- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Preparation of HeLa Total Membranes and Assay of Lipid-inhibition of Serine Palmitoyltransferase Activity
Published: Vol 10, Iss 12, Jun 20, 2020 DOI: 10.21769/BioProtoc.3656 Views: 4332
Reviewed by: Manjula MummadisettiTrinadh Venkata Satish TammanaPiyali Saha

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
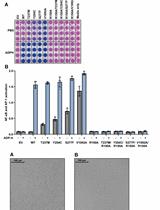

Measurement of the Activity of Wildtype and Disease-Causing ALPK1 Mutants in Transfected Cells With a 96-Well Format NF-κB/AP-1 Reporter Assay
Tom Snelling
Nov 20, 2024 1643 Views
Fluorescence Polarization-Based High-Throughput Screening Assay for Inhibitors Targeting Cathepsin L
Keyu Guo [...] Shuyi Si
Jul 20, 2025 2331 Views
Detecting the Activation of Endogenous Small GTPases via Fluorescent Signals Utilizing a Split mNeonGreen: Small GTPase ActIvitY ANalyzing (SAIYAN) System
Miharu Maeda and Kota Saito
Jan 5, 2026 543 Views
Abstract
Serine palmitoyltranferase (SPT) is a pyridoxal 5′ phosphate (PLP)-dependent enzyme that catalyzes the first and rate-limiting step of de novo synthesis of sphingolipids. SPT activity is homeostatically regulated in response to increased levels of sphingolipids. This homeostatic regulation of SPT is mediated through small ER membrane proteins termed the ORMDLs. Here we describe a procedure to assay ORMDL dependent lipid inhibition of SPT activity. The assay of SPT activity using radiolabeled L-serine was developed from the procedure established by the Hornemann laboratory. The activity of SPT can also be measured using deuterated L-serine but it requires mass spectrometry, which consumes money, time and instrumentation. The ORMDL dependent lipid inhibition of SPT activity can be studied in both cells and in a cell free system. This assay procedure is applicable to any type of mammalian cell. Here we provide the detailed protocol to measure SPT activity in the presence of either short chain (C8-ceramide) or long chain ceramide (C24-ceramide). One of the greatest advantages of this protocol is the ability to test insoluble long chain ceramides. We accomplished this by generating long chain ceramide through endogenous ceramide synthase by providing exogenous sphingosine and 24:1 acyl CoA in HeLa cell membranes. This SPT assay procedure is simple and easy to perform and does not require sophisticated instruments.
Keywords: Serine palmitoyltranferaseBackground
Serine palmitoyltranferase (SPT) is a multi-subunit enzyme that is widely expressed in eukaryotes and some prokaryotes (Hanada et al., 1997; Ikushiro et al., 2001; Hornemann et al., 2007). The first and rate-determining step of the sphingolipid biosynthetic pathway is catalyzed by SPT, producing 3-keto dihydrosphingosine from the condensation of serine and palmitoyl-CoA (Williams et al., 1984; Hanada, 2003). The mammalian SPT complex is composed of two large subunits and one small subunit. The large subunits are termed SPTLC1 and SPTLC2 and the small subunits are termed ssSPT (Nagiec et al., 1994; Lowther et al., 2012). SPT requires all three subunits to form a functional protein complex (Han et al., 2009). In some circumstances mammalian SPT is also composed of another large subunit termed SPTLC3 (Hornemann et al., 2009), which substitutes for SPTLC2. The functional SPT complex is formed by pairing SPTLC1 with either SPTCL2 or SPTLC3 and ssSPT. The active site of SPT resides in the SPTLC2 or SPTLC3 subunit. The substrate preference between the complex incorporating SPTLC2 and SPTLC3 differs. SPTLC2 prefers palmitoyl CoA. SPTLC3 prefers myristoyl-CoA (Hornemann et al., 2009). In certain situations SPT also incorporates alanine or glycine rather than serine into the precursor sphingolipid to generate a deoxysphingolipid base (Gable et al., 2010; Ferreira et al., 2018). SPT forms a stable complex with another ER resident protein termed ORMDL. This SPT-ORMDL interaction is crucial to maintain the homeostatic regulation of de novo biosynthesis of sphingolipids (Siow and Wattenberg, 2012; Davis et al., 2018). There are three closely related ORMDL family members found in mammals (Breslow et al., 2010; Siow and Wattenberg, 2012). We previously showed that increased levels of cellular ceramide inhibit SPT activity in an ORMDL-dependent manner in HeLa cells (Davis et al., 2019). The detailed mechanism behind the homeostatic regulation of SPT activity by the ORMDLs is largely unknown. SPT forms a stable complex with ORMDL, irrespective of cellular levels of ceramide. We propose that ceramides bind to either the ORMDLs or to the SPT-ORMDL complex to trigger the inhibition of SPT activity. We recently developed a cell free system in which membranes were isolated from HeLa cells and used to test the response of SPT activity to elevated sphingolipid. This system measures SPT activity in the presence or absence of ceramides with artificially short and natural chain lengths (Davis et al., 2019). We established the assay of ORMDL dependent lipid inhibition of SPT activity as a platform to test the functional interaction of SPT and ORMDL. This SPT assay procedure was developed from the procedure of Rutti et al. (2009). The cell free in vitro system has many advantageous over in vivo systems and both systems yield similar results (Davis et al., 2018). Using this cell free reconstitution system, we have determined that ORMDL dependent regulation of SPT activity is not mediated by post transcriptional or post translational mechanisms (Davis et al., 2018). This cell free system is a powerful tool to explore the detailed mechanism behind the homeostatic regulation of SPT by ORMDLs.
Materials and Reagents
- Vacuum filtration unit (0.2 μm) (VWR, catalog number: 10040-436 )
- T150 flask (Cyto One, catalog number: CC7682-4815 )
- 24-well plate (Cyto One, catalog number: CC7682-7524 )
- 26 gauge needle (BD, catalog number: 305110-269 )
- 1 ml syringe (BD, catalog number: 309659 )
- Thickwall Polycarbonate Tubes for Ultracentrifuge Rotors (Beckman Coulter, catalog number: 343776 )
- Screw cap tubes-2 ml (Axygen, catalog number: SCT-200-C-S )
- Scintillation tubes (PerkinElmer, catalog number: 6000292 )
- Disposable cell lifters-sterile (Fisher Scientific, catalog number: 08-100-240 )
- HeLa cells (ATCC, catalog number: CCL-2 )
- Dulbecco's minimal essential medium (DMEM Media) (Gibco, catalog number: 11960-044 )
- Fetal Bovine Serum (Gemini, catalog number: 900-108 )
- HEPES (Fisher Scientific, catalog number: BP310-1 )
- Penicillin-streptomycin, 10,000 U/ml (Gibco, catalog number: 15140-122 )
- Glutamine (Gibco, catalog number: 25030-081 )
- Trypsin-EDTA (Gibco, catalog number: 25200-056 )
- Collagen (Sigma, catalog number: C9791 )
- Sucrose (Fisher Scientific, catalog number: S5-3 )
- Myriocin (Cayman, catalog number: 63150 )
- Fumonisin B1 (Cayman, catalog number: 62580 )
- Tris (Fisher Scientific, catalog number: BP 152-5 )
- Complete mini EDTA-free Protease Inhibitor Cocktail (Roche, catalog number: 0 4693159001 )
- OptiMEM (Gibco, catalog number: 31985-070 )
- FA-Free BSA (Fisher, catalog number: BP9704-100 )
- Chloroform (Fisher, catalog number: C297-4 )
- Methanol (Fisher, catalog number: A452-4 )
- Liquid nitrogen
- C8 ceramide (Avanti, catalog number:860508)
- Sphingosine (Avanti, catalog number: 860490 )
- Palmitoyl CoA (Sigma, catalog number: P9716-10MG )
- 24:1 Coenzyme A (Avanti, catalog number: 870725 )
- Potassium hydroxide (Fisher, catalog number: P250-500 )
- DTT (Sigma, catalog number: D0632 )
- EDTA (Fisher, catalog number: S311-500 )
- Pyridoxal 5’ phosphate (Sigma, catalog number: P9255 )
- L-Serine (Sigma, catalog number:S4500)
- L-[3H(G)]-Serine (PerkinElmer, catalog number: 2477301 )
- Ecolite-Scintillation fluid (MP Biomedicals, catalog number: 882475 )
- Beta max-Scintillation fluid (MP Biomedicals, catalog number: 880020 )
- Serine free MEM (Gibco, catalog number: 11095-080 )
- Magnesium chloride (Fisher Scientific, catalog number: M87-100 )
- DMEM complete medium (see Recipes)
- C8-ceramide (see Recipes)
- Sphingosine (see Recipes)
- Palmitoyl CoA (see Recipes)
- 24:1 CoenzymeA (see Recipes)
- Fatty acid free BSA (see Recipes)
- 1 mM C8-ceramide (see Recipes)
- Methanol (see Recipes)
- Myriocin (see Recipes)
- Fumonisin B1 (see Recipes)
- Chloroform:methanol (2:1) (see Recipes)
- 10x Phosphate buffered saline (PBS) pH 7.4 (see Recipes)
- Alkaline methanol (see Recipes)
- Alkaline water (see Recipes)
- Digitonin (see Recipes)
- Pyridoxal 5’ phosphate (see Recipes)
- L-serine stock (see Recipes)
- 0.05% Trypsin (see Recipes)
- Scintillation fluid (see Recipes)
- Collagen (see Recipes)
- Collagen coating (see Recipes)
Equipment
- Cell culture Hood (LABCONCO, catalog number: 36213043726 )
- -80 °C freezer (Thermo Scientific, model: 5815 )
- CO2 incubator (Thermo Scientific, model: 3110 )
- KONTES Dounce glass homogenizer (VWR, catalog number: KT885300-0007 )
- Table top refrigerated centrifuge with swingout rotor (Eppendorf, model: 5810 )
- Table top Ultracentrifuge (Beckman Coulter, model: Optima MAX-XP )
- TLA 120.2 rotor ( Backman Coulter, S/N: 16U1971)
- Scintillation counter (Backman Coulter, model: LS6500 )
- Water bath (Thermo Scientific, model: 2354 )
- Nitrogen gas (Airgas, catalog number: NI UHP300 )
- CO2 gas (Airgas, catalog number: CD USP50 )
- Vortexer (Vortex-Genie 2, model: SI-0236 )
- Table top high speed centrifuge (Thermo Scientific, model: accuSpin Micro 17 )
Procedure
- Extraction of L-[3H(G)]-serine
- Handle and store L-[3H(G)]-Serine only in designated and authorized locations with proper precautions.
- Dispose radioactive waste in specially designated waste containers.
- Take 2 µl of serine before and after extraction and measure radioactivity to calculate the recovery of L-[3H(G)]-Serine.
- Dispense 0.2 ml of the [3H]-serine into a 2 ml screw cap tube, then add 0.5 ml of chloroform:methanol (2:1 ratio) to the tube.
- Vortex the tube for 1 min.
- Centrifuge the tube at 16,200 x g for 2 min at room temperature in a high speed centrifuge.
- Transfer the upper aqueous phase into fresh 2 ml screw cap tubes.
- Place the tubes into the house vacuum flask for 30 min to remove the traces of organic solvents.
- Measure the radioactivity after extraction to calculate specific activity of L-[3H(G)]-serine.
- Use this extracted [3H]-serine to measure SPT activity in cells and membranes.
- Extracted [3H]-serine will give less background, i.e., low counts with myriocin treatment in SPT assay.
- Assay of lipid inhibition of serine palmitoyltransferase (SPT) activity in intact HeLa cells
- Grow HeLa cells in DMEM complete medium and maintain cells in a CO2 incubator at 37 °C and 5% CO2.
- On day-1, plate HeLa cells (7 x 104 cells/well) in a collagen coated 24-well plate with 1 ml of DMEM complete medium.
- See Recipes for the procedure of collagen coating in 24-well plates.
- Divide cells into 4 groups for control, C8-ceramide, myriocin treatment and protein analysis.
- Prepare 4 wells for each group and use average value for calculations.
- Myriocin is a specific inhibitor for SPT and is a negative control for this assay.
- On day-2, add 10 µM C8-ceramide or 1 µM myriocin to the cells and incubate for 1 h in a CO2 incubator, see recipe for C8-ceramide and myriocin preparation.
- Incubate control cells with methanol-BSA complex.
- After 1 h remove C8-ceramide or myriocin and gently wash cells with 0.25 ml of PBS at room temperature.
- Measure SPT activity by adding 0.25 ml of extracted 3H-serine to the cells in serine free MEM media (5 µCi/ml of serine free media) and incubate cells in a CO2 incubator for 1 h at 37 °C.
- After 1 h, remove radioactive serine and gently wash cells with 0.25 ml of PBS at room temperature.
- Stop the reaction by adding 400 µl of alkaline methanol and scrape the cells using plastic cell lifters.
- Transfer cells into 2 ml screw cap tubes.
- Extract total sphingolipids under alkaline conditions as described below in Section H.
- Refer to Figure 1A in Davis et al. (2019) and Figure 2A in Siow et al. (2015) for results.
- Assay of SPT activity in Permeabilized HeLa cells
- Plate HeLa cells (7 x 104 cells/well) in a collagen coated 24-well plate and incubate in a CO2 incubator for 24 h.
- Divide cells into 4 groups for control, C8-ceramide, myriocin treatment and protein analysis.
- Prepare 4 wells for each group and use average value for calculations.
- After 24 h, permeablize HeLa cells with 0.02% digitonin prepared in Opti-MEM.
- Add 250 µl of 0.02% digitonin and incubate cells for 3 min at 37 °C.
- After 3 min, remove digitonin and gently wash cells with 0.25 ml of PBS at room temperature.
- Prepare preincubation medium, 50 mM HEPES pH8.0, 1 mM EDTA and 20 µM pyridoxyl 5′-phosphate.
- Dilute C8-ceramide or myriocin with preincubation media and pre-warmed to 37 °C.
- Add 200 µl of preincubation medium with or without 10 µM C8-ceramide or 1 µM myriocin to permeabilized cells.
- Incubate cells at 37 °C for 30 min in a CO2 incubator.
- After 30 min add 200 µl of labelling media to each well, contains 2 μCi of extracted [3H]-serine, 1 mM L-serine, and 50 μM palmitoyl-CoA prepared in preincubation medium.
- Incubate cells for 60 min in a CO2 incubator.
- After 60 min remove the labelling media.
- Stop the reaction by adding 400 µl of alkaline methanol and scrape the cells using plastic cell lifters.
- Transfer cells into 2 ml screw cap tubes for lipid extraction.
- Extract total sphingolipids under alkaline condition as described below in Section H.
- Refer to Figure 1A in Davis et al. (2019) and Figure 2A-B in Siow and Wattenberg (2012) for results.
- Preparation of HeLa cell lysate
- Seed 3 x 106 HeLa cells into one T-150 flask with DMEM complete medium.
- Grow HeLa cells for 48 h in a CO2 incubator to achieve 90-95% confluency.
- Remove the DMEM complete media and wash cells with 4 ml of PBS at room temperature.
- Add 4 ml of 0.05% trypsin to cells and incubate for 2 min at room temperature.
- After 2 min, remove trypsin and incubate cells at 37 °C for 5 min.
- Harvest cells with 10 ml of ice cold DMEM complete media.
- Collect cells by centrifugation for 10 min at 72 x g.
- Wash cells with 10 ml of ice cold PBS.
- Resuspend cells in 1.6 ml of swelling buffer (10 mM Tris pH 7.5, 15 mM KCl and 1 mM MgCl2) and incubate on ice for 15 min.
- To this, add 534 µl of 1 M sucrose, 7 µl of 200 mM EDTA and 80 µl of protease inhibitors cocktail from 25x stock.
Note: 25x stock made by dissolving 1 protease inhibitor tablet in 2 ml of water. - Homogenize the cells on ice with a 7 ml Dounce homogenizer using pestle-B with 30 to 40 strokes.
- Centrifuge the homogenate to remove unbroken cells and nuclei at 72 x g for 10 min at 4 °C with a swinging bucket rotor and collect the supernatant.
- Pass the cell free lysate through a 26 gauge needle with 1 ml syringe for 10 times.
- Aliquot 100 µl lysate into 1.5 ml tubes, snap freeze with liquid nitrogen and store at -80 °C until further analysis.
- Preparation of total membrane
- Prepare cell free lysate as described above in Section D.
- Transfer 1 ml of cell free lysate to each clean thick wall ultra-centrifuge tubes then centrifuge at 434,513 x g for 20 min at 4 °C in a TLA 120.2 rotor.
- Discard the supernatant.
- Resuspend the membrane pellet with 500 µl of membrane resuspension buffer (250 mM Sucrose, 25 mM Tris, pH 7.4 and 20 µl of protease inhibitors cocktail from 25x stock) and homogenize with the Dounce homogenizer using pestle-B with 10 strokes.
- Pass the membrane through a 26 gauge needle with 1 ml of syringe for 10 times.
- Aliquot 100 µl of membranes into 1.5 ml tubes and snap freeze with liquid nitrogen and store at -80 °C until further analysis.
- Assay of lipid inhibition of serine palmitoyltransferase activity in cell free lysate or total membranes
- Perform in vitro SPT activity in 2 ml screw cap tubes and total reaction volume of 200 µl.
- For assay of SPT activity with short chain ceramide, incubate cell free lysate (100 µg total protein) or membrane (50 µg total protein) in a pre-incubation buffer with or without 10 µM C8-ceramide or 1 µM myriocin on ice for 40 min.
- Pre-incubation buffer contains, 50 mM HEPES pH 8.0, 25 mM DTT, 2 mM EDTA and 20 µM PLP.
- After 40 min, add 100 µl of labelling mix then mix well.
- Labelling mix contains, 2 mM Serine, 100 µM palmitoyl CoA, 2 µCi extracted [3H]-serine.
- Incubate tubes for 60 min at 37 °C.
- Stop reaction by adding 400 µl of alkaline methanol.
- Extract total sphingolipid under alkaline conditions as described below in Section H.
- Refer to Figures 1A, B, 2A and 3A in Davis et al. (2019) for results.
- Assay of SPT activity with long chain ceramide
- For generation of long chain ceramide in membranes, incubate 50 µg of membrane with 20 μM sphingosine and 50 μM 24:1 CoA or 1 µM myriocin or 50 µM Fumonisin-B1 in a final volume of 100 µl in a buffer containing 20 mM HEPES pH 7.4, 25 mM KCl, 2 mM MgCl2 and 0.1% fatty acid free BSA.
- Fumonisin-B1 is an inhibitor of ceramide synthases.
- Incubate this reaction mix at 37 °C for 60 min to generate ceramide with 24:1 acyl chain by endogenous ceramide synthases.
- After 60 min, add 100 µl of labelling mix and incubate the tubes for additional 60 min at 37 °C.
- Labelling mix contains 50 mM HEPES pH 8.0, 1 mM DTT, 10 mM EDTA, 20 µM PLP, 2 mM serine,100 µM palmitoyl CoA and 2 µCi extracted [3H] serine.
- Stop the reaction by adding 400 µl of alkaline methanol and extract lipids under alkaline conditions as described below in Section H.
- Refer to Figure 2B-C in Davis et al. (2019) for results.
- Extraction of total sphingolipids under alkaline condition
- Add 100 µl of chloroform to the tube contains cells or membrane or lysate with 400 µl alkaline methanol.
- Vortex the tubes and centrifuge for 1 min at 16,200 x g in a high speed centrifuge.
- Add 500 µl of chloroform followed by 300 µl alkaline water and 100 µl of 2 N NH4OH to separate aqueous and organic phases.
- Vortex the tube for 1 min and spin at 16,200 x g for 1 min in a high speed centrifuge.
- Aspirate the upper aqueous phase.
- Add 1 ml of alkaline water to the lower organic phase, vortex well then centrifuge for 1 min at 16,200 x g.
- Aspirate the upper aqueous phase and repeat the above step 6 again.
- Transfer 350 µl of the lower organic phase into scintillation vials and dry under nitrogen gas.
- Add 5 ml of scintillation fluid, vortex well and count radioactivity using a scintillation counter.
Data analysis
- Expression of in vitro and in vitro SPT activity as [3H]-serine in total sphingolipids (CPM/mg/min).
- Here we provide the results obtained from previously performed experiments and detailed steps to calculate in vivo and in vitro SPT activity with results obtained from previously performed experiments.
- Table 1 shows results of in vitro SPT activity from HeLa cell membranes.
Table 1. Serine palmitoyltransferase activity in total membranes of HeLa cells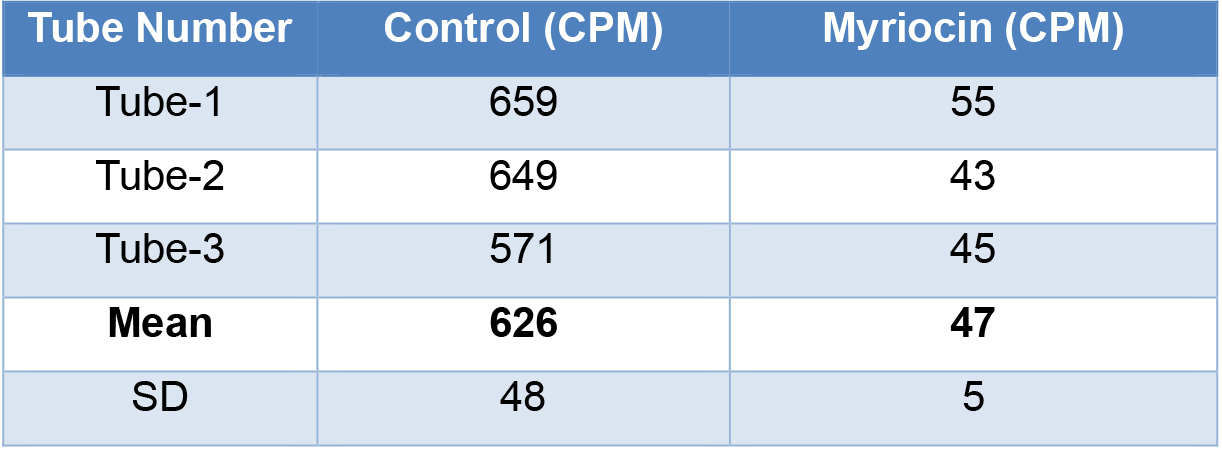
- Normalize the CPM values to mg of total protein (50 µg of protein used for SPT assay)
Control: 626.5/0.05 × 1 = 12,530 CPM/mg
Myriocin: 47.6/0.05 × 1 = 952 CPM/mg - Convert values to per min (SPT assay conducted for 60 min)
Control: 12530/60 = 208.8 CPM/mg/min
Myriocin: 952/60 = 15.8 CPM/mg/min - In vivo and in vitro SPT activity is expressed as [3H] serine in total sphingolipids (CPM/mg/min).
- Expression of in vitro SPT activity as [3H]-serine in total sphingolipids (pMol/mg/min)
- Total radioactivity in substrate (labelling mix) is 1,280,980 CPM.
- SPT reaction mix contains 200 nMol of serine.
- Consider 200 nMol of serine is equivalent to 1,280,980 CPM.
- Calculate specific activity from this value by using this formula,
nMol of serine/CPM in substrate × mean CPM from SPT assay = X nMol,
e.g., 200/1,280,980 × 626 = 0.097 nMol of [3H] serine in total sphingolipids. - Mean value of control SPT activity is 626 CPM.
- Convert the above value to per mg protein, 0.097/0.05 ×1 = 1.94 nMol/mg protein.
- Convert the above value to per min, 1.94/60 = 0.032 nMol/mg/min.
- Convert value from nmol to pmol by multiplying the above value by 1,000 (0.032 × 1,000 = 32 pMol/mg/min).
- In vitro SPT activity is expressed as [3H]-serine in total sphingolipids (pMol/mg/min).
Recipes
- DMEM complete medium
500 ml of DMEM media
50 ml of FBS
10 ml of HEPES pH 8.0
5 ml of glutamine and 5 ml Penicillin-streptomycin - C8-ceramide
10 mg of C8-ceramide dissolved in 1.174 ml of methanol as a 20 mM stock and store at -20 °C - Sphingosine
10 mg of sphingosine dissolved in 3.34 ml of methanol as a 10 mM stock store at -20 °C - Palmitoyl CoA
10 mg of palmitoyl CoA dissolved in 1.99 ml of DMSO as a 5 mM stock and store at -20 °C - 24:1 CoenzymeA
Dissolve 5 mg of 24:1 Coenzyme A in 857.2 µl of DMSO to make 5 mM Stock and store at -20 °C - Fatty acid free BSA
Fatty acid free BSA prepared in PBS as a 2% stock and filter sterilize using 0.2 µm syringe filter and store at 4 °C - 1 mM C8-ceramide
BSA complex 2.5 µl of 20 mM C8-ceramide mixed with 47.5 µl of 2% fatty acid free BSA, made every time fresh - Methanol:BSA complex
2.5 µl of methanol mixed with 47.5 µl of 2% fatty acid free BSA, made every time fresh - Myriocin
Dissolve myriocin in methanol as a 1 mM stock and store at -20 °C - Fumonisin B1
1 mg of Fumonisin B1 dissolved in 92.4 µl of DMSO as a 15 mM stock and store at -20 °C - Chloroform:methanol (2:1)
Mix 2 volume of chloroform and 1 ml volume of methanol - 10x Phosphate buffered saline (PBS) pH 7.4
15.4 mM KH2PO4, 1.55 M NaCl and 28 mM Na2HPO4·7H2O in water, if required adjust pH to 7.4 with HCl - Alkaline methanol
Dissolve 0.7 g potassium hydroxide in 100 ml of methanol - Alkaline water
Add 100 µl of 2 N NH4OH in 100 ml water - Digitonin
Prepare 1% digitonin in DMSO as a stock and dilute to 0.02% in Opti-Mem - Pyridoxal 5’ phosphate
Prepared in water as a 20 mM stock and store at -20 °C - L-serine stock
Prepare serine in water as a 50 mM stock and store at -20 °C - 0.05% Trypsin
Dilute stock 0.25% trypsin into 0.05% with Phosphate buffered saline 7.4 and filter sterilize and store at 4 °C - Scintillation fluid
We use Ecolite scintillation fluid to measure radioactivity in aqueous solutions and Beta max to measure radioactivity in organic solvents - Collagen
Dissolve 10 mg of collagen in 10 ml of 0.1 N acetic acid and mix it for 3 h at room temperature, then dilute collagen to 50 µg/ ml in PBS and store at 4 °C - Collagen coating
Equilibrate collagen to room temperature and add 1 ml to each well of 24-well plate, incubate at room temperature for 5 min. After incubation completely remove all collagen by pipetting. Collagen is reusable at least for 10 times.
Acknowledgments
This assay protocol is developed from the method of Rutti et al. (2009). This work was supported by funding from National Institute of Health grant RO1HL131340 to B.W.
Competing interests
Authors have no conflicts of interest to disclose
References
- Breslow, D. K., Collins, S. R., Bodenmiller, B., Aebersold, R., Simons, K., Shevchenko, A., Ejsing, C. S. and Weissman, J. S. (2010). Orm family proteins mediate sphingolipid homeostasis. Nature 463(7284): 1048-1053.
- Davis, D., Kannan, M. and Wattenberg, B. (2018). Orm/ORMDL proteins: Gate guardians and master regulators. Adv Biol Regul 70: 3-18.
- Davis, D. L., Gable, K., Suemitsu, J., Dunn, T. M. and Wattenberg, B. W. (2019). The ORMDL/Orm-serine palmitoyltransferase (SPT) complex is directly regulated by ceramide: Reconstitution of SPT regulation in isolated membranes. J Biol Chem 294(13): 5146-5156.
- Ferreira, C. R., Goorden, S. M. I., Soldatos, A., Byers, H. M., Ghauharali-van der Vlugt, J. M. M., Beers-Stet, F. S., Groden, C., van Karnebeek, C. D., Gahl, W. A., Vaz, F. M., Jiang, X. and Vernon, H. J. (2018). Deoxysphingolipid precursors indicate abnormal sphingolipid metabolism in individuals with primary and secondary disturbances of serine availability. Mol Genet Metab 124(3): 204-209.
- Gable, K., Gupta, S. D., Han, G., Niranjanakumari, S., Harmon, J. M. and Dunn, T. M. (2010). A disease-causing mutation in the active site of serine palmitoyltransferase causes catalytic promiscuity. J Biol Chem 285(30): 22846-22852.
- Han, G., Gupta, S. D., Gable, K., Niranjanakumari, S., Moitra, P., Eichler, F., Brown, R. H., Jr., Harmon, J. M. and Dunn, T. M. (2009). Identification of small subunits of mammalian serine palmitoyltransferase that confer distinct acyl-CoA substrate specificities. Proc Natl Acad Sci U S A 106(20): 8186-8191.
- Hanada, K. (2003). Serine palmitoyltransferase, a key enzyme of sphingolipid metabolism. Biochim Biophys Acta 1632(1-3): 16-30.
- Hanada, K., Hara, T., Nishijima, M., Kuge, O., Dickson, R. C. and Nagiec, M. M. (1997). A mammalian homolog of the yeast LCB1 encodes a component of serine palmitoyltransferase, the enzyme catalyzing the first step in sphingolipid synthesis. J Biol Chem 272(51): 32108-32114.
- Hornemann, T., Penno, A., Rutti, M. F., Ernst, D., Kivrak-Pfiffner, F., Rohrer, L. and von Eckardstein, A. (2009). The SPTLC3 subunit of serine palmitoyltransferase generates short chain sphingoid bases. J Biol Chem 284(39): 26322-26330.
- Hornemann, T., Wei, Y. and von Eckardstein, A. (2007). Is the mammalian serine palmitoyltransferase a high-molecular-mass complex? Biochem J 405(1): 157-164.
- Ikushiro, H., Hayashi, H. and Kagamiyama, H. (2001). A water-soluble homodimeric serine palmitoyltransferase from Sphingomonas paucimobilis EY2395T strain. Purification, characterization, cloning, and overproduction. J Biol Chem 276(21): 18249-18256.
- Lowther, J., Naismith, J. H., Dunn, T. M. and Campopiano, D. J. (2012). Structural, mechanistic and regulatory studies of serine palmitoyltransferase. Biochem Soc Trans 40(3): 547-554.
- Nagiec, M. M., Baltisberger, J. A., Wells, G. B., Lester, R. L. and Dickson, R. C. (1994). The LCB2 gene of Saccharomyces and the related LCB1 gene encode subunits of serine palmitoyltransferase, the initial enzyme in sphingolipid synthesis. Proc Natl Acad Sci U S A 91(17): 7899-7902.
- Rutti, M. F., Richard, S., Penno, A., von Eckardstein, A. and Hornemann, T. (2009). An improved method to determine serine palmitoyltransferase activity. J Lipid Res 50(6): 1237-1244.
- Siow, D. L. and Wattenberg, B. W. (2012). Mammalian ORMDL proteins mediate the feedback response in ceramide biosynthesis. J Biol Chem 287(48): 40198-40204.
- Williams, R. D., Wang, E. and Merrill, A. H., Jr. (1984). Enzymology of long-chain base synthesis by liver: characterization of serine palmitoyltransferase in rat liver microsomes. Arch Biochem Biophys 228(1): 282-291.
- Siow, D., Sunkara, M., Dunn T.M., Morris, A.J., Wattenberg, B. (2015). ORMDL/serine palmitoyltransferase stoichiometry determines effects of ORMDL3 expression on sphingolipid biosynthesis. J Lipid Res. 56 (4): 898-908.
Article Information
Copyright
© 2020 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Kannan, M., Davis, D. L., Suemitsu, J., Oltorik, C. D. and Wattenberg, B. (2020). Preparation of HeLa Total Membranes and Assay of Lipid-inhibition of Serine Palmitoyltransferase Activity. Bio-protocol 10(12): e3656. DOI: 10.21769/BioProtoc.3656.
- Davis, D. L., Gable, K., Suemitsu, J., Dunn, T. M. and Wattenberg, B. W. (2019). The ORMDL/Orm-serine palmitoyltransferase (SPT) complex is directly regulated by ceramide: Reconstitution of SPT regulation in isolated membranes. J Biol Chem 294(13): 5146-5156.
Category
Biochemistry > Protein > Synthesis
Biochemistry > Protein > Activity
Cell Biology > Cell-based analysis > Enzymatic assay
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link











