- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Quantification of Methylglyoxal Levels in Cowpea Leaves in Response to Cowpea Aphid Infestation
Published: Vol 10, Iss 20, Oct 20, 2020 DOI: 10.21769/BioProtoc.3795 Views: 3530
Reviewed by: Satyabrata NandaSaumik BasuAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
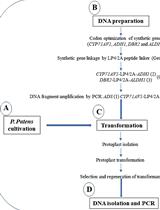
Heterologous Production of Artemisinin in Physcomitrium patens by Direct in vivo Assembly of Multiple DNA Fragments
Nur Kusaira Khairul Ikram [...] Henrik Toft Simonsen
Jul 20, 2023 2360 Views
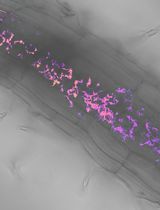
Utilizing FRET-based Biosensors to Measure Cellular Phosphate Levels in Mycorrhizal Roots of Brachypodium distachyon
Shiqi Zhang [...] Maria J. Harrison
Jan 20, 2025 2432 Views
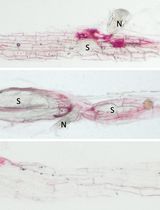
Image-Based Lignin Detection in Nematode-Induced Feeding Sites in Arabidopsis Roots
Muhammad Amjad Ali and Krzysztof Wieczorek
May 5, 2025 1841 Views
Abstract
Aphids are a serious pest of crops across the world. Aphids feed by inserting their flexible hypodermal needlelike mouthparts, or stylets, into their host plant tissues. They navigate their way to the phloem where they feed on its sap causing little mechanical damage to the plant. Additionally, while feeding, aphids secrete proteinaceous effectors in their saliva to alter plant metabolism and disrupt plant defenses to gain an advantage over the plant. Even with these arsenals to overcome plant responses, plants have evolved ways to detect and counter with defense responses to curtail aphid infestation. One of such response of cowpea to cowpea aphid infestation, is accumulation of the metabolite methylglyoxal. Methylglyoxal is an α,β-dicarbonyl ketoaldehyde that is toxic at high concentrations. Methylglyoxal levels increase modestly after exposure to a number of different abiotic and biotic stresses and has been shown to act as an emerging defense signaling molecule at low levels. Here we describe a protocol to measure methylglyoxal in cowpea leaves after cowpea aphid infestation, by utilizing a perchloric acid extraction process. The extracted supernatant was neutralized with potassium carbonate, and methylglyoxal was quantified through its reaction with N-acetyl-L-cysteine to form N-α-acetyl-S-(1-hydroxy-2-oxo-prop-1-yl)cysteine, a product that is quantified spectrophotometrically.
Keywords: MethylglyoxalBackground
The importance of methylglyoxal in plant response and signaling to various stresses is only just beginning to be understood. Various abiotic stresses have been shown to lead to an accumulation of methylglyoxal in plants (Yadav et al., 2005; Borysiuk et al., 2018). This accumulation suggests that methylglyoxal has a signaling role in plants (Hossain et al., 2009; Hoque et al., 2016; Mostofa et al., 2018). Exogenous application of methylglyoxal has been found to upregulate antioxidant and defense genes in plants corroborating the role methylglyoxal has as a signaling molecule (Kaur et al., 2015; Li et al., 2017). The accumulation of methylglyoxal in plants has also been described in response to biotic stresses including bacterial, viral and fungal infections (Melvin et al., 2017). Recently, it has been shown that cowpea aphid infestation also leads to an increase in methylglyoxal level expanding its role in defense against herbivore pests (MacWilliams et al., 2020). Three methods have been established for quantification of methylglyoxal. Of these three methods, the N-acetyl-L-cysteine method by Wild et al. (2012) has been found to measure methylglyoxal in the most economical and safest way. The other two methods involve expensive enzyme purification or derivatization with an explosive chemical (Racker, 1951; Gilbert and Brandt, 1975). N-acetyl-L-cysteine method involves mixing methylglyoxal with N-acetyl-L-cysteine to generate N-α-acetyl-S-(1-hydroxy-2-oxo-prop-1-yl)cysteine which is detected and measured at absorbance 288 nm by a spectrophotometer (Figure 1). In this protocol, we combine the method by Wild et al. (2012), that simply measures methylglyoxal levels, with extraction of methylglyoxal from plant tissues, and measuring its levels in cowpea (Vigna unguiculata) leaves after cowpea aphid infestation (Aphis craccivora).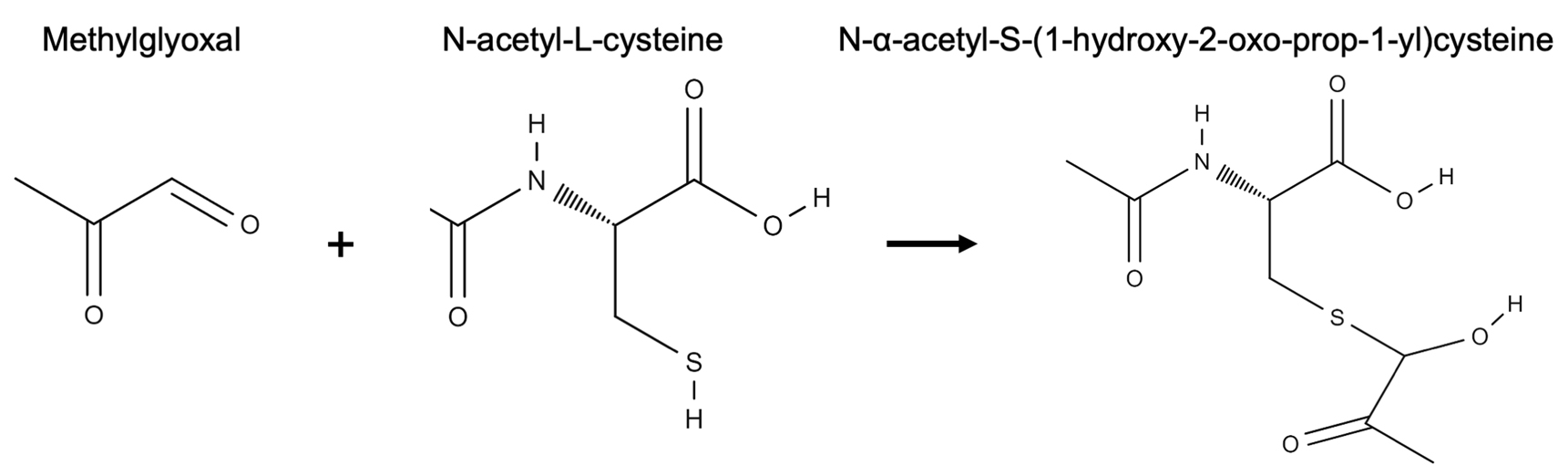
Figure 1. Reaction of methylglyoxal with N-acetyl-L-cysteine to form N-α-acetyl-S-(1-hydroxy-2-oxo-prop-1-yl)cysteine. Structures were drawn using the online resource at http://molview.org.
Materials and Reagents
- Disposable nitrile gloves (Fisher Scientific, catalog number: 19-130-1597 )
- Lab coat
- N95 mask (3M, catalog number: 19-033-524 )
- 32 oz (946 ml) plastifoam cups (First Street, model: SF32 )
- White printing paper letter size (8 1/2" x 11") (Office Depot, catalog number: 841195 )
- Bug domes (BioQuip)
- Fine tip paint brush (academy 775 round size 0; Grumbacher, catalog number: 14173361639 )
- Disposable Petri dish (Fisher Scientific, catalog number: FB0875713 )
- Pipette tips 2 µl, 10 µl, 200 µl, 1,000 µl
- Eppendorf tubes (1.5 ml)
- Eppendorf tubes (2 ml)
- Pellet pestles (Fisher Scientific, catalog number: 12-141-364 )
- Methacrylate disposable cuvettes (Fisher Scientific, catalog number: 14-955-127 )
- UC Mix 3 Soil (Plaster Sand 15.50 cu. ft., peat moss 11.50 cu. ft., KNO3 0.25 lb., limestone flour 1.50 lb., phosphate 1.25 lb., dolomite 3.75 lb., magnesium 0.07 lb., iron 0.13 lb., manganese 0.03 lb., zinc 0.05 lb., copper 0.11 lb.) (https://agops.ucr.edu/soil-mixing) or similar plaster sand-peat moss mix soil
- Cowpea seeds (cv CB46)
- Cowpea aphid colony
- Sodium phosphate monobasic (NaH2PO4) (Fisher Scientific, catalog number: S369-500 )
- Methylglyoxal (Sigma-Aldrich, catalog number: M0252-25ML )
- N-acetyl-L-cysteine (Alfa Aesar, catalog number: A15409 )
- 70% Perchloric acid (Macron Fine Chemicals, catalog number: MK-2766-500 )
- Potassium carbonate (CK2O3) (Fisher Scientific, catalog number: P208-500 )
- Sodium hydroxide (NaOH) (Fisher Scientific, catalog number: BP359-212 )
- Charcoal (Fisher Scientific, catalog number: C270C )
- Alkacid wide-range test ribbons (Fisher Scientific, catalog number: A979 )
- ddH2O
- Ice
- 100 mM Methylglyoxal (see Recipes)
- 100 mM NaH2PO4 buffer (pH 7.0) (see Recipes)
- 500 mM N-acetyl-L-cysteine (see Recipes)
- 5% Perchloric acid (see Recipes)
- 1 M CK2O3 (see Recipes)
- 10 mM NaOH (see Recipes)
Equipment
- Pipettes (P2, P20, P200, P1000)
- Microcentrifuge (Beckman Coulter, model: Microfuge 22R Centrifuge )
- Spectrophotometer (Beckman Coulter, model: Du 730 Life Science UV/Vis Spectrophotometer)
- pH meter (Mettler Toledo, model: MP 220 pH Meter )
- Electronic balance (Mettler Toledo, model: AG104 Electric Balance)
- Plant and aphid growth room or greenhouse maintained at 28 ± 2 °C and 16 h light/8 h dark photoperiod
- Fume hood
Procedure
- Planting and plant growth
- Fill up the plastifoam cups with UC Mix 3 soil. Use a minimum of three plants for each infestation time point and for non-infested control (biological replicates).
- Plant a single cowpea seed in each cup at around 2.5 cm depth.
- Maintain the plants at 28 ± 2 °C for 16 h light/8 h dark photoperiod and water as needed.
- After 2 weeks, plants will have full grown unifoliate leaves and will be ready for infestation.
- Aphid infestation
- Using a wet fine paintbrush, collect apterous (wingless) 4th stage nymphs and adult cowpea aphids from a colony and place the aphids in a Petri dish as seen in Figure 2.
Note: A cowpea aphid colony is best maintained on young plants with 16 h of light at 28 ± 2 °C.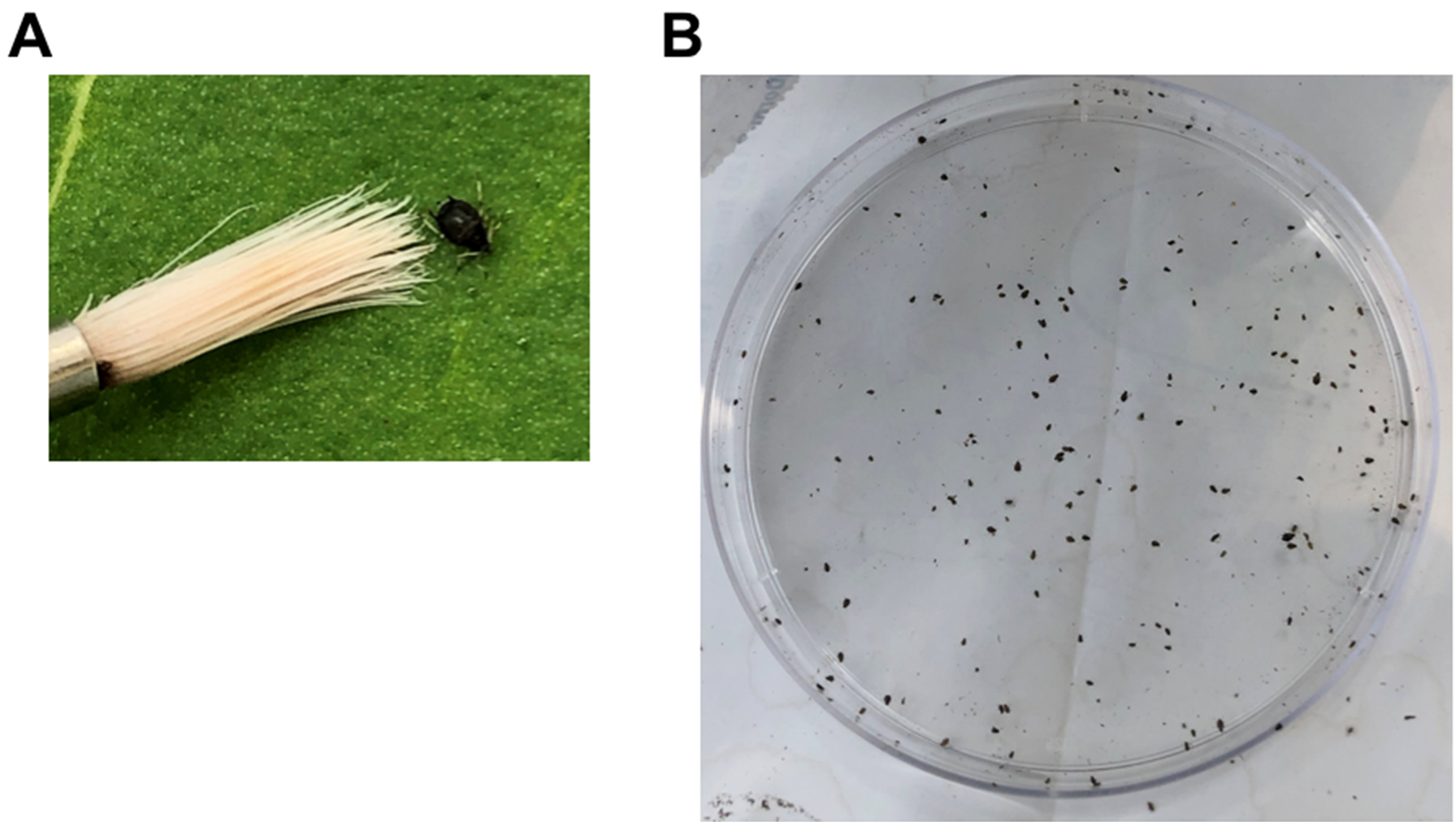
Figure 2. Collection of cowpea aphids. A. An adult cowpea aphid next to a paintbrush. B. Cowpea aphids in a 9-mm Petri dish. - Move the dish with the aphids to the plant growth room where the plants will be infested and maintained.
- Using the same paintbrush, transfer about 100 aphids to a single unifoliate leaf.
Notes:- Ensure that the leaf is evenly infest with aphids as seen in Figure 3C.
- For multiple time point infestations, infest the plants such that you are able to harvest the leaves, from all time points, at the same time. For example, for 24 h and 48 h time points, infest on day 1 and day 2 at a similar time of the day and harvest the leaves on day 3.
- Make a slit in a piece of paper of about 15-mm height and 7-mm width (Figure 3A), and place the paper around the cowpea leaf to form a barrier, to inhibit aphids from leaving the leaf, as seen in Figure 3. Place a paper barrier on the control non-infested plants as well.
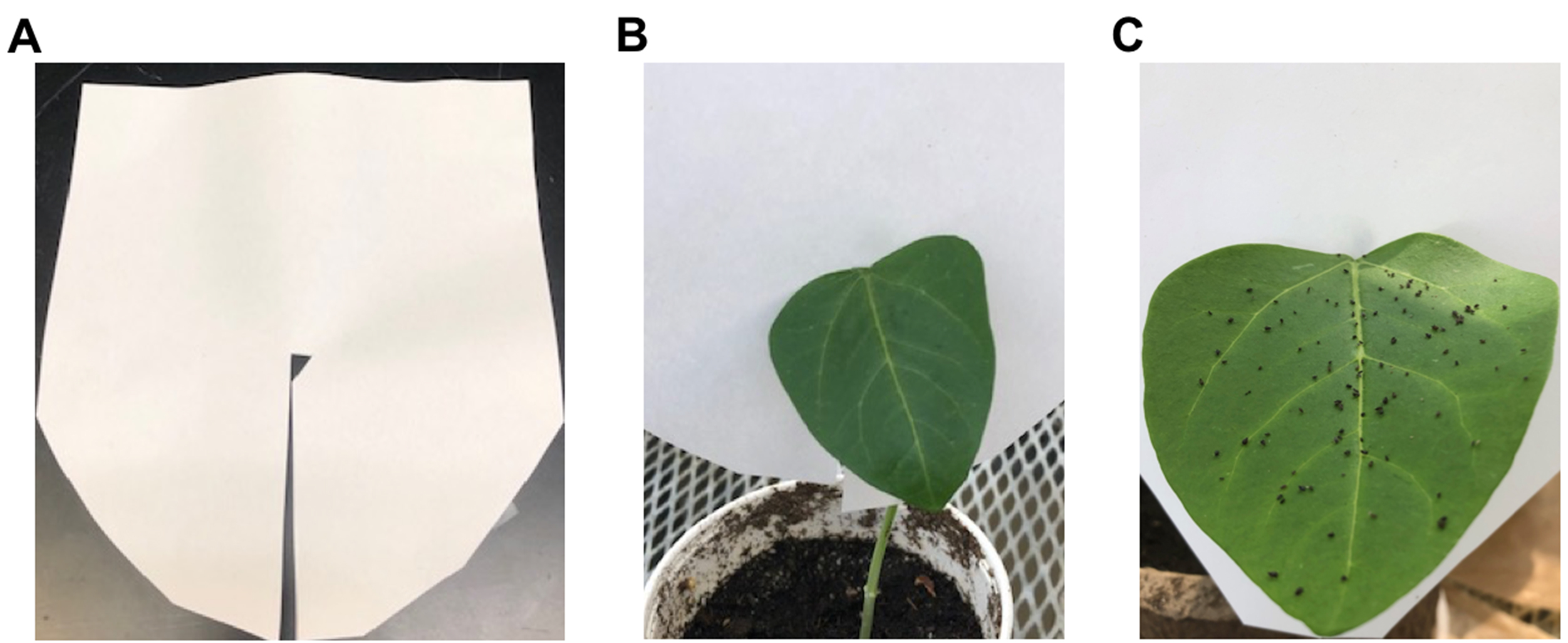

Figure 3. The paper barrier used to keep aphids on a single leaf. A. The slit in the paper allows to insert the paper around the petiole of a leaf. B. A cowpea plant with the paper barrier in place. C. A cowpea leaf evenly infested with cowpea aphids.
Note: The paper barrier discourages aphids from leaving the leaf, while limiting mechanical damage that could be caused by traditional clip cages, used to confine aphids on a given location on a leaf. - Move the infested plants into a bug dome and maintain at 28 ± 2 °C for 16 h light/8 h dark photoperiod.
- Place the non-infested control plants in a different bug dome, adjacent to the bug dome containing the aphid-infested plants.
- Using a wet fine paintbrush, collect apterous (wingless) 4th stage nymphs and adult cowpea aphids from a colony and place the aphids in a Petri dish as seen in Figure 2.
- Methylglyoxal extraction
Notes: Because perchloric acid and methylglyoxal are toxic, before starting the procedure, take the following safety measures:- Wear personal protective equipment (PPE), including a lab coat, gloves and N95 mask, for handling perchloric acid and methylglyoxal.
- Perform all steps in a fume hood.
- Have an easily accessible designated waste container, inside the hood, to dispose everything that comes in contact with perchloric acid.
- Move the plants and a balance, for weighing the leaves, near the fume hood.
- Mark and weigh appropriate number of empty 1.5 ml Eppendorf tubes and record their weights.
- Remove the paper barrier, cut the petiole of the infested leaf and immediately cut the leaf in half and place a half leaf in a 1.5 ml Eppendorf tube.
- Weigh the tube and determine the leaf tissue weight by subtracting the weight of the empty tube.
Note: The weight of half a cowpea 2-week-old unifoliate leaf is around 300 mg. - Add 300 µl of 5% perchloric acid and homogenize the leaf tissue using a pestle.
Notes:- Use 1:1 ratio of mg leaf tissue and µl volume of 5% perchloric acid.
- To avoid variability, steps 3-5 should be performed quickly.
- Incubate the samples for 15 min at room temperature.
- Centrifuge the samples at 13,000 x g for 10 min at 4 °C.
- Transfer the supernatant to a new 1.5 ml tube.
- Divide equally the supernatant from each sample into two 1.5 ml tubes to use as technical replicates.
- Add 10 mg of charcoal to each tube to decolorize the supernatant and incubate for 15 min at room temperature.
Note: Invert the tubes a couple of times during the 15 min incubation. - Neutralize the solution by adding an appropriate volume of 1 M CK2O3.
Note: Use caution when titrating with 1 M CK2O3 and monitor the pH of the sample with Alkacid wide-range test ribbons to prevent over-neutralization. For 250 µl of supernatant, around 45 µl of 1 M CK2O3 is needed. If additional titration is needed, use small volumes of 1 M CK2O3. - Centrifuge the samples at 13,000 x g for 10 min at 4 °C.
Note: If the pellet is floating and is not stuck at the bottom of the tube, repeat the centrifugation. - Transfer the supernatant from each tube to a new 1.5 ml tube and keep on ice until ready to measure methylglyoxal concentration.
- Methylglyoxal standard curve
- Prepare a standard curve by adding the reagents in 2 ml tubes as described in Table 1.
Table 1. Reagents and concentrations used for developing the methylglyoxal standard curve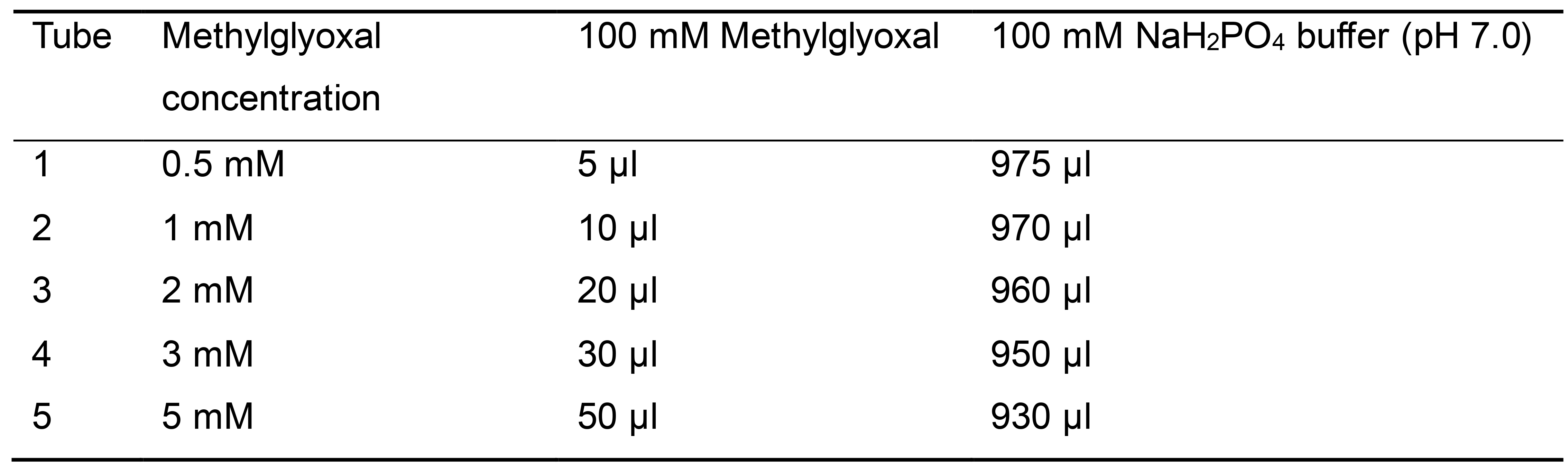
Notes:- Methylglyoxal is highly toxic and should be handled in a fume hood with proper PPE. All steps including the formation and reading of N-α-acetyl-S-(1-hydroxy-2-oxo-prop-1-yl)cysteine with the spectrophotometer should also be performed in the fume hood for maximum safety, with an easily accessible designated waste container for everything that comes in contact with methylglyoxal.
- 100 mM methylglyoxal is made fresh in ddH2O.
- Add 20 µl of 500 mM N-acetyl-L-cysteine to each tube to start the reaction shown in Figure 1.
- Invert the tubes to mix the reaction and incubate at room temperature for 10 min.
Notes:- Tubes should be inverted multiple times during the incubation period.
- If this protocol is used for measuring methylglyoxal under different stresses or different plant species, then, observe the kinetics of the reaction of a sample to ensure that the 10 min is a long enough incubation time for the reaction to reach a plateau.
- Move the samples into cuvettes and using a spectrophotometer detect the formation of N-α-acetyl-S-(1-hydroxy-2-oxo-prop-1-yl)cysteine by measuring the absorbance at 288 nm.
- Plot a standard curve as shown in Figure 4.
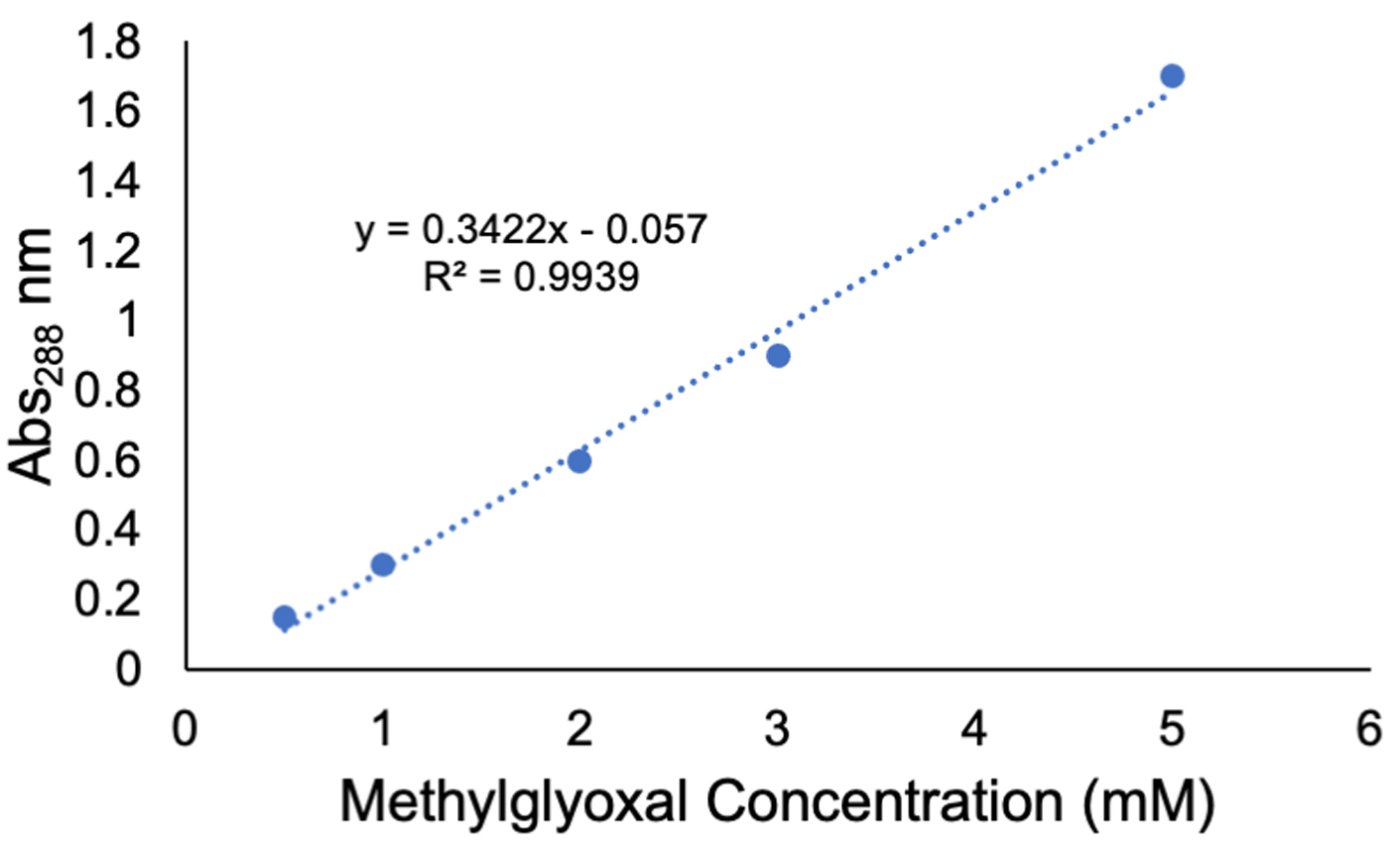
Figure 4. A standard curve generated by measuring absorbance at 288 nm using a spectrophotometer and the methylglyoxal concentrations and reagents in Table 1
- Prepare a standard curve by adding the reagents in 2 ml tubes as described in Table 1.
- Methylglyoxal estimation
- For each sample, use a 2 ml tube and label accordingly. In each tube, add 930 µl of 100 mM NaH2PO4 buffer (pH 7.0).
Note: 100 mM NaH2PO4 buffer (pH 7.0) should be prepared fresh. - Add 50 µl of each sample to the 2 ml tube with the 100 mM NaH2PO4 buffer.
- Add 20 µl of 500 mM N-acetyl-L-cysteine to each tube to start the reaction shown in Figure 1.
Note: 500 mM N-acetyl-L-cysteine is prepared fresh in 100 mM NaH2PO4 buffer (pH 7.0). - Invert the tubes to mix and incubate at room temperature for 10 min.
Note: Invert the tubes multiple times during the incubation period. - Move the samples into cuvettes and detect the formation of N-α-acetyl-S-(1-hydroxy-2-oxo-prop-1-yl)cysteine by measuring absorbance at 288 nm using a spectrophotometer.
Note: If the sample absorbance falls outside of the standard curve, dilute the sample and redo the reaction starting at Step E1.
- For each sample, use a 2 ml tube and label accordingly. In each tube, add 930 µl of 100 mM NaH2PO4 buffer (pH 7.0).
Data analysis
- Using the methylglyoxal standard curve shown in Figure 4, determine the methylglyoxal concentration (mM) in the samples by (Abs288 – intercept)/slope.
- Multiply the value obtained by 20 [20 being the dilution factor (1,000 µl total volume/50 µl of the sample) for the example given above].
Note: If the samples were diluted before measurement, include that dilution factor in the calculation. - Determine the total methylglyoxal content of a leaf sample (µmol g-1 fresh weight) by multiplying the methylglyoxal concentration (mM) by the extracted sample volume (ml) and dividing by the leaf fresh weight (g).
- Average the two technical replicates for the value of a biological replicate. Then, determine the value of each time point by calculating the average of the different biological replicates.
Recipes
- 100 mM Methylglyoxal (prepared fresh)
Add 15.4 µl of methylglyoxal to 984.6 µl ddH2O - 100 mM NaH2PO4 buffer (pH 7.0) (prepared fresh)
Dissolve 0.599 g NaH2PO4 in 50 ml ddH2O. Bring to pH 7 with 10 mM NaOH - 500 mM N-acetyl-L-cysteine (prepared fresh)
Dissolve 81.5 mg N-acetyl-L-cysteine in 1 ml NaH2PO4 buffer (pH 7.0) - 5% Perchloric Acid
Add 1.79 ml 70% Perchloric acid to 23.21 ml ddH2O - 1 M CK2O3
Dissolve 6.91 g CK2O3 in 50 ml ddH2O - 10 mM NaOH
Dissolve 0.4 g of NaOH pellets in 1,000 ml ddH2O
Acknowledgments
This protocol was adapted from Wild et al. (2012) and Borysiuk et al. (2018). This work in the I.K. group is funded by the USDA National Institute of Food and Agriculture Hatch project 1017522. Research in the B.S. group is supported by the grant 2014/14/E/NZ3/00155 from the Polish National Science Centre.
Competing interests
Authors have no conflict of interests or competing interests.
References
- References1.Borysiuk, K., Ostaszewska-Bugajska, M., Vaultier, M. N., Hasenfratz-Sauder, M. P. and Szal, B. (2018). Enhanced formation of methylglyoxal-derived advanced glycation end products in Arabidopsis under ammonium nutrition. Front Plant Sci 9: 667.
- Gilbert, R. P. and Brandt, R. B. (1975). Spectrophotometric determination of methylglyoxal with 2,4-dinitrophenylhydrazine. Anal Chem 47: 2418-2422.
- Hoque, T. S., Hossain, M. A., Mostofa, M. G., Burritt, D. J., Fujita, M. and Tran, L. S. (2016). Methylglyoxal: an emerging signaling molecule in plant abiotic stress responses and tolerance. Front Plant Sci 7: 1341.
- Hossain, M. A., Hossain, Z. and Masayuki, F. (2009). Stress-induced changes of methylglyoxal level and glyoxalase I activity in pumpkin seedlings and cDNA cloning of glyoxalase I gene. J Crop Sci 3(2): 53-64.
- Kaur, C., Kushwaha, H. R., Mustafiz, A., Pareek, A., Sopory, S. K. and Singla-Pareek, S. L. (2015). Analysis of global gene expression profile of rice in response to methylglyoxal indicates its possible role as a stress signal molecule. Front Plant Sci 6: 682.
- Li, Z. G., Duan, X. Q., Min, X. and Zhou, Z. H. (2017). Methylglyoxal as a novel signal molecule induces the salt tolerance of wheat by regulating the glyoxalase system, the antioxidant system, and osmolytes. Protoplasma 254(5): 1995-2006.
- MacWilliams, J. R., Dingwall, S., Chesnais, Q., Sugio, A. and Kaloshian, I. (2020). AcDCXR Is a cowpea aphid effector with putative roles in altering host immunity and physiology. Front Plant Sci 11(605).
- Melvin, P., Bankapalli, K., D'Silva, P. and Shivaprasad, P. V. (2017). Methylglyoxal detoxification by a DJ-1 family protein provides dual abiotic and biotic stress tolerance in transgenic plants. Plant Mol Biol 94(4-5): 381-397.
- Mostofa, M. G., Ghosh, A., Li, Z. G., Siddiqui, M. N., Fujita, M. and Tran, L. P. (2018). Methylglyoxal - a signaling molecule in plant abiotic stress responses. Free Radic Biol Med 122: 96-109.
- Racker, E. (1951). The mechanism of action of glyoxalase. J Biol Chem 190: 685-696.
- Wild, R., Ooi, L., Srikanth, V. and Munch, G. (2012). A quick, convenient and economical method for the reliable determination of methylglyoxal in millimolar concentrations: the N-acetyl-L-cysteine assay. Anal Bioanal Chem 403(9): 2577-2581.
- Yadav, S. K., Singla-Pareek, S. L., Ray, M., Reddy, M. K. and Sopory, S. K. (2005). Methylglyoxal levels in plants under salinity stress are dependent on glyoxalase I and glutathione. Biochem Biophys Res Commun 337(1): 61-67.
Article Information
Copyright
© 2020 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
MacWilliams, J. R., Ostaszewska-Bugajska, M., Borysiuk, K., Szal, B. and Kaloshian, I. (2020). Quantification of Methylglyoxal Levels in Cowpea Leaves in Response to Cowpea Aphid Infestation. Bio-protocol 10(20): e3795. DOI: 10.21769/BioProtoc.3795.
Category
Plant Science > Plant metabolism > Other compound
Plant Science > Plant immunity > Plant-insect interaction
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










