- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Preparing Viable Hippocampal Slices from Adult Mice for the Study of Sharp Wave-ripples
Published: Vol 10, Iss 19, Oct 5, 2020 DOI: 10.21769/BioProtoc.3771 Views: 5708
Reviewed by: Akira KarasawaCarey Y. L. HuhAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Cell-attached and Whole-cell Patch-clamp Recordings of Dopamine Neurons in the Substantia Nigra Pars Compacta of Mouse Brain Slices
Stefano Cattaneo [...] Stefano Taverna
Aug 5, 2021 5814 Views
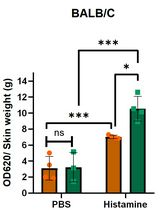
Analysis of Vascular Permeability by a Modified Miles Assay
Hilda Vargas-Robles [...] Michael Schnoor
Apr 5, 2025 2590 Views
Ultrafast Isolation of Synaptic Terminals From Rat Brain for Cryo-Electron Tomography Analysis
Rong Sun and Qiangjun Zhou
Sep 5, 2025 3582 Views
Abstract
We describe a protocol for preparing acute brain slices which can produce robust hippocampal sharp wave-ripples (SWRs) in vitro. The protocol is optimized for its simplicity and reliability for the preparation of solutions, slicing, and recovery incubation. Most slices in almost every mouse prepared though the protocol expressed vigorous spontaneous SWRs for ~24 h, compared to the 20-30% viability from "standard" low sodium slicing protocols. SWRs are spontaneous neuronal activity in the hippocampus and are essential for consolidation of episodic memory. Brain slices reliably expressing SWRs are useful for studying memory impairment and brain degeneration diseases in ex vivo experiments. Spontaneous expression of SWRs is sensitive to conditions of slicing and perfusion/oxygenation during recording. The amplitude and abundance of SWRs are often used as a biomarker for viable slices. Key improvements include fast circulation, a long recovery period (3-6 h) after slicing, and allowing tissue to recover at 32 °C in a well perfused incubation chamber. Slices in our custom-made apparatus can express spontaneous SWRs for many hours, suggesting a long period with balanced excitation and inhibition in the local networks. Slices from older mice (~postnatal 180 days) show similar viability to younger (postnatal 21-30) mice.
Keywords: Brain sliceBackground
Acute brain slices have been an invaluable preparation for electrophysiology and other neuroscience studies. While there is a vast body of literature describing the usage of brain slices, protocols for reliably producing viable slices are usually discussed in less detail. Slices from older animals are typically less viable. Viability of the brain slices is often described using healthy cell morphology and electrophysiological properties of the neurons. However, healthy looking neurons in the tissue do not always translate to normal network behaviors. For example, thin slices prepared with the standard sucrose replacement protocol (e.g., Aghajanian and Rasmussen, 1989; Ye et al., 2006) provide morphologically healthy cells for patch studies, but do not express spontaneous network activity such as hippocampal sharp-wave ripples (SWRs).
SWRs are spontaneous neuronal population events that occur in the hippocampus during sleep and quiet restfulness (for review see Buzsáki, 2015). An experience, such as an exploration of the environment, is registered in the hippocampus as sequential activations of neuronal assemblies (a.k.a “place cell assemblies”). These sequences are then re-activated in SWRs during sleep or quiet restfulness (off-line states), and contribute to the consolidation of the memory (Wilson and McNaughton, 1994; Lee and Wilson, 2002; Ji and Wilson, 2007). Suppression of SWRs impairs hippocampus dependent episodic memory (Ego-Stengel and Wilson, 2010), suggesting that SWRs are important for the reactivation of experience related to neuronal ensembles in the absence of related sensory input.
Spontaneous SWRs also occur in vitro in hippocampal slices (Kubota et al., 2003; Maier et al., 2003 and 2009; Colgin et al., 2004; Behrens et al., 2005; Miyawaki et al., 2014; Keller et al., 2015). However, in standard submerged-type recording chambers, SWRs are rarely seen, due to insufficient perfusion. Vigorous perfusion on both sides of the slice seems to be an essential condition for providing adequate oxygen that allows for the SWRs to occur (Wu et al., 2005; Hájos et al., 2009; Maier et al., 2009).
In our experience, slices prepared with standard high-sucrose low-sodium protocol (Aghajanian and Rasmussen, 1989) only have a lower probability (20-50%) of expressing spontaneous SWRs. Slices from mice older than postnatal (P) 30-days have a further reduced probability of expressing SWRs and often fail to induce LTP. Extending the recovery time after slicing often helped. Consistent with (Maier et al., 2009), we also observed that SWRs were rarely expressed in conventional slice incubation chambers, which have passive fluid convection or oxygen bubble induced convection (e.g., Brain Slice Keeper-4 apparatus, Automate Scientific). A forced fluid circulation with a small pump significantly increased SWR expression. We have tested 198 slices from 35 mice of P 35 to P180 days of age, and every animal tested showed spontaneous SWRs, including 6-month-old 5xFAD mice with obvious amyloid plaques and slices from mice that have undergone severe traumatic brain injuries.
Fast circulation may be important for two reasons: 1. Thick slices (350-500 μm) are essential for preserving SWR generating circuits. Fast circulation on both sides of the slice can improve the oxygen delivery into the thickness of the tissue. 2. SWRs are spontaneous activity occurring 1-2 times/second. Continuous activation of large number of neurons requires more oxygen provided by fast circulation.
Our protocol is also aimed at its simplicity, i.e., using stock solutions to minimize the solution preparation time (< 30 min), using simple low sodium high sucrose cutting solution without unstable compounds like NMDG (Tanaka et al., 2008) and antioxidants (Brahma et al., 2000), and omitting the complex procedure of intra-cardiac perfusion and protective recovery protocol (Ting et al., 2014 and 2018). Some of our research papers using the protocol have been published (Jiang et al., 2018; Sun et al., 2018; Li et al., 2019).
Materials and Reagents
- Paper towel
- Glass pipet for local field potential electrode, 1.5 mm outer diameter x 1.1 mm inner diameter, 7.5 cm long borosilicate glass pipet with inner filament (Sutter Instruments, catalog number: BF150-110-7.5 )
- Nylon mesh (Component supply UMN-600), glued onto the bottom of the slice holder
- Silicon tubing (1/4-3/8, VWR , catalog number: 89403-862), for connecting the pump and the top chamber
- Disposable Transfer Pipettes (Thermo Scientific, Samco 3.2 ml) for slice transferring between incubation and recording chamber
- Animals
Wildtype Mice (C57BL/6J, RRID: IMSR_JAX :000664, Jackson Labs) and transgenic 5xFAD mice (RRID:MMRRC_034840-JAX, Jackson Labs) (Oakley et al., 2006) were used. Mice for experiments were from our Georgetown colony, they were kept on a standard 12 h light/dark cycle, food and water were provided ad libitum, with all experimental procedures performed in accordance with the guidelines of the Georgetown University Animal Care and Use Committee. - Super glue (liquid form, Loctite, catalog number: 1775049) for gluing the brain during slicing
- Sucrose (ACS Reagents, Fisher Scientific, catalog number: S5-500 )
- KCl (ACS Reagents, Fisher Scientific, CAS: 7447-40-7 )
- MgSO4 (ACS Reagents, Fisher Scientific, CAS: 7487-88-9 )
- CaCl2 (ACS Reagents, Fisher Scientific, catalog number: C79-500 )
- Glucose (ACS Reagents, Sigma-Aldrich, catalog number: G8270-5KG )
- NaH2PO4 (ACS Reagents, Sigma-Aldrich, catalog number: S0751-1KG )
- HEPES (ACS Reagents, Fisher Scientific, catalog number: BP310-1 )
- NaCl (ACS Reagents, Sigma-Aldrich, catalog number: S5886-10KG )
- Carbogen (95% O2, 5% CO2, tank and regulator available from local gas suppliers)
- NaHCO3 powder (ACS Reagents, Fisher Scientific, catalog number: S233-500 )
- Slice cutting ACSF solution (see Recipes)
Stock 1: Sucrose
Stock 2: Cutting ACSF
Stock 3: NaHCO3 powder - Incubation/recording ACSF (see Recipes)
Equipment
For slice preparation
- Vibratome slice cutter (Leica, model: VT1000S )
- Water bath (e.g., Southwest Science , model: SHW10LD 10 Liter Bath)
- Digital Thermometer ( VWR Traceable Double Thermometer, catalog number: 23226-656 , or any type with 0.1 °C accuracy)
- Custom-made incubation chamber illustrated in Figure 2, made of 1/4 Plexiglass (acrylic) sheet, obtained from McMaster-Carr
- Slice holder is cut from an egg crate light diffusor panel with 1/2 x 1/2 x 1/2 cells (e-Plastics SKU: W/EGG.500X2X4)
- 600 ml glass beaker ( VWR , catalog number: 89001-078 )
- Carbogen blower (e.g., Air Stone Cylinder Bubble Diffuser from Amazon) three of them are needed (two for incubation gas blowers for incubation chamber and recording ASCF flask, one used as a filter for ACSF circulation in recording chamber)
- Low voltage, low flow rate pump, available from a solar powered mini water fountain/bird bath kits (Amazon). The pump provides a flow of ~1,000 ml/min, and uses a safe low voltage power supply (below)
- Variable DC power supply for the pump (Circuit specialist CSI5003XE, or B&K Precision 1550 DC Power Supply)
- Surgical scissors for mouse decapitation (WPI, catalog number: 501743G )
- Spring scissors (WPI, catalog number: 15905G ) for cutting through skull and extracting the brain
- Size 4 watercolor pointed brush (Office depot 596713) for transferring brain slices
- Dumont #7 student forceps (Fine Science Tools, catalog number: 97797-00 ) for extracting the brain
- Tapered Spatula (PTFE coated, VWR , catalog number: 10806-412) for extracting the brain
- Dumont #5 student forceps (Fine Science Tools, catalog number: 91150-20 ) for adjusting slice position in recording chamber
For SWR recording
While most electrophysiology rigs can be used for recording SWRs, perfusion of both sides of the slice with a high circulation rate is essential. See Maier et al. (2009) and Hájos et al. (2009) for the modification of conventional chambers on patch recording rigs. Listed below are parts for a low-cost local field potential recording system. The system is easy to use and reliable for recording many hours of SWRs from one slice.
- Local field potential recording chamber
A chamber that can perfuse both sides of slice is important. See procedures and data recording sections. Figure 3 illustrates the flow directions in the custom-made recording chamber. - LFP amplifier, 1000x amplification with a probed head stage (e.g., Warner Instruments, model: DP 311A )
- Data acquisition system to digitize the data (e.g., NeuronNexue smart box, allowing multiple channels with an electrode array)
- An oscilloscope (e.g., Digikey, 2250-SDS1022-ND ) for positioning electrode
- An audio monitor (A-M Systems 3300) for positioning the electrode
- Manual micromanipulators (SD Instruments SKU MX160 ) and magnetic stands for holding the manipulators (SD Instruments)
- 625 nm LED (Thorlabs, model: M625D3 ) for slice illumination under the chamber
- Pipet puller (Sutter, model: P97 ) for pulling ultra-low impedance pipet for local field potential (LFP) recordings
- LFP recording and Stimulation electrodes (low impedance (~500K) glass pipette pulled by the pipette puller). Low electrode impedance (50-500K) is important for low noise recording.
- Peristaltic pump, for ACSF circulation (e.g., Pulsatron Dolphin Series Peristaltic Metering Pump, 60.0 gpd/25 PSI, Norprene tubing, .25" ID x .44" OD).
- Plastic tubing for connecting pump and recording chamber perfusion ( VWR , catalog number: 89403-850 , 3/32-5/32 Tygon tubing)
- Tubing clamp flow regulator ( VWR Talon® Regular Hosecock Clamp), for regulating perfusion rate of the recording chamber
- 1000 ml glass flask for circulating ACSF perfusion of the recording chamber ( VWR , catalog number: 75804-652 )
- Stereo zoom dissecting microscope, 10-40x zoom, available from Olympus, Nikon, or American microscopes (SKU: V-SM-3 SD Instruments)
- Recording bench (Newport, catalog number: IG-22-2 )
This 2 x 2 ft optical breadboard can be set up on standard laboratory bench top. Recording the LFP from SWRs does not need a vibration isolation table. A Faraday cage may also be avoided if a low impedance glass electrode is used and there is no instrument power cord within 30-inches of the set up.
Procedure
- Preparation
Before slicing, prepare two artificial cerebral-spinal fluid (ACSF) solutions from the stock solutions (procedures in the recipe section below): The cutting ACSF (200 ml) and incubation ACSF (2,000 ml). Both are vigorously bubbled for ~3 min with carbogen gas. The cutting ACSF is then cooled down to 4 °C in an ice/water bath, and the incubation ACSF is warmed up to 32 °C in the incubation apparatus (see details in the Recipes section). - Slicing
- The procedures for using animals are strictly following a protocol approved by the Institutional Animal Care and Use Committee (IACUC) at Georgetown University Medical Center following the NIH guidelines.
- The animal is deeply anesthetized in an anesthesia jar (approved by IACUC) by freely breathing isoflurane vapor (> 5%) for ~20 s. After being fully anesthetized (loss of muscle tension and no response to tail pinch), the animal is quickly decapitated with a pair of surgical scissors.
- Use a pair of small surgical scissors to cut the skin along the midline of the scalp to expose the skull, then immerse the head in the cold (4 °C) cutting ACSF solution for ~1 min. While immersed, remove the tissue covering the skull to reduce heat insulation. The brain should be cooled down to 5-10 °C after this step.
- Use a pair of small surgical scissors to cut the skull following Figure 1. Use the Dumont #7 forceps to remove the skull fragments over the brain. Gently extract the brain with the spatula, starting from the rostral end, and let the brain slide into the cutting ACSF solution at ~4 °C.
Important: Make sure that the tip of spatula stays at the midline of the ventral side of the brain, to avoid touching the ventral temporal lobes which may lead to compression of the ventral hippocampus. The temperature of the cutting ACSF needs to be higher than 0 °C to avoid damage caused by hypothermia.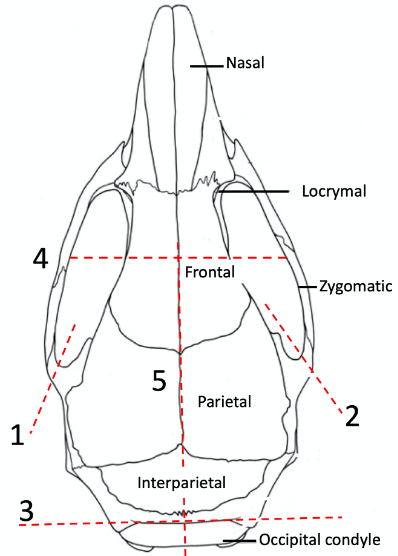
Figure 1. Skull cutting lines for brain extraction. Red lines mark the cuts through the skull. Numbers are the suggested sequence. Cut 5 should be not too deep to ensure no disturbances to the cortical tissue beneath the cut. The head needs to be cooled down to below 10 °C before brain extraction. As long as the brain is cold, it is not critically important for a short extraction time. Attention should be paid not to stretch or compress the hippocampi during extraction. - Apply a thin layer of super glue liquid on the surface of the vibratome cutting stage for gluing the brain on the stage.
- Use the spatula to pick up the brain from the cutting solution (dorsal side of the brain up), and use a small piece of paper towel to absorb excess solution from the ventral side, and then place the ventral side of the brain onto the surface of the cutting plate coated with the super glue liquid.
- Apply a small amount of pressure onto the dorsal side of the brain to secure a firm gluing contact between the brain and the cutting stage.
- Now mount the cutting stage onto the vibratome, and fill the chamber of the cutting stage with 4 °C cutting ACSF. The brain is sliced in a horizontal orientation, from the rostral end to the caudal end. A 12.7-degree tilting angle from the horizontal plain (high at the rostral end) may better preserve the connection between CA3 and CA1 areas for better SWR expression (Miyawaki et al., 2014). However, when cut at 480-500 μm, the 12.7 degree of tilting angle is not important. Thicker slices (> 350 μm, Maier et al., 2009) are very important for the expression of SWRs.
- Hippocampal slices (that we cut at 488 μm) are cut at a high vibrating speed (~70 Hz) and a low preceding speed (1-2 on the vibratome setting). It is OK to allow the temperature of the cutting ACSF solution to elevate from ~4 °C to 10 °C while cutting. Bubbling with carbogen is needed if the total cutting time is longer than 30 min, to prevent a change in the pH of the cutting solution.
- After each cut, the slice is gently transferred to the incubation ACSF in the incubation chamber using the fine paint brush while paying attention not to bend or compress the tissue. The whole horizontal slice (two hemispheres) can be trimmed into two halves before transferring. Usually we obtain 4-5 whole horizontal slices from each animal (10 hippocampal slices from both sides).
- Incubation
- Incubation is maintained at 32 °C in the incubation apparatus (Figure 2), with constant flow circulation. Adjust the variable power supply and maintain a rate of ~1,000 ml/min (3.5-4 V to the pump). The flow rate should not be higher than 1,000 ml/min, in order to reduce movement or floating of the slices in the solution which may cause mechanical damage to the tissue.
- Carbogen should be constantly bubbled into the ACSF in the beaker at a rate of 200-400 ml/min, to maintain saturation of O2/CO2 in the ACSF solution.
- After 2 h at 32 °C, reduce the water bath temperature to 27 °C and allow the ACSF to slowly cool down from 32 °C to 27 °C (it takes about 1.5 h). Continue incubating until the slices are used. Slices stay viable for ~24 h in the incubation apparatus.
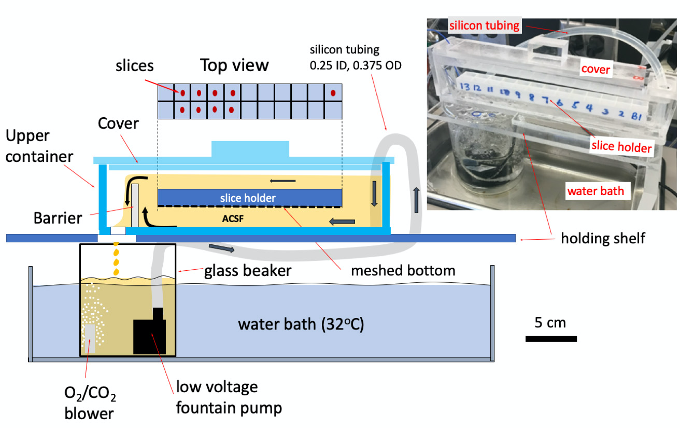
Figure 2. Construction of the incubation apparatus. The upper container of the apparatus is a container of flowing ACSF, made of acrylic plastic glass. Inside the container is a slice holding chamber (cut from an egg crate style lamp diffuser [see Equipment]). Slices are held in separated compartments (top view), and immersed in circulating ACSF solution. Nylon mesh (see Materials) is glued to the slice holder to allow good circulation. The bottom side of the slice holder has a higher flow rate than that of the top side, that provides a small downward current in each compartment to stabilize the slices against the mesh. A barrier in the top container maintains the fluid level. Below the apparatus is a 600 ml glass beaker, sitting in a water bath with water ~6 cm deep to maintain the temperature of 32 °C. ACSF in the beaker is continuously bubbled with carbogen at a rate of 200-400 ml/min. A low voltage pump (see Equipment) is used for circulating the ACSF. The pump is powered by a variable DC power supply (see Equipment). Adjust the voltage to the pump (3-4 V) to have a flow of rate of ~1,000 ml/min (about 1/2 of the pump capacity). The total fluid volume is 800-1,000 ml. The upper container is covered to reduce evaporation and a change in osmolarity after many hours.
- SWR recording
- Establish a circulation of the incubation/recording ACSF solution in the recording chamber: Put 800-1,000 ml of incubation ACSF into the 1,000 ml flask, put the flask on a shelf 1-1.5 meters above the level of the recording chamber to allow a gravitational flow and syphon it from the flask to the chamber via the 3/32x 5/32 Tygon tubing. An air stone is connected to the tubing end in the flask and serves as a fluid filter for the circulation and as an anchor for the tubing to stay at the bottom of the flask. Suction by laboratory vacuum or a syringe is used to prime the syphon. The flow rate is regulated by the tubing clamp. The normal circulation rate is 40-100 ml/min. The fluid dropped out of the recording chamber is collected in a 100 ml beaker below the chamber and sent back to the flask by a peristaltic pump. We use gravitational force for both inlet and outlet flow of the ACSF as this can completely remove the static electric noise generated by squeezing the tubing while running the peristaltic pump. The ACSF in the flask is continuously bubbled with carbogen at a rate of 50-100 ml/min, to maintain saturation of O2/CO2.
- SWRs gradually emerged after > 3 h of incubation. After 7 h of incubating, most of the slices should express SWRs from the dorsal most to the ventral most of the hippocampus.
- To position a slice in recording chamber: Carefully transfer a slice from the incubation chamber into the center of the recording chamber by a fluid transfer pipette (the small end of the pipette is cut out). The slice should be held stable at the center of the chamber by the vertical flow of the fluid. The position and orientation of the slice can be gently adjusted by a pair of #5 fine forceps. Pay great attention not to bend or compress the tissue. A mechanical disturbance to the tissue can stop or attenuate SWRs for a few minutes. The hippocampus should be positioned at the center of the view field of the stereo microscope at its full zoom magnification (40x).
- To prepare the recording electrode: Ultra-low resistance glass electrode is pulled by 5-7 pulls using the P-97 puller. Multiple pulls result in resistance of 50-100K Ohms when filled with ACSF, and the tip opening is about 10 μm. Low resistance leads to low noise for the LFP recording.
- Mount the electrode onto the head stage of the amplifier. Put the tip of the electrode into the fluid surface in the recording chamber over the top of the hippocampal slice. This step can be done without the microscope. Turn on the oscilloscope and the audio monitor. There should be a flat baseline in the oscilloscope and a small static noise in the audio monitor. A jumping baseline and large audio noise indicate the electrode is not filled well (air bubbles). Measure the resistance of the electrode, it should be less than 100K Ohms.
- To position the electrode tip: Turn the zoom magnification of the microscope down to the minimum (0.75x) and adjust the focus to find the electrode tip. Adjust the tip's position with the manipulator to lower the tip to approach the recording position (stratum radiatum of CA1, Figure 4A). When adjusting the electrode, always adjust the microscope focus so the electrode tip is in sight. Never move the electrode if its tip is not in the focus of the microscope. At this time, do not insert the tip into the tissue. Now increase the zoom magnification of the microscope to 2-3x, adjust the focus to find the electrode tip. At the high zoom power continue to lower down the electrode until it sits on the surface of the tissue (both the slice surface and the electrode tip is in focus). Now the sound of the ripple oscillation (~100 Hz) should be audible in the audio monitor.
- To insert the electrode tip into the tissue: Slowly advance the electrode tip to the surface of the tissue. The ripple sound becomes louder when approaching the tissue. When the tip touches the tissue, a loud noise can be heard in the audio monitor. At the contact position, slowly turn the knob of the insertion axis of the manipulator 1/4-1/2 turns, which advances the tip 90-180 μm into the tissue. Note that the mechanical disturbance of the electrode can attenuate the amplitude of the SWRs for a few minutes, so slow insertion is important (~20 s for 90 μm advance). Such small and slow movement of the electrode tip cannot be seen by the microscope, so guiding by audio monitor is needed. After a few minutes of electrode insertion, vigorous SWRs with high amplitude should appear and be sustained for hours (Figure 4). We usually recorded 100-200 μm below the surface where the signal is the largest.
- Slices from dorsal hippocampus have a much slower SWR rate than those from the ventral hippocampus. SWRs will only be sustained when both sides of the slice are well perfused in a submerged recording chamber. SWRs do not express well in conventional single sided perfusion chambers (e.g., Warner RC-22).
- The sustaining of SWRs during recording: In our two-sided perfusion chamber shown in Figure 3, SWRs can be sustained for ~24 h (Figure 4C, Figure 5).
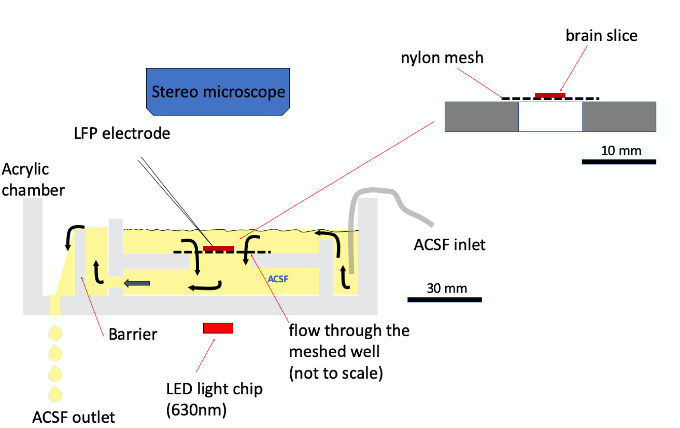
Figure 3. Recording chamber. The schematic drawing illustrates a custom-made recording chamber that creates a downward vertical fluid flow around the slice. The flow provides vigorous perfusion to both sides of the slice. Also, the downward flow keeps the slice stable (no need for a weight on the slice). We use a barrier to maintain the fluid level on top of the tissue. The outlet of the ACSF is gravity dripping, which significantly reduces the electrical noise introduced by vacuum suction. Also, the ACSF can be collected and re-circulated by a peristaltic pump. A light emitting diode (LED) chip on the bottom of the chamber is used to provide diffuse illumination. Using red LED (630 nm) can avoid photoactivation of the neurons expressing channel rhodopsin-2. Red light also increases the transparency of the thick tissue. This chamber is thick and will not fit onto a compound microscope for patching cells. However, SWRs do not express well in conventional single-side perfusion chambers (e.g., Warner RC-22). Ensuring that both sides of the tissue are perfused well is important (Hájos et al., 2009; Maier et al., 2009; Caccavano et al., 2020) for the SWRs to express well in chambers for the cell patch recordings.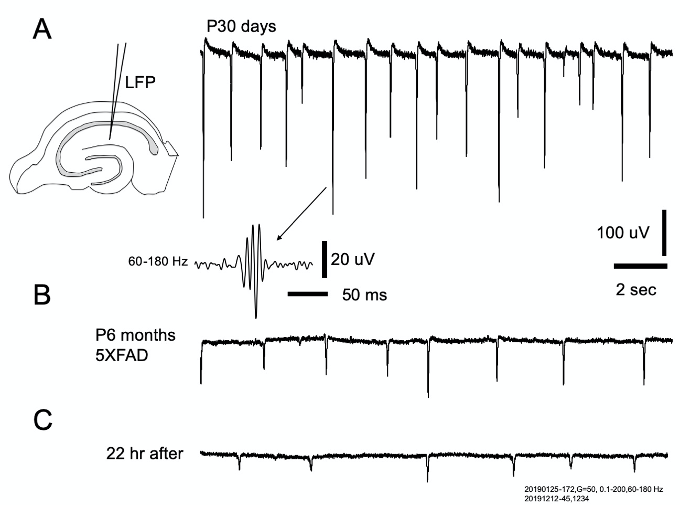
Figure 4. SWRs from young and old mice. A. left: SWRs were recorded from striatum radiatum by an ultra-low resistant glass pipet (< 100K ohms, filled with saline) that provides excellent signal-to-noise ratio. Right traces: Abundant SWRs were seen in young mice with high sharp wave amplitude (top trace), and ripple oscillations (bottom insert, filtered between 60-180 Hz for clarity). B. All older mice tested (P6 month, 4 out of 4) expressed SWRs. The SWRs had lower amplitude and rate compared to the young mice. C. SWRs from a slice of the same animal in B, recorded after overnight perfusion in the holding chamber. While the amplitude and repeating rate were further reduced, the occurrence rate of SWRs didn't show a large decline.
Data analysis
SWRs can be clearly distinguished from seizure/interictal spikes by its occurrence rate, LFP amplitude, and calcium signals (Li et al., 2019). SWRs have a high occurrence rate (1-3 events/s) while interictal spikes are much slower (0.3-0.01 events/s). SWRs are also accompanied by ~100 Hz ripple oscillations (Figure 4A insert), but interictal spikes are mostly accompanied by low gamma (20-50 Hz).
Recovery in the incubation apparatus is important for SWR expression. In our experience with 198 slices, SWRs were never seen from slices that recovered for less than 2 h, but always seen after 7 h of recovery time. Usually, younger mice (P28-P35) recover sooner, and 3-4 h of incubation is adequate. Older mice (P60-P180) take a longer time to recover from slicing, but after 5-7 h, most of the slices show SWRs. In animals genetically encoded with GCaMP-6f, hippocampal regions show strong green fluorescence after cutting, which suggests an elevated concentration of intracellular calcium. The fluorescence gradually reduced after a few hours of recovery in the incubation chamber. In conventional beaker-and-oxygen-blower type incubation chambers, often only a fraction of slices recovered well and expressed SWRs. The fluid circulation in conventional chambers were not well controlled. When oxygen delivery was less optimized, the viability of the tissue declined at the same time as the recovery from the trauma of slicing. Longer recovery time would not improve the viability if the ACSF circulation is compromised.
To improve the fluid circulation during the recovery period, we have fabricated a number of custom-made incubation chambers for longer recovery time. The common feature for these chambers is a forced circulation either by a stir bar or a pump. We found that ordinary peristaltic pumps do not provide adequate flow for a large incubation chamber which holds ~26 slices from 2-3 animals. Thus, we used a low voltage DC pump (obtained from a solar fountain) with a flow of ~1,000 ml/min. This high flow greatly improved the circulation of ACSF around all the slices. Better oxygenation also allows the slices to recover in a higher temperature (~32 °C). The apparatus brings a stable condition for recovery from cutting related cellular injury and elevated intracellular calcium. Part of our work using thick slices and forced circulation have been published (Sun et al., 2018; Jiang et al., 2018; Li et al., 2019). Slices from older (P3-6 month) animals also have apparent normal SWR expression prepared by this protocol (Figure 4B/C) despite traumatic brain injury (unpublished observations) or the development amyloid plaques in the 5XFAD animals (Caccavano et al., 2020).
In most slices, the amplitude of the SWRs gradually reduced with time in the recording chamber. It took ~5 h in the recording chamber for the SWR amplitude to reduce to half (Figure 5). It seems that 6-month old mice had more amplitude reduction than those in one-month old animals. It also appears that after long recording, the SWR amplitude was less diverse; more SWRs with similar amplitude were seen (Figure 5).
It seems that slices with no spontaneous SWRs also showed poor performance in evoked activities, e.g., failed to be potentiated by a tetanus stimulation (100 Hz x 100 pulses) and low amplitude in theta-gamma oscillations when bathed in carbachol (20 mM) containing ACSF. These less viable slices were often subjected to shorter recovery time after cutting (unpublished observations).
The biomarker of slice "viability" is an important issue. In this manuscript, we consider a slice viable if SWRs are vigorously expressed for hours. Expressing SWRs requires network integrity, with balanced synaptic excitation/inhibition (Schlingloff et al., 2014).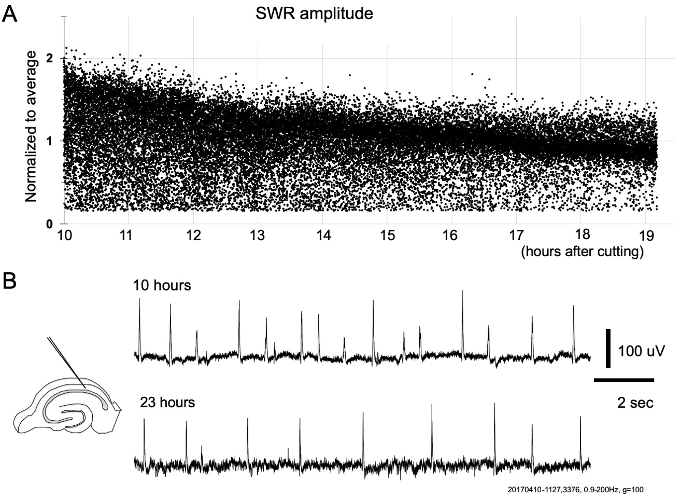
Figure 5. Recording time and amplitude declining. A. Continuous recording (>10 h) of SWR amplitude from a slice of P30 animal. The recording started at 10 h after slicing. SWR amplitude gradually reduced over the entire period. The reduction rate was to one-half in 5 h. B. Traces from 10 and 23 h after slicing. Note that the SWRs with lower amplitude displayed a disproportionate reduction after may hours. In this Figure we show SWR signals from striatum oriens (signal polarity is reversed from that in Figure 4).
Recipes
- Slice cutting ACSF solution (200 ml per experiment, made before use from two stock solutions and 0.43 g NaHCO3 powder)
Final composition: Sucrose (252.0 mM); KCl (3.0 mM); MgSO4 (4.0 mM); CaCl2 (1.0 mM); Glucose (10.0 mM); NaHCO3 (10.0 mM); NaH2PO4 (1.25 mM); HEPES (5.0 mM); Saturated with carbogen; pH = 7.4.
Stock 1: Sucrose
- 177 g sucrose, dissolved in 1,000 ml hot (65 °C) distilled water
- Then add distilled water to a final volume of 2,000 ml
- Keep the stock at 4 °C before use
Stock 2: Cutting ACSF
- Dissolve the chemicals in the following sequence in 180 ml distilled water
Reagent (Molecular weight) Grams for 200 ml KCl (74.55) 0.45 g MgSO4 (120.37) 0.96 g Glucose (180.8) 3.60 g NaH2PO4 (119.98) 0.30 g HEPES (238.3) 2.38 g CaCl2·2H2O (147.02) 0.29 g - After all chemicals are fully dissolved, add distilled water to final value of 200 ml
- Keep the stock in 4 °C refrigerator before use
Stock 3: NaHCO3 powder
Prepare 10 of 0.43 g NaHCO3 powder (each store in a 1.5 ml Eppendorf tube) for 10 experiments
This significantly reduce the preparation time before each experiment
Making 200 ml slice cutting ACSF solution before each experiment:
- Take 180 ml of sucrose stock solution and add 20 ml of cutting stock solution to reach a final volume of 200 ml. Vigorously stir-mix the two solutions with a magnetic stir bar.
- Put the NaHCO3 powder in one Eppendorf tube (0.43 g) into the 200 ml slice cutting ACSF. Add the powder slowly while vigorously stirring until all the fine crystals of NaHCO3 are dissolved. Blow carbogen with a fish tank air stone for 3 min to ensure full saturation of O2/CO2. Cool the solution down to 4 °C before use.
Important: Measure the temperature of the cutting solution before use. The optimum temperature for slicing is 3-6 °C. Because both stock solutions were at 4 °C before mixing, it should take only about 10 min to cool down the cutting ACSF solution. - Incubation ACSF (2,000 ml per each experiment, made from the stock solution below and 4.37 g of NaHCO3 powder).
- 177 g sucrose, dissolved in 1,000 ml hot (65 °C) distilled water
- Incubation/recording ACSF
Stock solution for incubation/recording ACSF
- Dissolve the chemicals in the following sequence in 1,600 ml distilled water
Reagent (Formula weight) Grams for 2,000 ml NaCl (58.44) 144.0 g KCl (74.55) 4.5 g MgSO4 (120.37) 4.8 g Glucose (180.8) 36.0 g NaH2PO4 (119.98) 3.0 g HEPES (238.3) 23.8 g CaCl2·H2O (147.02) 5.9 g - After all chemicals are fully dissolved, add distilled water to reach a final volume of 2,000 ml
- Keep the stock solution at 4 °C and use within one month. The 2,000 ml ACSF stock solution is good for 10 experiments
- Prepare 20 of 4.37 g NaHCO3 powder (each store in a 17 ml capped culture tube) for 20 experiments. This significantly reduce the preparation time before each experiment
Making 2,000 ml incubation ACSF before each experiment:
- Take 200 ml incubation ACSF stock and dissolve in 1,600 ml warm (35 °C) distilled water
- Stir-mix the solution with a magnetic stir bar
- Add NaHCO3 powder from one tube (4.37 g) into the solution while vigorously stirred until fine crystals of NaHCO3 are fully dissolved
- Add warm (35 °C) distilled water to a reach a final volume of 2,000 ml
- Bubble with carbogen for 3 min to ensure fully saturated with O2/CO2
- Dissolve the chemicals in the following sequence in 1,600 ml distilled water
Acknowledgments
Supported by NIH R03AG061645, Georgetown University Medical Center Dean's Toulmin grant FY2017, FY 2019.
Competing interests
The authors declare no competing financial interests.
Ethics
The procedures for using animals are strictly following a protocol approved by the Institutional Animal Care and Use Committee (IACUC) at Georgetown University Medical Center following the NIH guidelines.
References
- Aghajanian, G. K. and Rasmussen, K. (1989). Intracellular studies in the facial nucleus illustrating a simple new method for obtaining viable motoneurons in adult rat brain slices. Synapse 3(4):331-338.
- Behrens, C. J., van den Boom, L. P., de Hoz, L., Friedman, A. and Heinemann, U. (2005). Induction of sharp wave-ripple complexes in vitro and reorganization of hippocampal networks. Nat Neurosci 8(11): 1560-1567.
- Brahma, B., Forman, R. E., Stewart, E. E., Nicholson, C. and Rice, M.E. (2000). Ascorbate inhibits edema in brain slices. J Neurochem 74:1263-1270.
- Buzsáki, G. (2015). Hippocampal sharp wave-ripple: A cognitive biomarker for episodic memory and planning. Hippocampus 25(10): 1073-1188.
- Caccavano, A., Bozzelli, P. L., Forcelli, P. A., Pak, D. T. S., Wu, J. Y., Conant, K. and Vicini, S. (2020). Inhibitory parvalbumin basket cell activity is selectively reduced during hippocampal sharp wave ripples in a mouse model of familial Alzheimer's disease. J Neurosci doi: 10.1523/JNEUROSCI.0425-20.2020.
- Colgin, L. L., Kubota, D., Jia, Y., Rex, C. S. and Lynch, G. (2004). Long-term potentiation is impaired in rat hippocampal slices that produce spontaneous sharp waves. J Physiol 558(Pt 3): 953-961.
- Ego-Stengel, V. and Wilson, M.A. (2010). Disruption of ripple-associated hippocampal activity during rest impairs spatial learning in the rat. Hippocampus 20(1):1-10.
- Hájos, N., Ellender, T. J., Zemankovics, R., Mann, E. O., Exley, R., Cragg, S. J., Freund, T. F. and Paulsen, O. (2009). Maintaining network activity in submerged hippocampal slices: importance of oxygen supply. Eur J Neurosci 29(2): 319-327.
- Jiang, H., Liu, S., Geng, X., Caccavano, A., Conant, K., Vicini, S. and Wu, J. (2018). Pacing hippocampal sharp-wave ripples with weak electric stimulation. Front Neurosci 12: 164.
- Ji, D. and Wilson, M. A. (2007). Coordinated memory replay in the visual cortex and hippocampus during sleep. Nat Neurosci 10(1): 100-107.
- Keller, M. K., Draguhn, A., Both, M. and Reichinnek, S. (2015). Activity-dependent plasticity of mouse hippocampal assemblies in vitro. Front Neural Circuits 9: 21.
- Kubota, D., Colgin, L. L., Casale, M., Brucher, F. A. and Lynch, G. (2003). Endogenous waves in hippocampal slices. J Neurophysiol 89(1): 81-89.
- Lee, A. K. and Wilson, M. A. (2002). Memory of sequential experience in the hippocampus during slow wave sleep. Neuron 36(6): 1183-1194.
- Li, P., Geng, X., Jiang, H., Caccavano, A., Vicini, S. and Wu, J. Y. (2019). Measuring sharp waves and oscillatory population activity with the genetically encoded calcium indicator GCaMP6f. Front Cell Neurosci 13: 274.
- Maier, N., Morris, G., Johenning, F. W. and Schmitz, D. (2009). An approach for reliably investigating hippocampal sharp wave-ripples in vitro. PLoS One 4(9): e6925.
- Maier, N., Nimmrich, V. and Draguhn, A. (2003). Cellular and network mechanisms underlying spontaneous sharp wave-ripple complexes in mouse hippocampal slices. J Physiol 550(Pt 3): 873-887.
- Miyawaki, T., Norimoto, H., Ishikawa, T., Watanabe, Y., Matsuki, N. and Ikegaya, Y. (2014). Dopamine receptor activation reorganizes neuronal ensembles during hippocampal sharp waves in vitro. PLoS One 9(8): e104438.
- Oakley, H., Cole, S. L., Logan, S., Maus, E., Shao, P., Craft, J., Guillozet-Bongaarts, A., Ohno, M., Disterhoft, J., Van Eldik, L., Berry, R. and Vassar, R. (2006). Intraneuronal beta-amyloid aggregates, neurodegeneration, and neuron loss in transgenic mice with five familial Alzheimer's disease mutations: potential factors in amyloid plaque formation. J Neurosci 26(40): 10129-10140.
- Schlingloff, D., Káli, S., Freund, T. F., Hájos, N., Gulyás, A.I. (2014). Mechanisms of sharp wave initiation and ripple generation. J Neurosci 34(34):11385-11398.
- Sun, Z. Y., Bozzelli, P. L., Caccavano, A., Allen, M., Balmuth, J., Vicini, S., Wu, J. Y. and Conant, K. (2018). Disruption of perineuronal nets increases the frequency of sharp wave ripple events. Hippocampus 28(1): 42-52.
- Tanaka, Y., Furuta, T., Yanagawa, Y. and Kaneko, T. (2008). The effects of cutting solutions on the viability of GABAergic interneurons in cerebral cortical slices of adult mice. J Neurosci Methods 171:118-125.
- Ting, J. T., Daigle, T. L., Chen, Q. and Feng, G. (2014). Acute brain slice methods for adult and aging mice: application of targeted patch clamp analysis and optogenetics. Methods Mol Biol 1183:221-242.
- Ting, J. T., Lee, B. R., Chong, P., Soler-Llavina, G., Cobbs, C., Koch, C., Zeng, H. and Lein, E. (2018). Preparation of Acute Brain Slices Using an Optimized N-Methyl-D-glucamine Protective Recovery Method. J Vis Exp (132):53825.
- Wilson, M. A. and McNaughton, B. L. (1994). Reactivation of hippocampal ensemble memories during sleep. Science 265(5172): 676-679.
- Wu, C., Luk, W. P., Gillis, J., Skinner, F. and Zhang, L. (2005). Size does matter: generation of intrinsic network rhythms in thick mouse hippocampal slices. J Neurophysiol 93(4): 2302-2317.
- Ye, J. H., Zhang, J., Xiao, C. and Kong, J. Q. (2006). Patch-clamp studies in the CNS illustrate a simple new method for obtaining viable neurons in rat brain slices: glycerol replacement of NaCl protects CNS neurons. J Neurosci Methods 158(2):251-259.
Article Information
Copyright
© 2020 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Liu, L., Zhou, X. and Wu, J. (2020). Preparing Viable Hippocampal Slices from Adult Mice for the Study of Sharp Wave-ripples. Bio-protocol 10(19): e3771. DOI: 10.21769/BioProtoc.3771.
Category
Neuroscience > Basic technology > Acute slice preparation
Cell Biology > Tissue analysis > Physiology
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









