- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
SMART (Single Molecule Analysis of Resection Tracks) Technique for Assessing DNA end-Resection in Response to DNA Damage
Published: Vol 10, Iss 15, Aug 5, 2020 DOI: 10.21769/BioProtoc.3701 Views: 6093
Reviewed by: Alka MehraImre GáspárFarah Haque

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Measuring Real-time DNA/RNA Nuclease Activity through Fluorescence
Paulina Wyrzykowska [...] Richard Chahwan
Nov 5, 2021 3987 Views
Abstract
DNA double strand breaks (DSBs) are among the most toxic lesions affecting genome integrity. DSBs are mainly repaired through non-homologous end joining (NHEJ) and homologous recombination (HR). A crucial step of the HR process is the generation, through DNA end-resection, of a long 3′ single-strand DNA stretch, necessary to prime DNA synthesis using a homologous region as a template, following DNA strand invasion. DNA end resection inhibits NHEJ and triggers homology-directed DSB repair, ultimately guaranteeing a faithful DNA repair. Established methods to evaluate the DNA end-resection process are the immunofluorescence analysis of the phospho-S4/8 RPA32 protein foci, a marker of DNA end-resection, or of the phospho-S4/8 RPA32 protein levels by Western blot. Recently, the Single Molecule Analysis of Resection Tracks (SMART) has been described as a reliable method to visualize, by immunofluorescence, the long 3′ single-strand DNA tails generated upon cell treatment with a S-phase specific DNA damaging agent (such as camptothecin). Then, DNA tract lengths can be measured through an image analysis software (such as Photoshop), to evaluate the processivity of the DNA end-resection machinery. The preparation of DNA fibres is performed in non-denaturing conditions so that the immunofluorescence detects only the specific long 3′ single-strand DNA tails, generated from DSB processing.
Keywords: DNA repairBackground
Genomic instability is one of the enabling characteristics leading to tumour development (Hanahan and Weinberg, 2011). Many sources of DNA damage, either endogenous or exogenous, induce various DNA lesions including double strand breaks (DBSs), which activate the DNA damage response (DDR)–the cell process aimed at preserving DNA integrity. Two main repair pathways are involved in the DSBs repair: non-homologous end joining (NHEJ) and homologous recombination (HR) (Mao et al., 2008). The HR process prevents the loss of genetic information upon DNA damage through a faithful repair using a complementary DNA sequence. By inhibiting the error prone NHEJ and triggering homology-directed DSB repair, DNA end resection has a crucial function in directing the repair pathway choice towards a faithful repair. So, to promote a correct HR, the DSBs are processed through the DNA end-resection machinery which is necessary to generate the long 3′ single-strand DNA (ssDNA) tails essential for the homologous strand invasion. DNA end-resection is a finely regulated process; the first step consists of the recruitment of the MRN complex (MRE11-RAD50-NBSI) and CtIP (RBBP8) onto the DNA lesions to produce short stretches of ssDNA (Stracker and Petrini, 2011) followed by further end-resection mediated by exonuclease I (EXOI) or DNA replication helicase/nuclease 2 (DNA2) in complex with the Bloom syndrome helicases (BLM) (Nimonkar et al., 2011). Subsequently, the replication protein A (RPA) complex binds the ssDNA generated by DNA end-resection, preventing the formation of DNA hairpins (Chen et al., 2013) and to facilitate the loading of RAD51 required for the strand exchange process (Krogh and Symington, 2004).
The evaluation of DNA end-resection is crucial to dissect the molecular mechanisms underlying this finely tuned process and to identify the key players involved. Various methods can be used to assess the DNA end-resection process indirectly, analyzing phosphorylated S4/8 RPA32 as mentioned above, or RAD51 foci, or detection of BrdU in ssDNA following long incubation time (Tkáč et al., 2016). A method to determine the length of resection and the extent of ssDNA at a specific DSB site was developed in 2014 by Tania Paull, although this is based on the expression of an endonuclease (such as AsiSI) to induce DSBs at specific sites within the genome, which is then followed by PCR analysis (Zhou et al., 2014).
The SMART assay was first developed by Huertas and co-authors as a reliable method to measure the length of resected DNA following exposure to any damaging agent in any cellular system, at the level of single molecules (Cruz-García et al., 2014). The assay was based on the previously developed DNA combing assay, which enables the physical stretching of DNA fibres onto a solid support allowing visualization through immunofluorescence (Alfano et al., 2016 and 2017). We recently identified the RNA binding protein HNRNPD as a new player in the HR process and used the SMART assay, along with other techniques, to evaluate DNA end-resection (Alfano et al., 2019). We pulse-labeled HeLa cells with the halogenated IdU (5-Iodo-2′-deoxyuridine)–although any other pyrimidine analogue can be used, such as BrdU (Bromodeoxyuridine)or CldU (5-Chloro-2'-deoxyuridine)–which is incorporated during S phase into the newly synthesized DNA, for approximately 24 h followed by treatment with the DNA damaging agent, camptothecin. At the end of drug incubation time, cells were lysed and the DNA fibres were extracted in non-DNA denaturing conditions (avoiding hydrochloric acid treatment); this is an important criterion for the assay, because the anti-BrdU antibody is able to recognize the DNA-incorporated IdU only when DNA is in the single strand conformation, as is the end-resected DNA. Through the SMART assay, it is possible to visualize the long 3′ single-strand DNA tails and measure the DNA tracts evaluating the efficiency of the endogenous resection machinery without genetic manipulations. Finally, this technique can be used in any model cellular system from bacteria to humans.
The DNA fibres, stretched on a microscope slide, and challenged with the anti-BrdU Ab and the secondary fluorescinated Ab were then visualized through immunofluorescence microscopy. Analysis through the SMART assay allowed us to quantify the amount of resected DNA: by using an image analysis software (Photoshop CS5), we were able to measure the length of the DNA fibre tracts analysing the efficiency of the DNA end-resection machinery.
Materials and Reagents
- 6 cm dish (Corning Inc., catalog number: 430166 )
- Silane Prep-slide (Sigma-Aldrich, catalog number: S4651 )
- HeLa cells (ATCC, CCL2 , catalog number: CCL2 )
- Anti-BrdU clone BU1/75 (ICR1) (AbdSerotec, catalog number: OBT0030CX )
- Alexa Fluor 488-conjugated chicken anti-rat (Thermo Fisher Scientific, catalog number: A-21470 )
- Anti phospho-RPA32 S4/8 antibody (Bethyl Laboratories, catalog number: A300-245 )
- Roswell Park Memorial Institute (RPMI) 1640 Medium (Thermo Fisher Scientific, catalog number: 11875093 )
- 10% fetal bovine serum (FBS), EU Approved (South American) (Thermo Fisher Scientific, GibcoTM, catalog number: 10270-106 )
- Penicillin-streptomycin (Thermo Fisher Scientific, GibcoTM, catalog number: 15070-063 )
- Trypsin-EDTA (Thermo Fisher Scientific, catalog number: 25200056 )
Note: HeLa cells were cultured in RPMI 1640 supplemented with 10% FBS, 1 μg/ml penicillin and 1 µg/ml streptomycin. - 5-Iodo-2′-deoxyuridine (IdU) (Sigma-Aldrich, catalog number: I7125 , storage at 4 °C)
- (S)-(+)-Camptothecin (CPT) (Sigma-Aldrich, catalog number: C9911 , storage at -20 °C)
- Sodium dodecyl sulphate (SDS) (Sigma-Aldrich, catalog number: 71725 , storage at RT)
- TRIS (Sigma-Aldrich, catalog number: 17926 , storage at RT)
- EDTA 0.5 M pH 8.0 (Thermo Fisher Scientific, catalog number: AM9262 , storage at 4°C)
- Sodium phosphate dibasic (Na2HPO4) (Sigma-Aldrich, catalog number: 255793 , storage at RT)
- Potassium phosphate monobasic (KH2PO4) (Sigma-Aldrich, catalog number: P0662 , storage at RT)
- Potassium Cloride (KCl) (Sigma-Aldrich, catalog number: P9541 , storage at RT)
- Hydrochloric acid 39.5% (HCl) (Carlo Erba Reagents, catalog number: 405761 , storage at RT)
- Methanol (Carlo Erba Reagents, catalog number: 528101 , storage at RT)
- Acetic Acid Glacial (Carlo Erba Reagents, catalog number: 401424 , storage at RT)
- Ethanol Absolute (Carlo Erba Reagents, catalog number: 414607 , storage at RT)
- Bovine Serum Albumine (BSA) (Sigma-Aldrich, catalog number: A9418 , storage at 4 °C)
- Sodium Chloride (NaCl) (Sigma-Aldrich, catalog number: S9888 , storage at RT)
- ProLong Gold Antifade Reagent (Thermo Fisher Scientific, catalog number: P36935 , storage at RT)
- Phosphate buffered saline (PBS) (see Recipes)
- Spreading buffer (see Recipes)
Equipment
- Zeiss Axiovert LSM100M confocal microscope (Carl Zeiss, Germany)
- CO2 incubator (Panasonic, catalog number: MCO-18AC )
- Centrifuge
Software
- Adobe Photoshop CS5
- GraphPad Prism 8 software
Procedure
- Representation of SMART assay
Figure 1 shows a representative scheme of the SMART assay.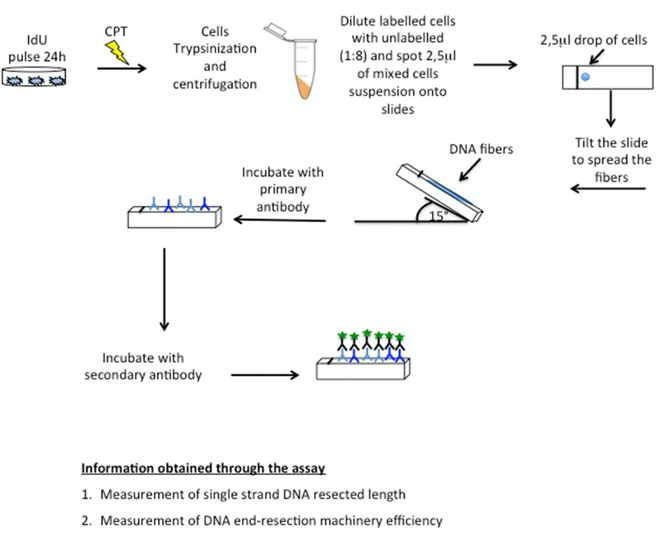
Figure 1. Schematics of SMART assay. First of all, label HeLa cells with IdU for 24 h followed by CPT treatment; at the end of incubation trypsinize cells and dilute them at a 2.5 x 105 cells/ml concentration. Mix labelled and unlabelled cells 1:8 and spot 2.5 μl of this mix onto the upper part of the slide; after incubation with 7.5 μl of spreading buffer, tilt the slide form the end to allow the stretching of the DNA fibres.
Figure 2 shows representative images acquired through confocal microscopy.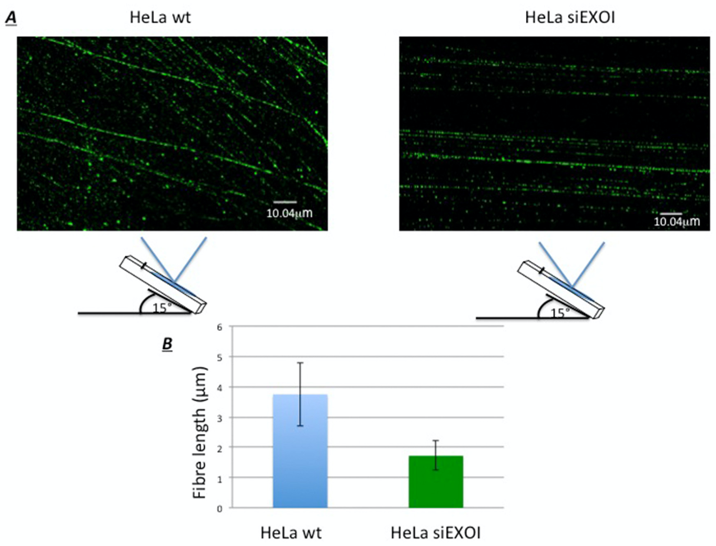
Figure 2. Representative images of the SMART technique. Panel A shows two images from HeLawt or siEXOI cells treated with 1 μM of campthotecin for one hour; following the SMART assay images were analyzed as reported in the protocol through the Fiji software. As shown in panel B the silencing of EXOI, one of the principal exonucleases involved in the generation of single strand DNA upon DNA damage, reduces the length of the processed DNA up to two times. The Figure 3B reports the mean ± standard deviation of the fibre length measured in μm, for each condition. - SMART assay protocol
- Culture HeLa cells in RPMI 1640 medium supplemented with 10% FBS, 1 μg/ml penicillin and 1 µg/ml streptomycin.
- Treat HeLa cells in a 6 cm dish at 80% confluency with 10 μM of IdU (we made a 28 mM IdU stock solution in 0.2 N NaOH) and incubate at 37 °C in a standard 5% CO2 incubator for 24 h.
Note: The length of incubation with BrdU or its analogs such as IdU has to be determined for each different cell type, depending on the cell cycle length. A pulse labelling time needs to allow BrdU incorporation during one complete cell cycle. - Prepare a 6 cm dish of HeLa cells at 80% confluence as non-IdU-marked.
Note: The dilution of IdU marked cell with the non-marked cells is essential to reduce the presence of marked fibres onto the glass slides favouring a correct immunofluorescence analysis. - Treat HeLa cells with 1 μM of camptothecin in RPMI 1640 medium supplemented with 10% FBS, 1 μg/ml penicillin and 1 µg/ml streptomycin and incubate at 37 °C in a 5% CO2 incubator for 2 h.
Note: The incubation time and drug concentration for the DNA damage agent can change in different cell types. As control, perform a DNA damage response analysis with different conditions evaluating DNA damage and resection through Western blot using an anti phospho-RPA32 S4/8 antibody (Bethyl Laboratories). - Detach HeLa cells from the dish with 1 ml of Trypsin-EDTA solution for 5 min at 37 °C in a standard 5% CO2 incubator.
- Inactivate the trypsin-EDTA solution with 3 ml of RPMI 1640 medium supplemented with 10% FBS, 1 μg/ml penicillin and 1 µg/ml streptomycin.
- Centrifuge HeLa cells for 5 min at 800 x g (rcf).
- Wash the cell pellet with 5 ml of ice-cold PBS followed by centrifugation for 5 min at 800 x g (rcf). Repeat Step B8 twice.
- Resuspend HeLa labelled cells with the ice-cold PBS at 2.5 x 105 cells/ml. Prepare another mix of unlabelled cells at the same concentration (2.5 x 105 cells/ml).
- Dilute labelled cells by mixing them in a 1:8 proportion with unlabelled cells.
- Mix 2.5 μl of cells with 7.5 μl of spreading buffer directly on the upper part of the slides.
- Incubate at room temperature (RT) for 10 min.
- Tilt the slide, from the ends, to 15°.
- DNA spreads are air-dried, approximately 10 min.
- Fix DNA fibres in 3:1 methanol/acetic acid at -20 °C for 15 min.
- Wash slides in PBS twice.
Note: The volume of PBS depends on the container used for the immersion of the slides - Incubate DNA fibres in 70% ethanol and 30% water overnight at 4 °C.
- Wash slides with PBS twice.
Note: This step is essential to remove all the ethanol residues before blocking DNA fibres. The volume of PBS depends on the container used for the immersion of the slides. - Block DNA fibres with PBS + 5% BSA for 30 min at RT.
Note: The volume of PBS depends on the container used for the immersion of the slides - Incubate slides with 1:300 Anti-BrdU clone BU1/75 (ICR1) for one hour at 37 °C.
- Wash slides with PBS twice.
Note: The volume of PBS depends on the container used for the immersion of the slides. - Incubate slides with 1:400 Alexa Fluor 488-conjugated chicken anti-rat.
- Wash slides with PBS twice.
Note: The volume of PBS depends on the container used for the immersion of the slides. - Mount the slide with ProLong Gold Antifade Reagent.
- Analyse the slide through a confocal microscope (we used the Zeiss LSM100) with a minimum of 63x oil immersion objective.
Data analysis
Analyse approximately 150 fibres for each condition and measure the length of fibres in pixel through the Fiji image software analysis.
To obtain the fibre length in μm:
- First of all download the LSM toolbox form the plugin window.
Note: This is an important step to open directly the confocal image in the .lsm format without jpg conversion. - Open the .lsm file, as reported in the Figure 3 below, there is the resolution of the picture and the conversion in μm.
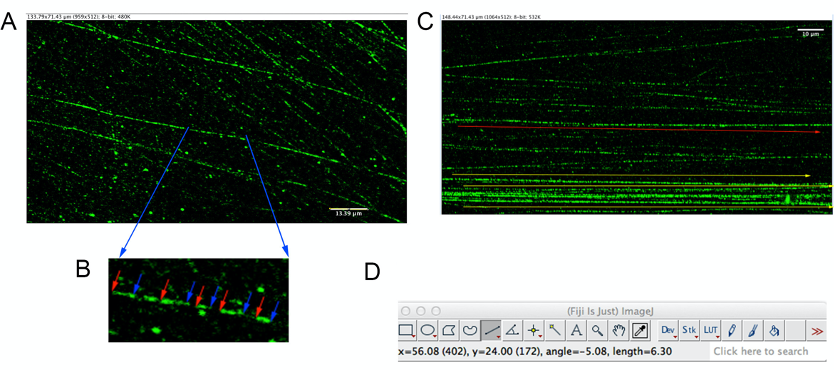
Figure 3. A representative image opened with the Fiji software showing in the upper left part the pixel resolution and μm conversion; in the lower corner of the figure the μm dimension. A. Micrometer conversion into kilobase depends on the DNA type, the nucleosome and chromatin fibre assembly; for HeLa cells and the combing assay 1 μm of DNA fibre length, during the S-phase, should approximately correspond to 2.59 kb as reported originally by Jackson and Pombo (Jackson and Pombo, 1998) and others (Bianco et al., 2012; Jamroskovic et al., 2020; Petermann et al., 2008). B. A magnification image of DNA fibre showing in red (start) and blue (end) the dimension of the analysed DNA tracts. C. The yellow arrows indicate, in lower part of the figure, the direction and the length of the DNA fibres; these fibres cannot be used for the measurement given the high number of other DNA fibres, which reduce the identification of single DNA traits. The red arrow indicates a fibre at a density and fluorescence, which are optimal for the analysis of the single DNA traits. D. Example of a screenshot of the Straight Line tool from the Fiji software analysis. - Select the Straight Line from the main window and measure the length of the fibres. The Straight Line tool of the Fiji software provides the length of the selected fibre identified through the x, y and angle coordinates; see an example of the Straight Line tool screen in Figure 3D.
- In this example, the mean fibre length ± standard deviation, calculated out of three independent experiments was plotted. Statistical significance was evaluated through the Student’s t-test using the GraphPad Prism 8 software in accordance with previous work (Cruz-García et al., 2014). The more informative way to represent data and the appropriate test for statistical analysis, however, will have to be determined according to the experimental plan upon checking data for normality through GraphPad Prism or similar programs.
Recipes
- Phosphate buffered saline (PBS)
137 mM NaCl
2.7 mM KCl
10 mM Na2HPO4
1.8 mM KH2PO4 - Spreading Buffer
0.5% Sodium dodecyl sulphate (SDS)
200 mM Tris-HCl pH 7.4
50 mM EDTA pH 8
Acknowledgments
We are grateful to the Sbarro Health Research Organization (http://www.shro.org), for its support.
We are grateful to Andres Cruz-García, AnaLópez-Saavedra and Pablo Huertas who first developed the SMART assay (Cruz-García et al., 2014), from which this protocol was adapted.
Competing interests
The authors declare no conflict of interest.
References
- Alfano, L., Costa, C., Caporaso, A., Altieri, A., Indovina, P., Macaluso, M., Giordano, A. and Pentimalli, F. (2016). NONO regulates the intra-S-phase checkpoint in response to UV radiation. Oncogene 35(5): 567-576.
- Alfano, L., Giordano, A. and Pentimalli, F. (2017). DNA Fiber Assay upon Treatment with Ultraviolet Radiations. Bio-protocol 7(11): e2301.
- Alfano, L., Caporaso, A., Altieri, A., Dell'Aquila, M., Landi, C., Bini, L., Pentimalli, F. and Giordano, A. (2019). Depletion of the RNA binding protein HNRNPD impairs homologous recombination by inhibiting DNA-end resection and inducing R-loop accumulation. Nucleic Acids Res 47(8): 4068-4085.
- Bianco, J. N., Poli, J., Saksouk, J., Bacal, J., Silva, M. J., Yoshida, K., Lin, Y. L., Tourriere, H., Lengronne, A. and Pasero, P. (2012). Analysis of DNA replication profiles in budding yeast and mammalian cells using DNA combing. Methods 57(2): 149-157.
- Chen, H., Lisby, M. and Symington, L. S. (2013). RPA coordinates DNA end resection and prevents formation of DNA hairpins. Mol Cell 50(4): 589-600.
- Cruz-García, A., López-Saavedra, A. and Huertas, P. (2014). BRCA1 accelerates CtIP-mediated DNA-end resection. Cell Rep 9(2): 451-459.
- Hanahan, D. and Weinberg, R. A. (2011). Hallmarks of cancer: the next generation. Cell 144(5): 646-674.
- Jackson, D. A. and Pombo, A. (1998). Replicon clusters are stable units of chromosome structure: evidence that nuclear organization contributes to the efficient activation and propagation of S phase in human cells. J Cell Biol 140(6): 1285-1295.
- Jamroskovic, J., Doimo, M., Chand, K., Obi, I., Kumar, R., Brannstrom, K., Hedenstrom, M., Nath Das, R., Akhunzianov, A., Deiana, M., Kasho, K., Sulis Sato, S., Pourbozorgi, P. L., Mason, J. E., Medini, P., Ohlund, D., Wanrooij, S., Chorell, E. and Sabouri, N. (2020). Quinazoline Ligands Induce Cancer Cell Death through Selective STAT3 Inhibition and G-Quadruplex Stabilization. J Am Chem Soc 142(6): 2876-2888.
- Krogh, B. O. and Symington, L. S. (2004). Recombination proteins in yeast. Annu Rev Genet 38: 233-271.
- Mao, Z., Bozzella, M., Seluanov, A. and Gorbunova, V. (2008). Comparison of nonhomologous end joining and homologous recombination in human cells. DNA Repair (Amst) 7(10): 1765-1771.
- Nimonkar, A. V., Genschel, J., Kinoshita, E., Polaczek, P., Campbell, J. L., Wyman, C., Modrich, P. and Kowalczykowski, S. C. (2011). BLM-DNA2-RPA-MRN and EXO1-BLM-RPA-MRN constitute two DNA end resection machineries for human DNA break repair. Genes Dev 25(4): 350-362.
- Petermann, E., Helleday, T. and Caldecott, K. W. (2008). Claspin promotes normal replication fork rates in human cells. Mol Biol Cell 19(6): 2373-2378.
- Stracker, T. H. and Petrini, J. H. (2011). The MRE11 complex: starting from the ends. Nat Rev Mol Cell Biol 12(2): 90-103.
- Tkáč, J., Xu, G., Adhikary, H., Young, J.T.F., Gallo, D., Escribano-Díaz, C., Krietsch, J., Orthwein, A., Munro, M., Sol, W., et al. (2016). HELB Is a Feedback Inhibitor of DNA End Resection. Mol Cell 61: 405-418.
- Zhou, Y., Caron, P., Legube, G. and Paull, T. T. (2014). Quantitation of DNA double-strand break resection intermediates in human cells. Nucleic Acids Res 42(3): e19.
Article Information
Copyright
© 2020 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Altieri, A., Dell'Aquila, M., Pentimalli, F., Giordano, A. and Luigi, A. (2020). SMART (Single Molecule Analysis of Resection Tracks) Technique for Assessing DNA end-Resection in Response to DNA Damage. Bio-protocol 10(15): e3701. DOI: 10.21769/BioProtoc.3701.
Category
Cancer Biology > Genome instability & mutation > Biochemical assays > DNA structure and alterations
Molecular Biology > DNA > DNA damage and repair
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link