- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Preparation of Bacterial Outer Membrane Vesicles for Characterisation of Periplasmic Proteins in Their Native Environment
Published: Vol 10, Iss 24, Dec 20, 2020 DOI: 10.21769/BioProtoc.3853 Views: 4602
Reviewed by: David PaulElías Barquero-CalvoAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
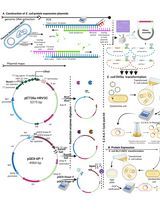
Thermus thermophilus CRISPR Cas6 Heterologous Expression and Purification
Junwei Wei [...] Yingjun Li
Jul 20, 2025 2202 Views
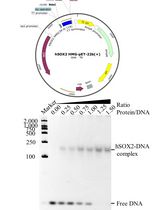
Prokaryotic Expression and Purification of the hSox2-HMG Domain
Lijie Yang [...] Jingjun Hong
Aug 20, 2025 2439 Views
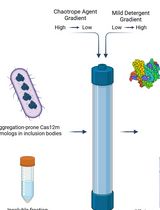
On-Column Dual-Gradient Refolding for Efficient Recovery of Insoluble Affinity-Tagged Recombinant Proteins
Anna Vlaskina [...] Maxim Patrushev
Feb 5, 2026 187 Views
Abstract
Bacterial outer membrane vesicles (OMVs) are naturally formed by budding from the outer membrane of Gram-negative bacteria. OMVs consist of a lipid bilayer identical in composition to the original outer membrane and contain periplasmic content within their lumen. Enriched with specific envelope proteins, OMVs make for an excellent native-like platform to study these proteins in-situ using biophysical methods. Here, we describe in detail the preparation of OMVs from Escherichia coli, which are luminally enriched with periplasmic proteins and uniformly labeled with stable isotopes (2H and 15N), suitable for the subsequent characterisation of proteins at atomic resolution in their native environment by solution-state NMR spectroscopy. The ability to perform structural studies of periplasmic components in-situ clears the way to reaching an in-depth understanding of the functional and mechanistic details of this unique cellular compartment.
Keywords: Outer membrane vesiclesBackground
The periplasm of Gram-negative bacteria is a quite remarkable cellular compartment. This space, incarcerated between the inner and outer bacterial membrane, contains proteins at an extraordinarily high concentration exceeding 300 mg ml−1 (Oliver, 1996) and, in the absence of cellular sources such as ATP, functions almost energetically independent from its cytosolic counterpart. So far, structural knowledge about periplasmic proteins has been exclusively obtained using purified proteins isolated from their native environment. Thus, any structural and functional influence this special environment might impose on a protein are lost during purification. Efforts to study periplasmic proteins in-situ using biophysical methods such as in-cell NMR spectroscopy have been hampered by the low volume ratio of the periplasm, which contributes only 5-10% to the total bacterial volume (Brass et al., 1986).
We recently described a new method to characterize proteins within bacterial outer membrane vesicles (OMVs) using solution-state NMR spectroscopy (Thoma and Burmann, 2020). OMVs, which are naturally released by Gram-negative bacteria, contain periplasmic content in their lumen and can be non-destructively purified from bacterial growth cultures (Chutkan et al., 2013; Schwechheimer and Kuehn, 2015). Thus, OMVs represent an excellent platform to study periplasmic proteins, enriched in the luminal space, together with their natural environment. To be amendable for NMR studies, OMVs are uniformly labeled with stable isotopes (2H and 15N) and prepared in large quantities. Our approach proved particularly useful for the study of soluble periplasmic proteins within this environment, as the membrane components of OMVs remain largely invisible to solution-state NMR, due to slower molecular tumbling rates.
One key factor for obtaining large amounts of luminally enriched OMVs is the use of a suitable expression strain. Widely used expression hosts such as E. coli BL21(DE3) do not produce sufficient amounts of OMVs to be feasible for these types of experiments (Thoma et al., 2018). In contrast, strains lacking the major outer membrane protein OmpA have been shown to have a 26-fold increased OMV production (Schwechheimer et al., 2014). Moreover, deletion of OmpA omits possible background signals originating from its soluble peptidoglycan binding domain in NMR experiments (Thoma and Burmann, 2020). In our experimental set-up we relied on the strain BL21(DE3)ompA, which was kindly provided by Guido Grandi (Fantappiè et al., 2014) and which reproducibly yielded high amounts of luminally enriched OMVs. However, other alternatives such as BL21A are available (Meuskens et al., 2017) or could easily be tailor-made by deletion of the ompA gene in the expression strain of choice.
Another critical factor for obtaining large amounts of suitable OMVs is the time point, at which the bacteria are removed from the growth media. Bacterial growth should be maintained as long as possible, since the amount of OMVs present in the culture supernatant correlates directly with the cell density and consequentially with the length of the growth period. However, OMVs should be collected before bacterial growth ceases and cultures enter the stationary phase, because this will substantially increase contaminations with cytoplasmic proteins, inner membranes, and cell debris resulting from stress-induced explosive cell lysis (Turnbull et al., 2016; Thoma et al., 2018). Depending on whether cells are grown in H2O or D2O based media this stage is typically achieved after 3 to 5 h, respectively. Since bacterial growth is not only retarded by the use of D2O but also underlies various influences such as the strain and plasmid used for expression, it is advisable to determine the growth behavior of the bacterial cultures prior to OMV preparation. Whereas deuteration is not strictly necessary for small to medium sized proteins (< 25 kDa), luminally enriched OMVs prepared from cells grown in D2O based media typically yield NMR spectra of higher quality due to a decrease in background signals and a smaller NMR line-width (Thoma and Burmann, 2020).
Materials and Reagents
0.22 µm vacuum filtration unit (such as Steritop Quick Release, EMD Millipore, catalog number: S2GPT05RE )
0.45 µm vacuum filtration unit (such as Stericup-HV, EMD Millipore, catalog number: SCHVU11RE )
≥ 50 ml syringe (such as Fisherbrand Sterile Syringes, Thermo Fisher Scientific, catalog number: 15899152 )
0.45 µm syringe filter (such as Millex-HP, EMD Millipore, catalog number: SLHP033R )
Microscale NMR tubes (such as 5mm Shigemi Tube Set, Shigemi, catalog number: BMS-005B )
Tryptone (Sigma-Aldrich, catalog number: T9410 )
Yeast Extract (Sigma-Aldrich, catalog number: Y1625 )
NaCl (Sigma-Aldrich, catalog number: S7653 )
Na2HPO4 (Sigma-Aldrich, catalog number: S7907 )
KH2PO4 (Sigma-Aldrich, catalog number: P0662 )
KCl (Sigma-Aldrich, catalog number: P9333 )
CaCl2·2H2O (Sigma-Aldrich, catalog number: C5080 )
MgCl2·6H2O (Sigma-Aldrich, catalog number: M2670 )
15NH4Cl (Sigma-Aldrich, catalog number: 299251 )
D2O (Sigma-Aldrich, catalog number: 617385 )
D-Glucose (Sigma-Aldrich, catalog number: G7528 )
MgSO4 (Sigma-Aldrich, catalog number: M7506 )
Thiamine (Sigma-Aldrich, catalog number: T1270 )
Isopropyl-β-D-thiogalactoside (IPTG; Sigma-Aldrich, catalog number: I6758 )
LB-medium (Lennox) (see Recipes)
M9 medium (see Recipes)
Dulbeccos phosphate buffered saline with added salts (DPBSS)/NMR sample buffer (see Recipes)
Equipment
Shaking incubator (such as New Brunswick Innova® 42, Eppendorf AG, catalog number: M1335-0012 )
500 ml baffled culture flask (such as ROTILABO baffled flasks, Carl Roth, catalog number: LY96.1 )
5,000 ml baffled culture flask (such as ROTILABO baffled flasks, Carl Roth, catalog number: PK29.1 )
Cell density meter (such as GE Healthcare Ultrospec, Thermo Fisher Scientific, catalog number: 10704417 )
High speed centrifuge (such as Avanti J-series, Beckman Coulter, catalog number: B22989 )
High speed large-volume fixed angle rotor (such as J-LITE JLA-16.250, Beckman Coulter, catalog number: 363934) equipped with suitable centrifugation bottles (such as 250 ml Polycarbonate Bottle, Beckman Coulter, catalog number: 356013)
High speed small-volume fixed angle rotor (such as JA-25.50, Beckman Coulter, catalog number: 363055) equipped with suitable centrifugation tubes (such as 50 ml Polycarbonate Bottle, Beckman Coulter, catalog number: 357000)
Vacuum pump, sufficiently strong for use with vacuum filtration units (ultimate vacuum ~0.01 mPa)
Vortex mixer (such as Vortex-Genie 2, Scientific Industries, catalog number: SI-0246)
Microvolume spectrophotometer (such as Nanodrop One, Thermo Fisher Scientific, ND-ONE-W)
Gel electrophoresis system for SDS page (such as Mini-PROTEAN Tetra, Bio-Rad, catalog number: 1658004) including power supply, polyacrylamide gels, and necessary buffers and staining solutions (refer to manufacturers specifications for details)
Software
Any spreadsheed or scientific graphing software allowing basic calculations (Excel, Numbers, LabPlot, Origin, etc.)
Procedure
Gene cloning and transformation
The complete gene coding for the protein of interest including its preceding periplasmic export signal should be cloned into a suitable plasmid backbone of the pET expression system (such as pET21a). Thereby, additional features such as affinity tags are not necessary and should be omitted. The final plasmid should be transformed into an OMV-overproducing strain such as BL21(DE3)ompA (Fantappiè et al., 2014). These steps can be carried out following standard molecular biology techniques. The description of detailed cloning and transformation procedures lies outside the scope of this protocol.
Preparation of growth media and buffers
Per sample requirements are: 5 ml LB medium, 1 L M9 medium, 25 ml DPBSS, 0.2 ml NMR sample buffer (see Recipes for formulations). Supplement growth media with appropriate antibiotics for selection of the strain/plasmid used for expression. When using the pET expression system, use Ampicillin to a final concentration of 100 µg ml-1 or Kanamycin to a final concentration of 50 µg ml-1. Stock solutions of antibiotics (Ampicillin: 100 mg ml-1; Kanamycin: 50 mg ml-1) and IPTG (1 M) can be prepared in H2O/D2O, corresponding to the solvent used for growth media.
Bacterial growth
Inoculate 5 ml LB medium with a single colony.
Grow shaking at 37 °C for 10 h.
Inoculate 100 ml M9 in a 500 ml baffled culture flask using 1 ml of the culture grown in LB medium.
Grow shaking at 37 °C overnight (~80 rpm, 25 mm orbit).
Add the entire preculture to 900 ml M9 in a 5,000 ml baffled culture flask.
Grow shaking at 37 °C (~80 rpm, 25 mm orbit).
Check the optical density OD600 every 30 min.
When the optical density reaches OD600 ~0.4 add IPTG to a final concentration of 0.2 mM to induce protein expression.
Continue to monitor the optical density OD600 every 30 min.
When bacterial growth ceases (after ~3 h in H2O/~5 h in D2O based media, see Figure 1) remove the cells by centrifugation at 10,000 x g for 10 min at 10 °C.
Filter the supernatant with 0.45 µm vacuum filtration units. The filtered supernatant can be stored at 4 °C for up to 24 h.

Figure 1. Media-dependent growth behavior. Exemplary growth curves in H2O based (dark blue) and D2O based (light blue) M9 minimal media. Triangles mark the time of induction, squares mark the time of harvest.
OMV Preparation
Collect OMVs from the filtered supernatant by centrifugation at 38,400 x g for 2 h at 10 °C. After centrifugation, centrifuge bottles should contain a small but clearly visible whitish, slightly cloudy pellet (Figure 2A).
Immediately discard the supernatant (OMV pellets will begin to suspend shortly).
Resuspend OMV pellets in a total volume of 25 ml DPBSS by pipetting.
Filter the OMV suspension with a 0.45 µm syringe filter.
Collect OMVs by centrifugation using a single centrifuge tube at 74,500 x g for 45 min at 10 °C (Figure 2B).
Immediately discard the supernatant.
Resuspend the OMV pellet in 200 µl NMR sample buffer by gentle vortexing.
Save 2 µl of the OMV suspension for subsequent analysis and transfer the remaining suspension to a microscale NMR tube (Figure 2C). OMV suspensions are in our hands stable at 4 °C for up to 2 weeks.

Figure 2. Preparation of OMVs. OMV pellets after the first (A) and second centrifugation (B), and resuspended in NMR buffer in a 5 mm Shigemi NMR tube (C).
OMV Analysis
Dilute 2 µl of the final OMV suspension with 18 µl DPBSS.
Record a UV-Vis spectrum of the dilute OMV suspension using a microvolume spectrophotometer (range: 220-360 nm) and export the spectrum data for further analysis.
Correct the exported spectrum for the light-scattering contribution of the OMVs and estimate the protein concentration from the absorbance at 280 nm (see Data analysis and Figure 3A). OMV preparations should contain >15 mg/ml total protein to yield NMR spectra of good quality.
Prepare two samples for SDS page using denaturing Laemmli Sample Buffer, containing ~6 µg total protein each.
Heat-denature one of the samples at 98 °C for 15 min, keep the second sample at room temperature.
Analyze samples by SDS page. Samples of protein-enriched OMVs should result in two prominent bands. One band corresponding to the major porins OmpF/OmpC present in the OMV membrane runs at 37 kDa in the heat-denatured sample and results in a contorted, ladder-like pattern between 60 and 70 kDa in the non-heated sample (Figure 3B). A second band, usually not subject to heat-modifiability, should be present at the weight of the overexpressed protein of choice, indicating successful enrichment in the OMVs (Figure 3B).

Figure 3. Analysis of OMVs by spectrophotometry and SDS-page. A. Exemplary UV-Vis spectrum of a dilute OMV suspension before (dashed line) and after light-scattering correction (solid line). B. Exemplary SDS page gel of OMVs enriched with a periplasmic protein (dark blue triangle). Major porins OmpF/OmpC are marked by the light blue open triangles.
Data analysis
The spectrophotometric estimation of the protein concentration in a sample based on absorbance measurement at a wavelength of 280 nm is a simple, fast, and routinely used method. However, UV-Vis spectra recorded of OMVs contain a strong light-scattering component. In order to estimate the protein concentration within an OMV sample, light-scattering due to the vesicles needs to be corrected. A light-scattering corrected UV-Vis spectrum can be obtained using the following equation, originally used to estimate the nucleic acid content of viruses (Porterfield and Zlotnick, 2010):

The total protein concentration in the OMV sample can be estimated from the corrected value of A280, assuming 1 Abs = 1 mg ml-1 for the mixture of proteins present in OMVs. Due to the presence of nucleic acids in OMVs (Brown et al., 2015), which strongly influences the absorbance at 280 nm, it is useful to estimate the amount of nucleic acids in the sample from the A260/A280 ratio of the light-scattering corrected spectrum, which can be done using the following empirical equation (Glasel, 1995):

Typically, protein-enriched OMVs contain ≤ 3% nucleic acids. If nucleic acids are detected ([%N] > 0), the total protein concentration in the OMV sample can be estimated from the corrected values of A280 and A260, using the following equation (Layne, 1957):

Notes
Typically a sufficient amount of enriched OMVs can be obtained from a growth culture volume of one liter using the culture conditions described above. Nevertheless, it should be noted that bacterial growth in D2O based media can be less reproducible than in H2O based media, occasionally resulting in irregular growth behavior and/or lower yields. Moreover, it should be noted that the overexpression of proteins in the periplasmic space can result in cytotoxicity (Schlegel et al., 2013), which might require specific adaption of the induction levels to the expression system used.
Recipes
LB-medium (Lennox)
10 g/L Tryptone
5 g/L Yeast extract
5 g/L NaCl
Dissolve in H2O and sterilize by autoclaving
M9 medium
6 g/L Na2HPO4
3 g/L KH2PO4
1 g/L 15NH4Cl
0.5 g/L NaCl
2 g/L D-Glucose
1 ml/L CaCl2 (0.1 M) solution*
1 ml/L MgSO4 (1 M) solution*
0.1 ml/L Thiamine (10 mg/ml) solution*
Dissolve solid components in H2O/D2O, add salt/Thiamine solutions and sterilize by filtration using 0.22 µm vacuum filtration units.
*Note: Prepare stock solutions in H2O/D2O, respectively.
Dulbeccos phosphate buffered saline with added salts (DPBSS)/NMR sample buffer:
1.15 g/L Na2HPO4
0.2 g/L KH2PO4
8 g/L NaCl
0.2 g/L KCl
0.45 ml/L CaCl2 (1 M) solution
0.25 ml/L MgCl2 (1 M) solution
Dissolve solid components in H2O*, add salt solutions and sterilize by filtration using 0.22 µm vacuum filtration units.
*Note: For NMR sample buffer, dissolve in H2O to 90% final volume, then add 10% D2O.
Acknowledgments
We thank G. Grandi (University of Trento) for providing E. coli strain BL21(DE)ΔompA. J.T. was supported by an EMBO Long-Term Fellowship (ALTF 413-2018). B.M.B. gratefully acknowledges funding from the Swedish Research Council (Vetenskapsrådet Starting Grant 2016-04721) and the Knut och Alice Wallenberg Foundation through a Wallenberg Academy Fellowship (2016.0163) as well as through the Wallenberg Centre for Molecular and Translational Medicine, University of Gothenburg, Sweden.
This protocol was originally described in Thoma and Burmann (2020).
Competing interests
The authors declare no competing interest.
References
- Brass, J. M., Higgins, C. F., Foley, M., Rugman, P. A., Birmingham, J. and Garland, P. B. (1986). Lateral diffusion of proteins in the periplasm of Escherichia coli. J Bacteriol 165(3): 787-795.
- Brown, L., Wolf, J. M., Prados-Rosales, R. and Casadevall, A. (2015). Through the wall: extracellular vesicles in Gram-positive bacteria, mycobacteria and fungi. Nat Rev Microbiol 13(10): 620-630.
- Chutkan, H., Macdonald, I., Manning, A. and Kuehn, M. J. (2013). Quantitative and qualitative preparations of bacterial outer membrane vesicles. Methods Mol Biol 966: 259-272.
- Fantappiè, L., de Santis, M., Chiarot, E., Carboni, F., Bensi, G., Jousson, O., Margarit, I. and Grandi, G. (2014). Antibody-mediated immunity induced by engineered Escherichia coli OMVs carrying heterologous antigens in their lumen. J Extracell Vesicles 3.
- Glasel, J. A. (1995). Validity of nucleic acid purities monitored by 260nm/280nm absorbance ratios. Biotechniques 18(1): 62-63.
- Layne, E. (1957). Spectrophotometric and Turbidimetric Methods for Measuring Proteins. In: Colowick, P. S. and Kaplan, N.O. (Eds.). Method in Enzymology. Academic Press, Inc., New York, 447-454.
- Meuskens, I., Michalik, M., Chauhan, N., Linke, D. and Leo, J. C. (2017). A new strain collection for improved expression of outer membrane proteins. Front Cell Infect Microbiol 7: 464.
- Oliver, D. B., (1996). Periplasm. In. Escherichia coli and Salmonella p. 26. Available at: papers3://publication/uuid/D3FF17F1-8B39-4CD7-B786-D17CA6614412.
- Porterfield, J. Z. and Zlotnick, A. (2010). A simple and general method for determining the protein and nucleic acid content of viruses by UV absorbance. Virology 407(2): 281-288.
- Schlegel, S., Rujas, E., Ytterberg, A. J., Zubarev, R. A., Luirink, J. and de Gier, J. W. (2013). Optimizing heterologous protein production in the periplasm of E. coli by regulating gene expression levels. Microb Cell Fact 12:24.
- Schwechheimer, C. and Kuehn, M. J. (2015). Outer-membrane vesicles from Gram-negative bacteria: biogenesis and functions. Nat Rev Microbiol 13(10): 605-619.
- Schwechheimer, C., Kulp, A. and Kuehn, M. J. (2014). Modulation of bacterial outer membrane vesicle production by envelope structure and content. BMC Microbiol 14: 324.
- Thoma, J., Manioglu, S., Kalbermatter, D., Bosshart, P. D., Fotiadis, D. and Müller, D. J. (2018). Protein-enriched outer membrane vesicles as a native platform for outer membrane protein studies. Commun Biol 1: 23.
- Thoma, J. and Burmann, B. M. (2020). High-resolution in situ NMR spectroscopy of bacterial envelope proteins in outer membrane vesicles. Biochemistry 59(17): 1656-1660.
- Turnbull, L., Toyofuku, M., Hynen, A. L., Kurosawa, M., Pessi, G., Petty, N. K., Osvath, S. R., Carcamo-Oyarce, G., Gloag, E. S., Shimoni, R., Omasits, U., Ito, S., Yap, X., Monahan, L. G., Cavaliere, R., Ahrens, C. H., Charles, I. G., Nomura, N., Eberl, L. and Whitchurch, C. B. (2016). Explosive cell lysis as a mechanism for the biogenesis of bacterial membrane vesicles and biofilms. Nat Commun 7: 11220.
Article Information
Copyright
© 2020 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Thoma, J. and Burmann, B. M. (2020). Preparation of Bacterial Outer Membrane Vesicles for Characterisation of Periplasmic Proteins in Their Native Environment. Bio-protocol 10(24): e3853. DOI: 10.21769/BioProtoc.3853.
Category
Biochemistry > Protein > Isolation and purification
Microbiology > Microbial biochemistry > Protein
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










