- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Establishing an Adult Mouse Brain Hippocampal Organotypic Slice Culture System that Allows for Tracing and Pharmacological Manipulation of ex vivo Neurogenesis
Published: Vol 11, Iss 1, Jan 5, 2021 DOI: 10.21769/BioProtoc.3869 Views: 5972
Reviewed by: Narayan SubramanianShai BerlinAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Microfluidic Cultures of Basal Forebrain Cholinergic Neurons for Assessing Retrograde Cell Death by Live Imaging
Srestha Dasgupta [...] Wilma J. Friedman
Jan 5, 2025 1888 Views
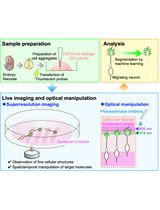
Time-Lapse Super-Resolution Imaging and Optical Manipulation of Growth Cones in Elongating Axons and Migrating Neurons
Masato Sawada [...] Kazunobu Sawamoto
Mar 20, 2025 2174 Views
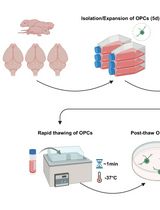
Cryopreservation of Bulk-Produced Primary Rat Oligodendrocyte Progenitor Cells
Hanki Kim [...] Jun Young Choi
Jun 20, 2025 1465 Views
Abstract
The function of the hippocampus depends on the process of adult hippocampal neurogenesis which underpins the exceptional neural plasticity of this structure, and is also frequently affected in CNS pathologies. Thus, manipulation of this process represents an important therapeutic goal. To identify potential strategies, organotypic adult brain slices are emerging as a valuable tool. Over the recent years, this methodology has been refined and here we present a combined protocol that brings together these refinements to enable long-term culture of adult hippocampal slices. We employ a sectioning technique that retains essential afferent inputs onto the hippocampus as well as serum-free culture conditions, so allowing an extended culture period. To sustain the neurogenic potential in the slices, we utilize the gliogenesis-inhibitor Indomethacin. Using EdU retention analysis enables us to assess the effects of pharmacological intervention on neurogenesis. With these improvements, we have established an easy and reliable method to study the effects of small molecules/drugs on proliferation and neuron formation ex vivo which will facilitate future discovery driven drug screenings.
Keywords: Adult hippocampal neurogenesisBackground
The hippocampus is a unique region of the brain with a high degree of plasticity as a result of the ongoing neurogenesis in the dentate gyrus throughout life. This process of adult hippocampal neurogenesis starts with the asymmetric division of neural stem cells (NSCs) in the subgranular zone (SGZ) that preserves the stem cell pool and generates progenitor cells poised for neuronal differentiation (Kempermann et al., 2004; Anacker and Hen, 2017; Toda and Gage, 2018). These latter cells go through a well-defined sequence of distinct stages: NSC-generated transiently amplifying progenitor cells divide rapidly and give rise to neuroblasts. Neuroblasts are characterized by their expression of the immature neuronal marker Doublecortin (Dcx). They exit the cell cycle to differentiate into immature post-mitotic neurons that in addition to Dcx transiently express the Calcium-binding protein Calretinin as well as the neuronal marker NeuN. In the final step, these newly generated neurons migrate and integrate functionally into the existing neuronal network as mature granule cell neurons which cease to express Dcx and Calretinin, but maintain NeuN expression.
Adult hippocampal neurogenesis plays a fundamental role in physiological CNS functions such as memory consolidation and cognitive flexibility. Thus, not surprisingly, it is implicated in pathologies like Alzheimer’s disease, depression, or schizophrenia (Winner and Winkler, 2015; Anacker and Hen, 2017; Moreno-Jimenez et al., 2019; Park, 2019) and therefore represents an important target for pharmaceutical intervention. However, drug testing is currently often based on cell culture systems that lack the complex architecture and connectivity of the intact brain (Pena, 2010; Humpel, 2015). On the other hand, drug testing in the intact brains of hundreds of animals is not practicable. Organotypic brain slices thus constitute a potential intermediate step as they maintain higher brain cyto-architecture and, as multiple sections can be obtained from one brain, enable comparison of different time points and/or drug concentrations with greatly reduced inter-animal variability. To study the effects of these drugs on neurogenesis, however, a reliable and easy lineage tracing approach is required.
Previously, Kim and colleagues established a protocol to culture adult mouse organotypic hippocampal slices for an extended period of four weeks by employing a serum-free culture medium (Kim et al., 2013). Moreover, they introduced a Hibernate-A based dissection medium as an alternative to more complicated media that required constant gassing with CO2. However, they were unable to trace the fate of single cells, nor did their slices maintain the essential afferent connection to the entorhinal cortex. Kleine Borgmann and co-workers used a different sectioning approach to overcome the latter problem and employed retrovirus labelling to study lineage progression in the SGZ (Kleine Borgmann et al., 2013). Here, we combined and refined these existing organotypic adult slice culture protocols. Moreover, we established EdU label tracing as a suitable read-out for neurogenesis. To circumvent the problem of a decreased neurogenic efficiency following a prolonged culture period, we utilized the Cyclooxygenase-inhibitor Indomethacin that has been shown before to reduce gliogenesis and exert protective effects on neurogenesis in vivo and ex vivo (Gerlach et al., 2016; Hain et al., 2018; Melo-Salas et al., 2018). Finally, establishing proof of concept as to the value of this protocol we compared the effects of Silychristin, iopanoic acid, and BCH on neurogenesis. These drugs are small molecular inhibitors of the monocarboxylate transporter 8 (Mct8), deiodinase type 2 (Dio2), and L-type amino acid transporters 1/2 (Lat 1/2) respectively–components of central thyroid hormone signalling. While neither iopanoic acid nor BCH treatment altered ex vivo neurogenesis, the number of new neurons was significantly reduced in Silychristin-treated cultures (Mayerl et al., 2020).
Taken together, these improvements enabled us to establish a method that allows for i) an extended culture period of adult brain slices for at least three weeks with good neurogenic efficiency, ii) easy lineage tracing using EdU that also provides a defined and controlled starting point, and iii) pharmacological manipulation of ex vivo hippocampal neurogenesis. We believe that our advanced methodology will be useful for future drug screening approaches to sustain or improve hippocampal neurogenesis in various pathological conditions or physiological alterations such as ageing.
Materials and Reagents
10 cm Petri dish (Thermo Fisher Scientific, Corning, catalog number: BP94A01 )
6-well-plate (Corning, catalog number: 3516 )
12-well-plate (Corning, catalog number: 3513 )
Tin foil
Microscope slides SuperFrost PlusTM (Thermo Fisher Scientific, catalog number: 10149870 )
Coverslips (e.g., 24 x 50 mm) (Thermo Scientific Menzel x1000 Coverslip 24 x 50 mm #1, catalog number: 15737592 )
Peel-A-Way® 22 x 40 mm x 20 mm deep, rectangular embedding molds (Ted Pella, catalog number: 27114 )
Millicell Cell Culture Insert, 30 mm, hydrophilic PTFE, 0.4 µm pore size (Merck, catalog number: PICMORG50 ), shelf storage
Mice (e.g., C57/Bl6N)
EdU (5-ethynyl-2-deoxyuridine) (Thermo Fisher Scientific, InvitrogenTM, catalog number: A10044 (50 mg), stored in 500 µl aliquots at -20 °C
HibernateTM-A Medium (Thermo Fisher Scientific, GibcoTM, catalog number: A1247501 ), stored at 4 °C for a maximum of 4 months
B-27TM supplement (50x) (Thermo Fisher Scientific, GibcoTM, catalog number: 17504001 ), aliquots stored at -20 °C, after defrosting stored at 4 °C for a maximum of 1 month
Agarose, low melting point (Merck, Calbiochem, catalog number: 2070-100GM ). 4% solution in PBS can be stored on shelf for up to 6 months and re-used 3-4x
NeurobasalTM-A Medium (Thermo Fisher Scientific, GibcoTM, catalog number: 10888022 ), stored at 4 °C for a maximum of 4 months
Indomethacin (Merck, Sigma-Aldrich, catalog number: I7378-5G ), stored at 4 °C as stock solution of 100 mM in DMSO for a maximum of 4 months
Silychristin (Merck, Sigma-Aldrich, catalog number: 51681-10MG ), stored at 4 °C as a stock solution of 100 mM in DMSO for a maximum of 4 months
Iopanoic acid (Merck, Sigma-Aldrich, catalog number: I0330000 ), stored at 4 °C as a stock solution of 100 mM in DMSO for a maximum of 4 months
BCH (R&D Systems, catalog number: 5027/50 ), stored at 4 °C as a stock solution of 100 mM in Neurobasal A for a maximum of 4 months
Formaldehyde 37% (Merck, Sigma-Aldrich, catalog number: 252549-500ML ), stored in a ventilated fume hood
Click-iT® EdU Alexa Fluor® 647 Imaging Kit (Thermo Fisher Scientific, Molecular Probes, catalog number: C10340 ), components diluted and stored according to manufacturer’s instructions
Rabbit-anti-Ki67 antibody (Abcam, catalog number: ab16667 ), aliquots stored at -20 °C
Guinea pig-anti-Doublecortin (Dcx) antibody (Merck, Millipore, catalog number: ab2253 ), aliquots stored at -20 °C
Mouse-anti-NeuN antibody (Merck, Millipore, catalog number: mab377 ), stored at 4 °C
Fluoromount-G® (Southern Biotech, catalog number: 0100-01 ), stored at room temperature
Super glue
Triton X-100 (Merck, Sigma-Aldrich, catalog number: X100-1L ), stored at room temperature
Glycine (Merck, Sigma-Aldrich, catalog number: G7126-1KG ), stored at room temperature
10x PBS (Thermo Fisher Scientific, Life Technologies, catalog number: 70011-036 500ml), stored at room temperature; dilute with distilled water
BSA (Merck, Sigma-Aldrich, catalog number: A7906-50G ), stored at 4 °C
Goat serum (Merck, Sigma-Aldrich, catalog number: G6767-100ML ), stored at -20 °C
Hoechst33258 (Thermo Fisher Scientific, Life Technologies, catalog number: 62249 5ML ), stored at 4 °C
L-Glutamine (Invitrogen, catalog number: 25030-024 ), aliquots stored at -20 °C
Penicillin/Streptomycin (10,000 U/ml Pen; 10,000 µg/ml Strep; Invitrogen, catalog number: 15140-122 ), aliquots stored at -20 °C
DMSO (Thermo Fisher Scientific, catalog number: D12345 3ML ), stored at room temperature
7 ml tubes (Sarstedt, Tube 7ml 47x20PC+Cap, catalog number: 71.9923.610 ), stored at room temperature
Dissection buffer (see Recipes)
Serum-free slice culture medium (see Recipes)
Equipment
Pipettes
Brush
Spatula (Thermo Fisher Scientific, FisherbrandTM Nickel Chattaway Spatula, catalog number: 11583462 "> 11583462 )
Scalpel (disposable scalpel, Swann Morton No. 11, Medisave, catalog number: SKU0503 )
Tissue culture hood
Water bath
Forceps (Fine Science Tools, Dumont #5 Forceps Standard Carbon, catalog number: 11251-10 )
Scissors (Fine Science Tools, Fine Scissors, Sharp, catalog number: 14060-10 )
Rocker
Confocal microscope (Leica SP8 )
Vibratome (Leica VT1000 S )
Software
ImageJ (NIH, https://imagej.nih.gov/ij/)
Procedure
Preparation of slices
For EdU label tracing studies, inject a 6-8-week old C57/Bl6N mouse i.p. with 100 µl EdU (10 mg/ml). To increase number of labeled cells, we injected EdU twice: 4 h and 2 h before sacrifice.
Prepare 4% low melting agarose in 1x PBS and keep it in a water bath at 37 °C until use.
Sacrifice the animal by exposing it to rising concentrations of CO2.
Isolate brain rapidly and transfer it into 2 ml serum-free dissection buffer (Kim et al., 2013) in a 7 ml tube on ice. Allow brain to equilibrate for at least 5 min.
Prepare for sectioning (Figure 1A): Transfer brain into a 10 cm Petri dish (Figure 1B), remove cerebellum and brainstem with a scalpel (Figure 1C) and cut along the midline of the brain (Figure 1D). According to the instructions provided in Kleine Borgmann et al. (2013) place hemispheres on the cut surface (Figure 1E) and remove a small part of the cerebral cortex cutting from dorsolateral to ventromedial at an angle of approximately 70° (Figures 1F, 1G). Flip the hemispheres onto these new cut surfaces and place them into an embedding mold on ice. Gently fill the mold with 4% low melting agarose and wait till it solidifies (Figure 1H). Trim the agarose block using a scalpel and glue it onto a vibratome tissue platform using super glue (Figure 1I). Hemispheres can be embedded, glued and cut separately or in parallel. We obtained the best results by cutting them in parallel oriented in a lateral-to-medial direction (i.e., the lateral part facing the vibratome and being cut first).
Transfer platform into the vibratome tissue container filled with chilled dissection buffer. To keep the temperature low, cover the container with ice (if possible) or store it on ice if not in use.
Cut 300 µm thick sections using high frequency (level 7-8) and low speed (level 1-2).
Carefully transfer sections containing the hippocampus into a 10 cm Petri dish containing 4-5 ml dissection buffer on ice using a brush and spatula (Figure 1J). Note examples of a well cut section with preserved brain cyto-architecture (Figure 1K) and a badly sectioned slice (Figure 1L). We did not trim the sections as we did not find any discernable difference in long term viability between trimming them so that they only contain the hippocampus and entorhinal cortex or culturing the entire sections.

Figure 1. Dissection procedure. (A) Schematic representation of brain sectioning. Panels B-L show photographs of the key steps, as follows. Place the brain in a Petri dish (B). Having cut away cerebellum and brainstem (C), bisect brain along the midline (D). Place hemispheres onto the new cut surfaces (E) and cut off a small part of the cerebral cortex in a dorsolateral to ventromedial direction at an angle of 70° (F). Flip onto the newest cut surface (G), embed in 4% low melting agarose (H), glue the block onto a tissue platform (I) and section hemispheres using a vibratome. Collect slices in a 10 cm Petri dish (J). Representative pictures of a well-cut slice suitable for culturing (K) and of a badly cut slice (L).
Slice culture
Before sacrificing the animal, place the required amount of Millicell inserts into 6-well-plate wells.
Prepare stock solutions. Dissolve Indomethacin in DMSO to obtain a 100 mM stock solution. Dissolve drugs in DMSO or medium.
Add 1 ml serum-free culture medium (Kim et al., 2013) containing Indomethacin and inhibitor (or solvent control) per well. Work in a tissue culture hood. Equilibrate at 37 °C and 5% CO2 for one to two hours in a standard cell culture incubator.
Add a small drop of culture medium (approx. 50 µl) from the well onto the insert using a standard 200 µl pipette to facilitate transfer of the sections. Using a brush and spatula transfer the vibratome sections from chilled dissection medium onto the Millicell inserts so that they are at the air/liquid-interface. Culture 2-3 sections per insert at 37 °C and 5% CO2.
On the next day, remove the culture medium containing inhibitors or solvent control completely using a 1 ml pipette. Keep the insert in the well and carefully raise it on one side using a pipette tip or forceps to allow for the removal of residual medium. Replace immediately with 1 ml of fresh medium (containing inhibitors) carefully pipetted into the well so that the sections on the insert are once again at the air/liquid interface. Repeat every other day.
Fixation
Fix slices by removing the culture medium and adding 1 ml 3.7% formaldehyde in PBS into the well and an additional 1 ml carefully on top of the slices. Fix for 1 h at ambient temperature.
Wash slices 3x 20 min in PBS.
Store the fixed slices in PBS at 4 °C or proceed with immuno-histochemistry.
Note: Slices may detach from the membrane during fixation. This does not affect further steps.
Immuno-histochemistry
Use forceps, scissors or a scalpel to cut the Millicell insert membrane from the plastic ring.
For EdU detection, transfer all slices from the same animal and condition (on the membrane or free-floating) into a 12-well-plate well and permeabilize them in PBS containing 0.1% Triton X100/0.1 M glycine. Block slices for 2 h in PBS containg 3% BSA. Incubate slices with 300 µl Click-iT reaction cocktail prepared according to the manufacturer’s instructions (Thermo Fisher Scientific Click-iT® EdU Alexa Fluor® 647 Imaging Kit, chapter EdU detection) at room temperature on a rocker for 3 h. For this and all subsequent steps protect the plate from light (e.g., by covering it with tin foil).
Block slices in 10% normal goat serum in PBS containing 0.5% Triton X100 overnight at 4 °C.
Prepare primary antibodies in blocking buffer (400 µl per well). Incubate slices at 4 °C on a rocker for 48 h. In our experiments, we used rabbit-anti Ki67 (1:250) to visualize proliferating cells, mouse-anti-NeuN (1:300) as a marker for neurons, and guinea pig-anti Dcx (1:300) to label neuroblasts and immature neurons
Wash slices 3x 1 h in PBS.
Incubate slices with secondary antibodies (1:500 in blocking buffer) on a rocker at 4 °C over night.
Remove secondary antibodies, add 400 µl Hoechst33258 (in water; 5 µg/ml) per well and incubate for 30 min.
Wash slices 3x 1 h in PBS.
Using forceps, transfer membranes (slices facing upwards) onto microscope slides or, in case of detached slices, use a brush and spatula.
Place the slices in the center of the slide. Add 200 µl Fluoromount-G onto the slices and gently apply a coverslip (e.g., 24 x 50 mm). Be careful not to squeeze the slices as they are extremely thick. Dry at 4 °C.
Imaging
Image slices using a confocal microscope. We employed a Leica SP8 confocal microscope and a 20x objective and imaged a 30-50 µm thick stack in the middle of the 300 µm thick slice.
Data analysis
For data analysis, use the freely available software ImageJ/Fiji. Enumerate marker positive cells (Ki67, Dcx, EdU, NeuN) in the SGZ in one picture every 10 µm from the z-stack taken and normalize these numbers to the length of the SGZ (measured using ImageJ’s segmented line tool) for quantification (Figure 2). For nuclear markers (Ki67, EdU, NeuN), ascertain that there is a clear overlay with Hoechst33258 staining. Average results from all slices that were obtained from the same animal and cultured under the same condition. In our studies, we quantified EdU incorporation into different cell populations at different time points and then determined statistical significance by unpaired two-tailed Student’s t-test using slices from 4-6 animals per time point and condition. Depending on the focus of the analysis, one-way or two-way ANOVA may be required, e.g., for longitudinal analysis of the same parameter such as proliferative capacity. To view our results, see Mayerl et al. (2020).
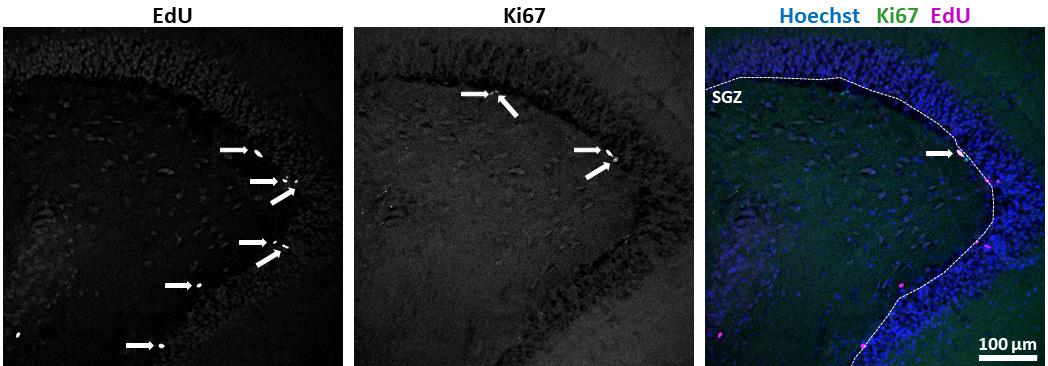
Figure 2. Quantification of results. Marker single or double positive cells along the SGZ were enumerated (arrows). Length of the SGZ was measured using ImageJ’s segmented line tool (dotted line). Quotients of positive cell numbers over length were calculated. (scale bar = 100 µm)
Notes
Note that sacrificing the animal by overexposure to CO2 can be accompanied by a reduction in brain tissue pH and increase in tissue stiffness (Holtzmann et al., 2016). Whether this change in stiffness alters the quality of the sectioning process in comparison to standard methods of euthanasia such as cervical dislocation or decapitation has not been investigated and may be determined individually.
While extracting the brain from the skull be careful not to cut into the brain tissue. Smaller, superficial cuts do not compromise the procedure, but larger incisions will result in cutting problems and ultimately lead to fractured and unsuitable slices. If the latter happens, do not proceed. Likewise, when preparing brain hemispheres for sectioning, take care not to pierce the tissue while trying to hold it in place with forceps. Better to use a blunt spatula to fix the tissue for cutting with the scalpel. When removing a small part of the cortex at a 70° angle you should be able to perceive a small strip of white matter in the removed part (see Figure 1G). In a few cases, brain hemispheres may not be cut as desired and the slices may be damaged (for comparison see Figures 1K and 1L). Problems can include ripped, fractured sections, severing of the hippocampus-cortex connection and visibly unequal thickness over the slice. The latter is common if agarose penetrates underneath a hemisphere while solidifying. As a result, brain tissue will not be glued to the vibratome tissue platform and is free to move if pressure from the blade is applied. If problems occur during cutting our experience is that trouble-shooting possibilities are very limited. We recommend repeating the procedure with another animal instead of using damaged slices.
One of the major points that needs considering is the thickness of the slices that–unlike neonatal slices–do not noticeably flatten over the culture period. Thus, the slices produce a pronounced background fluorescent intensity which will be a problem for some antibodies. In our hands, well-established, highly specific antibodies worked reliably, while other less specific antibodies did not give staining that could be used for quantifications or did not work at all. Moreover, we usually imaged on the day after mounting. Delayed imaging, however, may lead to the formation of air-filled gaps in the mounting medium that may dry out the sections. Thus, for longer storage before imaging sealing the coverslip e.g. with nail varnish constitutes a feasible option though we never further explored this possibility.
We never investigated whether the first block step (3% BSA in PBS) is sufficient and the second block step using 10% goat serum can be skipped. This likely depends on the performance of the antibodies employed.
Moreover, the protocol conditions are optimized for adult stem cells and neurons (hence using Hibernate A, Neurobasal A and Indomethacin) while they might be less efficient in keeping glia cells alive. Therefore, we do not recommend this protocol for the analysis of glia cells unless the culture conditions are further optimized.
Recipes
Following buffer recipes are based on Kim et al. (2013):
Dissection buffer (100 ml)
96 ml Hibernate A
2 ml B-27TM supplement (50x)
1 ml 200 mM L-Glutamine
1 ml Penicillin/Streptomycin
Serum-free slice culture medium (100 ml)
96 ml Neurobasal A
2 ml B-27TM supplement (50x)
1 ml 200 mM L-Glutamine
1 ml Penicillin/Streptomycin
80 µl 100 mM Indomethacin
Acknowledgments
This work was supported by a grant from the DFG to SM (MA7212/ 2-1), by a Wellcome Trust Senior Investigator award (104783/Z/14/Z) and funding from the BBSRC (BB/L00402X/1) to CffC. We thank SCRM animal facility staff (Luke McPhee, Chris Wilson, Lorraine McNeil, John Agnew, and Jamie Kelly) and SCRM imaging facility staff (Bertrand Vernay) for their excellent help. The protocol was first published in our original research paper “Hippocampal Neurogenesis Requires Cell-Autonomous Thyroid Hormone Signaling” in Stem Cell Reports (Mayerl et al., 2020).
Competing interests
The authors declare that no conflict of interests exists.
Ethics
All studies were executed under Project Licence PPL 70/8748, valid from 6 Dec 2015 to 5 Dec 2020, in compliance with UK Home Office regulations and local guidelines by The University of Edinburgh.
References
- Anacker, C. and Hen, R. (2017). Adult hippocampal neurogenesis and cognitive flexibility- linking memory and mood. Nat Rev Neurosci 18(6): 335-346.
- Gerlach, J., Donkels, C., Münzner, G. and Haas, C. A. (2016). Persistent gliosis interferes with neurogenesis in organotypic hippocampal slice cultures. Front Cell Neurosci 10: 131.
- Hain, E. G., Sparenberg, M., Rasinska, J., Klein, C., Akyuz, L. and Steiner, B. (2018). Indomethacin promotes survival of new neurons in the adult murine hippocampus accompanied by anti-inflammatory effects following MPTP-induced dopamine depletion. J Neuroinflammation 15(1): 162.
- Holtzmann, K., Gautier, H. O. B., Christ, A. F., Guck, J., Káradóttir, R. T. and Franze, K. (2016). Brain tissue stiffness is a sensitive marker for acidosis. J Neurosci Methods 271: 50-54.
- Humpel, C., (2015). Organotypic brain slice cultures: A review. Neuroscience 305: 86-98.
- Kempermann, G., Jessberger, S., Steiner, B. and Kronenberg, G. (2004). Milestones of neuronal development in the adult hippocampus. Trends Neurosci 27(8): 447-452.
- Kim, H., Kim, E., Park, M., Lee, E. and Namkoong, K. (2013). Organotypic hippocampal slice culture from the adult mouse brain: a versatile tool for translational neuropsychopharmacology. Prog Neuropsychopharmacol Biol Psychiatry 41: 36-43.
- Kleine Borgmann, F. B., Bracko, O. and Jessberger, S. (2013). Imaging neurite development of adult-born granule cells. Development 140(13): 2823-2827.
- Mayerl, S., Heuer, H. and ffrench-Constant, C. (2020). Hippocampal neurogenesis requires cell-autonomous thyroid hormone signaling. Stem Cell Reports 14(5): 845-860.
- Melo-Salas, M. S., Perez-Dominguez, M. and Zepeda, A. (2018). Systemic inflammation Impairs proliferation of hippocampal type 2 intermediate precursor cells. Cell Mol Neurobiol 38(8): 1517-1528.
- Moreno-Jimenez, E. P., Flor-Garcia, M., Terreros-Roncal, J., Rabano, A., Cafini, F., Pallas-Bazarra, N., Avila, J. and Llorens-Martin, M. (2019). Adult hippocampal neurogenesis is abundant in neurologically healthy subjects and drops sharply in patients with Alzheimer's disease. Nat Med 25(4): 554-560.
- Park, S. C. (2019). Neurogenesis and antidepressant action. Cell Tissue Res 377(1): 95-106.
- Pena, F. (2010). Organotypic cultures as tool to test long-term effects of chemicals on the nervous system. Curr Med Chem 17(10): 987-1001.
- Toda, T. and Gage, F. H. (2018). Review: adult neurogenesis contributes to hippocampal plasticity. Cell Tissue Res 373(3): 693-709.
- Winner, B. and Winkler, J. (2015). Adult neurogenesis in neurodegenerative diseases. Cold Spring Harb Perspect Biol 7(4): a021287.
Article Information
Copyright
© 2021 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Mayerl, S. and ffrench-Constant, C. (2021). Establishing an Adult Mouse Brain Hippocampal Organotypic Slice Culture System that Allows for Tracing and Pharmacological Manipulation of ex vivo Neurogenesis. Bio-protocol 11(1): e3869. DOI: 10.21769/BioProtoc.3869.
Category
Neuroscience > Cellular mechanisms > Cell isolation and culture
Stem Cell > Adult stem cell
Cell Biology > Cell-based analysis > Drug screenings
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link