- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
An Efficient Inoculation Technique to Assess the Pathogenicity of Pantoea Species Associated to Bacterial Blight of Rice
Published: Vol 10, Iss 17, Sep 5, 2020 DOI: 10.21769/BioProtoc.3740 Views: 5573
Reviewed by: Mohammad Malek Faizal AziziAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Quantitative Estimation of Auxin, Siderophore, and Hydrogen Cyanide Production in Halo and Drought-Tolerant Bacterial Isolates for Cucumber Growth
Zeinab Fotoohiyan and Ali Salehi Sardoei
Oct 5, 2025 1389 Views
Reproducible Emu-Based Workflow for High-Fidelity Soil and Plant Microbiome Profiling on HPC Clusters
Henrique M. Dias [...] Christopher Graham
Jan 20, 2026 524 Views
Abstract
Bacteria blight diseases of rice due to several genera of pathogenic bacteria are one of the major constraints worldwide for rice production. The disease can be best managed through host plant resistance sources. For most of these bacteria such as Xanthomonas oryzae pv. oryzae, X. oryzae pv. oryzicola, Pseudomonas fuscovaginae, Burkholderia glumae, Burkholderia gladioli and Acidovorax avenae subsp. avenae, specific diagnostic techniques that include molecular and pathogenicity tests have been developed.
However, for Pantoea spp., information on pathogenicity assay is very limited and protocols used are not uniform. Most authors use the leaf clipping method. In this paper, we describe the protocol for mechanical inoculation of rice seedlings aged 35 days. The method consists of infiltrating bacterial suspensions at concentrations of 108 CFU/ml, with a needleless syringe into the intercellular and interveinal spaces of rice leaves underside at about 4-5 cm below the leaf tip.
This method can be used for a standardized pathogenicity assessment, germplasm resistance evaluation for identifying and characterizing resistance sources.
Background
The rice species Oryza sativa is the most widely consumed staple food for a large part of the world's human population, especially in Asia and Africa (Nwanze et al., 2006; Somado et al., 2008; Gnanamanickam, 2009). Its production contributes to fight against food insecurity and reduce poverty. In Africa, rice self-sufficiency remains so far impossible. This situation is explained by several socio-economic, abiotic and biotic factors that constitute a brake for the rice development sector. Among the biotic stresses, plant diseases such as bacterial blights (BB) represent a threat to production (Sere et al., 2005; Verdier et al., 2012).
BB of rice caused by Pantoea spp. is becoming a significant biotic constraint for rice production (Doni et al., 2019). Indeed, since 2017, several cases of these diseases have been reported worldwide (Kini, et al., 2017a and 2017b; Aksoy and Boluk, 2019; Azizi et al., 2019; Doni et al., 2019). The bacterial complex composed of 27 species is widely distributed around the world. Its biological characteristics (versatile and ubiquitous) have enabled its establishment in several agro-ecosystems spanning from regions with high precipitation to semi-arid and arid irrigated zones. Likewise, management of this bacterial threat is challenging because the bacteria can persist in soil, irrigation water, seed or plant debris (Walterson and Stavrinides, 2015; Weller-Stuart et al., 2017). Furthermore, its parasitic association with rice and other important crops enhances possible interactions with other rice pathogenic microorganisms (Dossou and Silue, 2018) adding another constraint to its management. Undoubtedly, using Pantoea-resistant cultivars would be the most cost-effective and sustainable control method (Health et al., 2018).
Several pathogenicity assays for resistance/susceptibility characterization in germplasm and breeding lines exist that use inoculation techniques such as leaf clipping, inocula spraying or injection (Gonzalez et al., 2007; Nandakumar et al., 2009; Adorada et al., 2013; Yang and Bogdanove, 2013; Weny et al., 2019). These were applied differently with the six major rice bacterial pathogens, namely Xanthomonas oryzae pv. oryzae (Xoo), X. oryzae pv. oryzicola (Xoc), Pseudomonas fuscovaginae, Burkholderia glumae, B. gladioli and Acidovorax avenae subsp. avenae (Cui et al., 2016).
Depending on the etiology of some bacteria and the physiology of the host plant, certain bacteria inoculation techniques are more suitable and produce consistent results when screening germplasm. For example, leaf infiltration is more adapted for Xoc while leaf clipping is more appropriate for Xoo (Yang and Bogdanove, 2013). However, for Pantoea spp., the information for a standard pathogenicity protocol does not exist. Few available publications refer only to leaf clipping that is briefly described in few new disease reports (Mondal et al., 2011; González et al., 2014; Egorova et al., 2015; Aksoy and Boluk, 2019; Azizi et al., 2019). In addition, the literature lacks a standard disease resistance screening protocol. To fill this gap, the leaf clipping, spraying and injection of bacterial inocula into rice leaves were therefore tested in our lab. The first two techniques revealed some difficulties: for leaf clipping, it is mainly about (i) less complete symptoms development (ii) difficulty to monitor and evaluate symptom progression (iii) possible confusion of disease symptoms with abiotic necrosis/senescence. In addition, results gathered in our laboratory indicated that the use of the leaf clipping technique does not lead to symptom development for P. stewarthii and for P. agglomerans or to symptoms that are indistinguishable from natural leaf senescence ones (Figure 1). In addition, for the inoculum spraying technique, difficulties were about (i) persistent contaminations of inoculation chambers and the development unwanted symptom (off-type). When the appearance and progression of symptoms obtained with these techniques were somehow interesting, their evaluation and scoring were difficult to perform.
The third technique, i.e., injection of inoculum with a blunt-ended (needleless) syringe proved to be the most effective and suitable (Kini et al., 2017a and 2017b). An accurate evaluation of pathogenicity phenotype are a pre-requisite for the diagnosis of plant pathogens with reference to Koch postulate, but also a better assessment under artificial infection conditions of resistance/susceptibility of plants to pathogens. This paper therefore aims at standardizing and making available the protocol described by Kini et al. (2017a and 2017b).
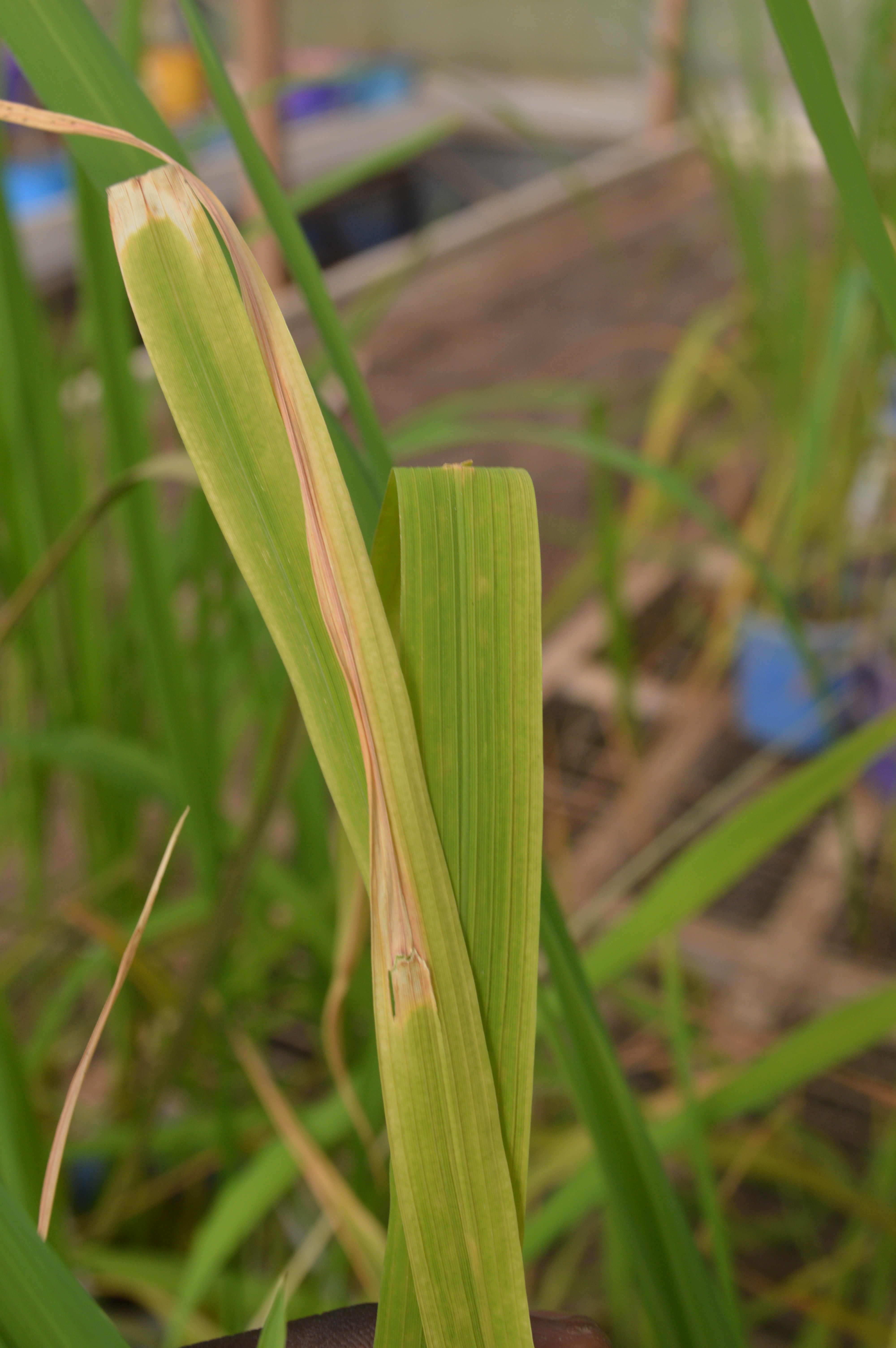

Figure 1. Typical Pantoea-induced symptoms developed on leaves inoculated with either P. stewartii or P. agglomerans using the leaf infiltration method (front leaf) compared to the absence of symptoms on rice leaves inoculated using the leaf clipping method (back leaf)
Materials and Reagents
- Latex gloves
- 2 ml micro-centrifuge tubes (Dominique DUTSCHER, catalog number: 0 33297 )
- 1 liter plastic pots
- 50 ml Falcon tube (Sigma, catalog number: CLS430828 )
- 500 ml wash bottle
- Plates 96 wells (VWR, catalog number: 735-0083 )
- 10, 20, 200, and 1,000 μl tips (Eppendorf, catalog number: 2231300008 )
- Paper towel
- Petri dishes
- Inoculating loop
- Marker
- Parafilm
- Fertilizer (urea and NPK 15-15-15 beaded and pelletized)
- Blunt-ended (needleless) syringes
- Sterilized field Soil
- Brand® UV cuvettes macro, chamber volume 2.5-4.5 ml (Sigma, catalog number: Z637157 )
- Icebox
- Plant materials: Rice seeds, including control accessions (susceptible, partially and highly resistant)
- Pantoea spp. Strains
- Hydrochloric acid (HCl)
- Alcohol
- Sterilized ultra-pure water
- Peptone (EuroMedex, catalog number: P3300 )
- Beef extract (Merck, CAS no. 68990-09-0)
- Sucrose (Merck, CAS no. 57-50-1)
- Glucose (Sigma, CAS no. 50-99-7)
- Yeast extract (Merck, CAS no. 8013-01-2)
- Tryptone (Merck, CAS no. 91079-40-2)
- Glutamic acid (Merck, CAS no. 56-86-0)
- Peptone Sucrose Agar (PSA plates) (see Recipes)
- 10 N HCl (see Recipes)
- 70% ethanol (see Recipes)
- Inoculum (see Recipes)
Equipment
- Tweezers
- Autoclave machine
- Nethouse, greenhouses/screenhouses/growth chambers
- Mortars and pestles
- MilliQ sterile ultrapure water system
- Laminar flow cabinet
- 10, 20, 200, and 1,000 μl pipettes
- -20 °C or deep freezer
- 28 °C growing incubator
- Spectrophotometer
- Vortexer
- pH meter
- Microcentrifuge
- Precision balance
Procedure
- Rice plant growth conditions
- Grow rice plants under semi-controlled environment greenhouses/screenhouses/growth chambers with the following conditions during: approximately 12 h of light, 25 ± 5 °C (day) and 20 ± 5 °C (night), and 75% relative humidity.
Note: Use the following recommended rice varieties as checks: C101A51, Azucena, Sahel 108, Moroberekan. - Make 3 holes in 20 cm diameter pots containing autoclaved (121 °C for 20 min) field soil or pre-conditioned potting compost.
- Sow 3 surface disinfected seeds per hole, use 3 holes per pots and 3 pots/rice accession (27 seeds/variety/replication).
- About 30-35 days after sowing, inoculate the plants as described below.
- Apply urea (recommended quantity for the rice varieties used) after inoculation for optimal plant growth.
- Grow rice plants under semi-controlled environment greenhouses/screenhouses/growth chambers with the following conditions during: approximately 12 h of light, 25 ± 5 °C (day) and 20 ± 5 °C (night), and 75% relative humidity.
- Inoculum preparation and plant inoculation (see Video 1)
- Streak the bacterial strains kept at -20 °C or -80 °C freezer on freshly prepared PSA agar plates and incubate them at 28 °C for 1-3 days. Use a strain diagnosed as Pantoea spp. as control.
- Then, scrape off the cells from the plates and suspend them in MilliQ sterile ultrapure water.
- Adjust the concentration of this suspension at 108 cells/ml (OD = at 600 nm) using a spectrophotometer.
- Then, when plants are 4-5-week-old, use a needleless syringe to pump inoculum and infiltrate 1 to 2 ml of the bacterial suspension inoculum into 3-6 youngest fully expanded leaves using the needleless syringe. Infiltrate leaves at about 3-5 cm below the tip of the leaves (Figure 2).
- Perform the infiltration on the underside of the leaves by pressing the mouth of the syringe on the leaf surface.
Note: The gloved fingers should gently block and support the opposite side of the leaf and gently press the plunger. Take care not to crush the leaf. Also, be sure not to overlap the midrib as this will raise the syringe and break the necessary seal. - Then, gently wipe the leaf surface with a sterilized paper towel to remove the excess liquid from the leaf surface.
- Finally, hold the leaf briefly against the light to confirm that infiltration was successful as such infiltrated spots will show water-soaking features. Use sterile distilled water as a negative control.Video 1. Peptone Sucrose Agar (PSA plates) preparation, plating and growth of bacteria, inoculum preparation and inoculation of rice plants
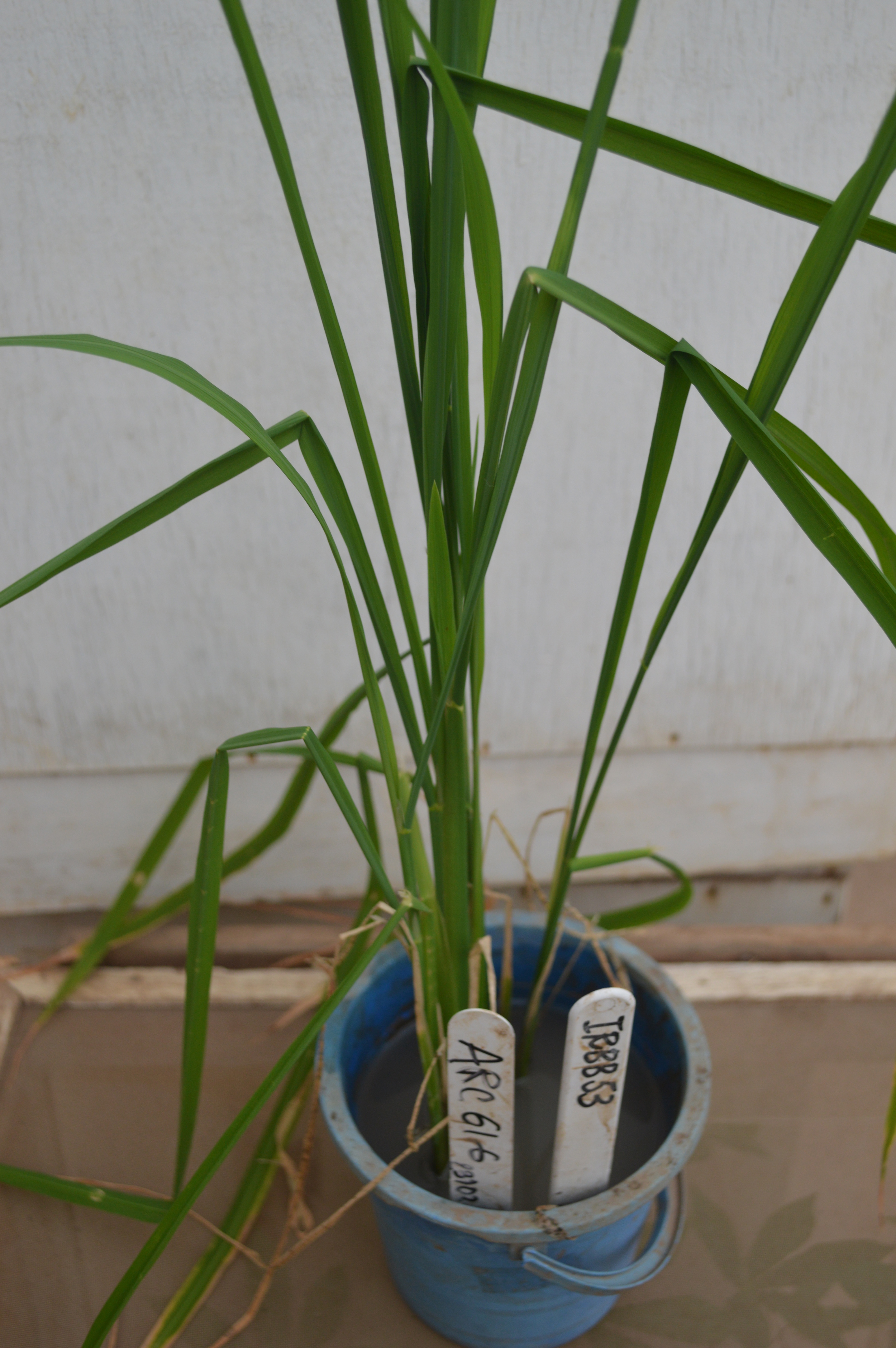

Figure 2. Rice plants aged 4-5 weeks after sowing that were just inoculated (note that the inoculated leaves that are bending)
- Symptoms progression
After inoculation, monitor symptom development on a daily basis.
Note: In susceptible reactions, a spectacular water-soaking will appear 4-5 days after inoculation (DAI) at the infiltration spots of the leaves while it is much weaker in resistant reaction. About 8 DAI, the resulting necrotic lesions at the inoculation site will increase lengthwise along the principal veins towards to the tip of the leaf. These lesions later expand and turn from straw yellow to light brown color and ultimately develop into typical blight symptoms at 15 to 21 DAI (Figure 3).
Figure 3. Scoring of the Pantoea blight symptom progression on rice plants inoculated at 35 days after sowing using a scale we developed. A = 2-4 DAI: Water-soaking at the inoculation point. B = 5-7 DAI: Yellow necrotic lesions at the inoculation points spreading in both directions (length at width). C = 10-12 DAI: Yellow necrotic lesions at the inoculation points increasing lengthwise along the principal veins towards to the tip of the leaf. D = 14-17 DAI: Ascending asymmetrical yellow necrosis of the tips of the leaves on 0.5 to 1 cm. E = 21-23 DAI: Presence of symptoms on three leaf areas, i) a grey necrotic area (at the tips of the leaves); ii) green area, iii) a yellowish area (which separates area i from ii). Area ii can evolve and areas i and ii can partially merge. F = 28-30 DAI: Total grey necrotic leaf area spreading from the tip of the leaf to the inoculation point. - Symptoms scoring
Pathogenicity assessment was made as follow: record the disease severity at 14 and 21 DAI according to the symptom severity scale below (Figure 4):- Water-soaking at the inoculation point.
- Yellow necrotic lesions at the inoculation point spreading in both directions (length and width).
- Yellow necrotic lesions at the inoculation point increasing lengthwise along the principal veins and towards to the tip of the leaf.
- Ascending asymmetrical yellow necrosis of 0.5 to 1 cm on the tip of the leaf.
- Presence of symptoms on three leaf areas, i) grey necrotic area (on the end of the leaf); ii) green area, iii) yellowish area (which separates area i from area ii). Green area can evolve and partially merge with the necrotic area.
- Total grey necrotic leaf area seen from the tip of the leaf to the inoculation point.
- Total necrotic greyish/brown leaf with leaf curling.
Accessions are then classified for resistance as follow:
•Resistant (R)
•Moderate Resistant (MR)
•Moderate Susceptible (MS)
•Susceptible (S)
Resistance/susceptibility phenotypes of plants from pure lines will be more or less the same compared to those of mixed seeds. In all cases, select the most representative response for each accession.
Pathogenicity of the isolates were classified as non-, moderate and fully pathogenic. As regards to their virulence, they were classified as avirulent when they induced only R and MR pathogenicity phenotypes and virulent when they induced MS and S ones.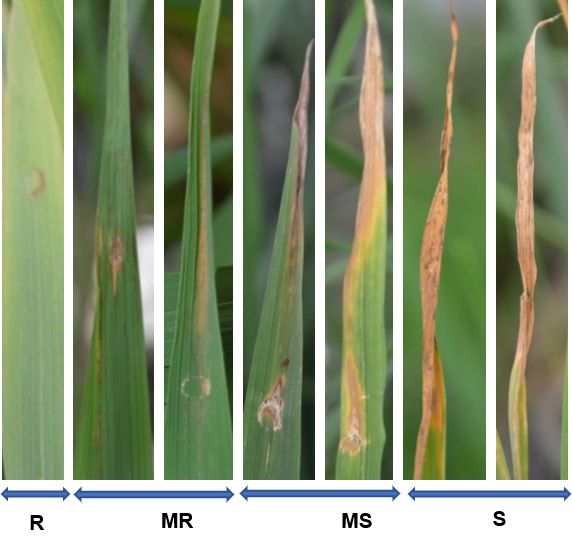
Figure 4. Scale developed and used to score symptom severity
- Assessment of bacterial population
At 7, 14 and 21 DAI, bacterial populations can be assessed as a function of colony-forming units (cfu) recovered from the inoculated leaves. In practice, cut 1 cm2 leaf pieces 2 cm below and above the inoculation points and make two corresponding samples. Ground these pieces in distilled sterile water (1 ml per leaf piece) using sterilized mortar and pestle or automatic ball mills. Make a 10 fold dilution of the resulting leaf saps and plate in triplicate on PSA or Pantoea Genus specific Agar (PGSA) described by Kini et al. (2019). Count the colonies on the three plates of each dilution and convert the data into cfu per leaf. Finally, perform, Analysis of Variances (ANOVA) on the collected data and calculate a correlation function.
Notes
- Other media are also suitable for culturing Pantoea spp. and include nutrient broth or NB (per liter: beef extract, 3 g; peptone, 5 g; sucrose, 10 g), glucose yeast extract or GYE (per liter: glucose, 20 g; yeast extract, 10 g), and Tryptone Sucrose or TS (per liter: Tryptone, 10 g; sucrose, 10 g; glutamic acid, 1 g). Solidification of each medium is achieved by supplementation with 15 g agar/L.
- To avoid contamination and ensure purity of bacterial strains, the PGSA medium can be used to select and purify the bacteria strains.
- The genus Pantoea is composed of about 27 species. Before inoculation, ensure that the species of the strains used is known. To this end, a multiplex PCR (Kini et al., 2018) scheme can therefore be used.
- Certain Pantoea spp. are quarantine pests in several countries (Health et al., 2018). So, the manipulation of such species may be restricted by biosafety measures. Always, manipulate the bacteria in accordance with your country’s legislation.
- Resistance or susceptibility depends on specific interactions between the rice plant and Pantoea spp. The choice of resistant or susceptible control accessions may also depend on the species and races of Pantoea spp. and the objectives of the experiment to be carried out.
- Plants of different stages may be used for different purposes, such as measuring plant disease resistance that is developmentally regulated.
- For easiest syringe inoculum infiltration in leaves, maintain rice plants at high humidity and inoculate at the beginning of the light cycle as this keeps stomata open.
Recipes
- Peptone Sucrose Agar (PSA plates) (see Video 1)
- In 1 L of MilliQ water, add 10 g peptone, 10 g sucrose, 16 g agar, and 1 g glutamic acid
- Adjust pH to 7.1 ± 0.2 using 1 M KOH and NaOH buffers
- Autoclave at 121 °C for 20 min
- Let the medium cool down but not solidify, then pour it into Petri dishes
- 10 N HCl
To prepare 1,000 ml of 10 M HCl, proceed as follow:- Poor 83 ml d’ HCl in a test tube
- Complete to 1,000 ml of water
- 70% ethanol
Using Gay-Lussac dilution table and 96° ethanol available in the laboratory:- Poor 40.85 ml of this ethanol were poured in an test tube
- Complete to 100 ml with distilled sterile water
- Inoculum
- Collect bacterial colonies in 50 ml Falcon tube containing 45 ml of MilliQ water
- Then, mix and take 2 ml inoculum in a Brand® UV cuvettes macro, chamber volume 2.5-4.5 ml and introduce it in a spectrophotometer and take the reading at OD600
- Then, adjust the inoculum by diluting it in MilliQ sterile water
- For each dilution, make sure the concentration is of 108 CFU/ml
Acknowledgments
This protocol was developed from procedures used previously in different laboratories (Yang and Bogdanove, 2013) for inoculation and virulence assay for bacterial blight and bacterial leaf streak of rice. Dr. Kossi Kini benefited a PhD scholarship from the French Research Institute for Sustainable Development (IRD) within the program “Allocation de Recherche pour une Thèse au Sud” (ARTS), a grant from the International Foundation for Science (IFS) and financial support from the Ministry of Foreign Affairs (Japan) and the Global Rice Science Partnership (GRiSP) through AfricaRice. All these institutions are gratefully acknowledged.
Competing interests
Authors have no competing interest.
References
- Adorada, D. L., Stodart, B. J., Cruz, C. V., Gregorio, G., Pangga, I. and Ash, G. J. (2013). Standardizing resistance screening to Pseudomonas fuscovaginae and evaluation of rice germplasm at seedling and adult plant growth stages. Euphytica 192(1): 1-16.
- Aksoy, H. M. and Boluk, E. (2019). First report of Pantoea agglomerans on Oryza sativa in Turkey. J Plant Pathol 101(2): 449-449.
- Azizi, M. M. F., Zulperi, D., Rahman, M. A. A., Abdul-Basir, B., Othman, N. A., Ismail, S. I., Hata, E. M., Ina-Salwany, M. Y. and Abdullah, M. A. F. (2019). First report of Pantoea ananatis causing leaf blight disease of rice in Peninsular Malaysia. Plant Disease 103(8): 2122.
- Cui, Z., Ojaghian, M. R., Tao, Z., Kakar, K. U., Zeng, J., Zhao, W., Duan, Y., Vera Cruz, C. M., Li, B., Zhu, B. and Xie, G. (2016). Multiplex PCR assay for simultaneous detection of six major bacterial pathogens of rice. J Appl Microbiol 120(5): 1357-1367.
- Doni, F., Suhaimi, N. S. M., Mohamed, Z., Ishak, N. and Mispan, M. S. (2019). Pantoea: a newly identified causative agent for leaf blight disease in rice. J Plant Dis Protect 126(6): 491-494.
- Dossou, B. and Silue, D. (2018). Rice pathogens intercepted on seeds originating from 11 African countries and from the USA. Seed Sci Technol 46(1): 31-40.
- Egorova, M., Mazurin, E. and Ignatov, A. N. (2015). First report of Pantoea ananatis causing grain discolouration and leaf blight of rice in Russia. New Dis Rep 32: 21-21.
- Gnanamanickam, S. S. (2009). Rice and its importance to human life. In: Biological control of rice diseases. Springer, pp. 1-11.
- González, A. D., Franco, M. A., Contreras, N., Galindo-Castro, I., Jayaro, Y. and Graterol, E. (2014). First report of Pantoea agglomerans causing rice leaf blight in Venezuela. Plant Dis 99(4): 552-552.
- Gonzalez, C., Szurek, B., Manceau, C., Mathieu, T., Sere, Y. and Verdier, V. (2007). Molecular and pathotypic characterization of new Xanthomonas oryzae strains from West Africa. Mol Plant Microbe Interact 20(5): 534-546.
- Kini, K., Agnimonhan, R., Afolabi, O., Milan, B., Soglonou, B., Gbogbo, V., Koebnik, R. and Silué, D. (2017a). First report of a new bacterial leaf blight of rice caused by Pantoea ananatis and Pantoea stewarthii in Benin. Plant Disease 101(1): 242-242.
- Kini, K., Agnimonhan, R., Afolabi, O., Soglonou, B., Silué, D. and Koebnik, R. (2017b). First report of a new bacterial leaf blight of rice caused by Pantoea ananatis and Pantoea stewarthii in Togo. Plant Dis 101(1): 241.
- Kini, K., Agnimonhan, R., Dossa, R., Silué, D. and Koebnik, R. (2018). A diagnostic multiplex PCR scheme for identification of plant-associated bacteria of the genus Pantoea. bioRxiv: 456806.
- Kini, K., Dossa, R., Dossou, B., Mariko, M., Koebnik, R. and Silué, D. (2019). A semi-selective medium to isolate and identify bacteria of the genus Pantoea. J Gen Plant Pathol 85(6): 424-427.
- Mondal, K. K., Mani, C., Singh, J., Kim, J. G. and Mudgett, M. B. (2011). A new leaf blight of rice caused by Pantoea ananatis in India. Plant Dis 95(12): 1582-1582.
- Nandakumar, R., Shahjahan, A. K. M., Yuan, X. L., Dickstein, E. R., Groth, D. E., Clark, C. A., Cartwright, R. D. and Rush, M. C. (2009). Burkholderia glumae and B. gladioli cause bacterial panicle blight in rice in the Southern United States. Plant Dis 93(9): 896-905.
- Nwanze, K. F., Mohapatra, S., Kormawa, P., Keya, S. and Bruce-Oliver, S. (2006). Rice development in sub-Saharan Africa. J Sci Food Agr 86(5): 675-677.
- Health, E. Panel O. P., Jeger, M., Bragard, C., Candresse, T., Chatzivassiliou, E., Dehnen-Schmutz, K., Gilioli, G., Grégoire, J. C., Jaques Miret, J. A., MacLeod, A., Navajas Navarro, M., Niere, B., Parnell, S., Potting, R., Rafoss, T., Rossi, V., Urek, G., Van Bruggen, A., Van der Werf, W., West, J., Winter, S., Manceau, C., Pautasso, M. and Caffier, D. (2018). Pest categorisation of Pantoea stewartii subsp. stewartii. EFSA J 16(7): e05356.
- Sere, Y., Onasanya, A., Verdier, V., Akator, K., Ouedraogo, L. S., Segda, Z., Coulibaly, M. M., Sido, A. Y. and Basso, A. (2005.) Rice bacterial leaf blight in West Africa: preliminary studies on disease in farmers’ fields and screening released varieties for resistance to the bacteria. Asian J Plant Sci 4: 577-579.
- Somado, E. A., Guei, R. G. and Nguyen, N. (2008). Overview: rice in Africa. Africa Rice Center, Bouaké.
- Verdier, V., Cruz, C. V. and Leach, J. E. (2012). Controlling rice bacterial blight in Africa: needs and prospects. J Biotech 159(4): 320-328.
- Walterson, A. M. and Stavrinides, J. (2015). Pantoea: insights into a highly versatile and diverse genus within the Enterobacteriaceae. FEMS Microbiol Rev 39(6): 968-984.
- Weller-Stuart, T., De Maayer, P. and Coutinho, T. (2017). Pantoea ananatis: genomic insights into a versatile pathogen. Mol Plant Pathol 18(9): 1191-1198.
- Weny, Safni, I., Lisnawita and Lubis, K. (2019). Screening for disease resistance in rice varieties against bacterial panicle blight disease (Burkholderia glumae) in Northern Sumatra of Indonesia. IOP Conference Series: Earth and Environmental Science 260: 012118.
- Yang, B. and Bogdanove, A. (2013). Inoculation and virulence assay for bacterial blight and bacterial leaf streak of rice. Methods Mol Biol 956: 249-255.
Article Information
Copyright
© 2020 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Kini, K., Agnimonhan, R., Wonni, I. and Silue, D. (2020). An Efficient Inoculation Technique to Assess the Pathogenicity of Pantoea Species Associated to Bacterial Blight of Rice. Bio-protocol 10(17): e3740. DOI: 10.21769/BioProtoc.3740.
Category
Plant Science > Plant immunity > Disease symptom
Plant Science > Plant immunity > Host-microbe interactions
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link