas, D. and Poumarat, F. (2010). Mycoplasmoses of ruminants in France: recent data from the national surveillance network. BMC Vet Res 6: 32.
- Dordet-Frisoni, E., Marenda, M. S., Sagne, E., Nouvel, L. X., Guerillot, R., Glaser, P., Blanchard, A., Tardy, F., Sirand-Pugnet, P., Baranowski, E. and Citti, C. (2013). ICEA of Mycoplasma agalactiae: a new family of self-transmissible integrative elements that confers conjugative properties to the recipient strain. Mol Microbiol 89(6): 1226-1239.
- Dordet-Frisoni, E., Faucher, M., Sagne, E., Baranowski, E., Tardy, F., Nouvel, L. X. and Citti, C. (2019). Mycoplasma Chromosomal Transfer: A Distributive, Conjugative Process Creating an Infinite Variety of Mosaic Genomes. Front Microbiol 10: 2441.
- Dordet-Frisoni, E., Sagne, E., Baranowski, E., Breton, M., Nouvel, L. X., Blanchard, A., Marenda, M. S., Tardy, F., Sirand-Pugnet, P. and Citti, C. (2014). Chromosomal transfers in mycoplasmas: when minimal genomes go mobile. mBio 5(6): e01958.
- Faucher, M., Nouvel, L. X., Dordet-Frisoni, E., Sagne, E., Baranowski, E., Hygonenq, M. C., Marenda, M. S., Tardy, F. and Citti, C. (2019). Mycoplasmas under experimental antimicrobial selection: The unpredicted contribution of horizontal chromosomal transfer. PLoS Genet 15(1): e1007910.
- Lederberg, J. and Tatum, E. L. (1946). Gene recombination in Escherichia coli. Nature 158(4016): 558.
- Marenda, M., Barbe, V., Gourgues, G., Mangenot, S., Sagne, E. and Citti, C. (2006). A new integrative conjugative element occurs in Mycoplasma agalactiae as chromosomal and free circular forms. J Bacteriol 188(11): 4137-4141.
- Meygret, A., Peuchant, O., Dordet-Frisoni, E., Sirand-Pugnet, P., Citti, C., Bebear, C., Beven, L. and Pereyre, S. (2019). High Prevalence of Integrative and Conjugative Elements Encoding Transcription Activator-Like Effector Repeats in Mycoplasma hominis. Front Microbiol 10: 2385.
- Rosengarten, R., Behrens, A., Stetefeld, A., Heller, M., Ahrens, M., Sachse, K., Yogev, D. and Kirchhoff, H. (1994). Antigen heterogeneity among isolates of Mycoplasma bovis is generated by high-frequency variation of diverse membrane surface proteins. Infect Immun 62(11): 5066-5074.
- Sirand-Pugnet, P., Citti, C., Barre, A. and Blanchard, A. (2007). Evolution of mollicutes: down a bumpy road with twists and turns. Res Microbiol 158(10): 754-766.
- Tardy, F., Mick, V., Dordet-Frisoni, E., Marenda, M. S., Sirand-Pugnet, P., Blanchard, A. and Citti, C. (2015). Integrative conjugative elements are widespread in field isolates of Mycoplasma species pathogenic for ruminants. Appl Environ Microbiol 81(5): 1634-1643.
- Zimmerman, C. U. and Herrmann, R. (2005). Synthesis of a small, cysteine-rich, 29 amino acids long peptide in Mycoplasma pneumoniae. FEMS Microbiol Lett 253: 315-321.
Article Information
Copyright
© 2021 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Sagné, E., Citti, C. and Dordet-Frisoni, E. (2021). Bacterial Conjugation Protocol for Ruminant Mycoplasmas . Bio-protocol 11(2): e3893. DOI: 10.21769/BioProtoc.3893.
Category
Microbiology > Microbial genetics > Transformation
Microbiology > Microbial physiology
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link








