- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Precise Targeting of Single Microelectrodes to Orientation Pinwheel Centers
(*contributed equally to this work) Published: Vol 10, Iss 11, Jun 5, 2020 DOI: 10.21769/BioProtoc.3643 Views: 3965
Reviewed by: Miao HeEhsan KheradpezhouhAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Operant Self-medication for Assessment of Spontaneous Pain Relief and Drug Abuse Liability in Mouse Models of Chronic Pain
David Cabañero [...] Rafael Maldonado
Mar 5, 2022 3065 Views
Developing a Ministroke Model in Mouse Barrel Cortex
Song Wang [...] Yihan Wu
Mar 20, 2025 2366 Views
Mouse Vestibulo-Ocular Reflex Testing for Otolith Organs and Horizontal Semicircular Canal
Tong Zhao [...] Fangyi Chen
Nov 20, 2025 1243 Views
Abstract
In the mammalian visual system, early stages of visual form perception begin with orientation selective neurons in primary visual cortex (V1). In many species (including humans, monkeys, tree shrews, cats, and ferrets), these neurons are organized in pinwheel-like orientation columns. To study the functional organization within orientation pinwheels, it is important to target pinwheel subdomains precisely. We therefore developed a technique to provide a quantitative determination of the location of pinwheel centers (PCs). Previous studies relied solely on blood vessel images of the cortical surface to guide electrode penetrations to PCs in orientation maps. However, considerable spatial error remained using this method. In the present study, we improved the accuracy of targeting PCs by ensuring perpendicularity of electrodes and by utilizing the orientation tuning of local field potentials (LFP) recorded at or near the optically determined positions.
Keywords: Orientation domainBackground
Within the visual cortex, orientation pinwheels (PCs) are singularity centers around which neuronal orientation preference systematically shifts. PCs have been the focus of many studies on how the brain encodes different aspects of object shape, such as linear oriented segments, curved segments, corners, and T junctions (Hubel and Wiesel, 1974; Das and Gilbert, 1999; Hashemi-Nezhad and Lyon, 2012). Single-unit recording has been the most common method for studying organization within orientation pinwheels (Hubel and Wiesel, 1974). However, due to limitations of blood vessel guided localization of microelectrodes and uncertainties of electrode perpendicularity (Nauhaus and Ringach, 2007), it is quite challenging to accurately target microelectrodes to pinwheel centers (PCs). Hashemi-Nezhad and Lyon (2012) used electrodes coated with Dil to nicely demonstrate that electrodes were truly perpendicular; however, the vasculature-based method still contained localization error. Nauhaus and Ringach (2007), carefully matched the electrode locations in a Utah array with locations in an optical imaged orientation map and achieved accurate PC localization, but such an approach offers little control over where the electrodes end up within orientation domains or pinwheel centers. Using 2-photon methods, Ohki et al. (2006) reported that the true PC is only 130 µm in diameter, underscoring the need for highly accurate electrode targeting (see also Nauhaus et al., 2008).
In this study, we developed a method for targeting PCs with high accuracy. This was achieved by: (1) Assuring perpendicular penetrations by imaging a large field of view with a narrow depth of field (~50 μm, front-to-front lens with f1.2 (Ratzlaff and Grinvald, 1991), and then ensuring the electrode paralleled exactly the axis of the optical imaging. Using this procedure, we have estimated that the deviation of electrodes from perpendicular is less than 2 deg (see Figure 3D). (2) Carefully targeting electrodes to PCs based on blood vessel guidance. (3) Further determining the precise PC center by assessing whether the local field potentials (LFP) exhibit ‘non-sine-like’ responses. We have previously shown (Li et al., 2019) using this methodology that penetrations with sine-like LFP response and small scatter in orientation preference are not true PC locations, while those with non-sine-like response and large scatter in orientation preference (> 60 deg, Maldonado et al., 1997) are the true PC locations. This additional non-sine-like criterion is essential for highly accurate determination of PC locations and has provided new understanding of orientation subdomain organization (Li et al., 2019). These detailed procedures are described below.
Materials and Reagents
- Cover glass
- Tungsten-in-glass microelectrodes (impedance: ~5 MΩ; tip diameter: ~1 μm):
These electrodes were homemade. Detailed description of procedures for electrode construction can be found in previous paper (Li et al., 1995). Briefly, tungsten-in-glass electrodes were prepared from tungsten wire (diameter 200 µm, Shanghai Tungsten and molybdenum products company, type: S-EST6135) and capillary glass (inner diameter: 0.3 mm, outer diameter: 1 mm, borosilicate glass, Shanghai Glass Works, type: SGW-95). The desired impedance of ~5 MΩ was measured (Ωmega-Tip-Z, World Precision Instruments,USA ) and corresponded to tungsten tip length of ~10 µm and the tip diameter of ~1 µm (Figure 1).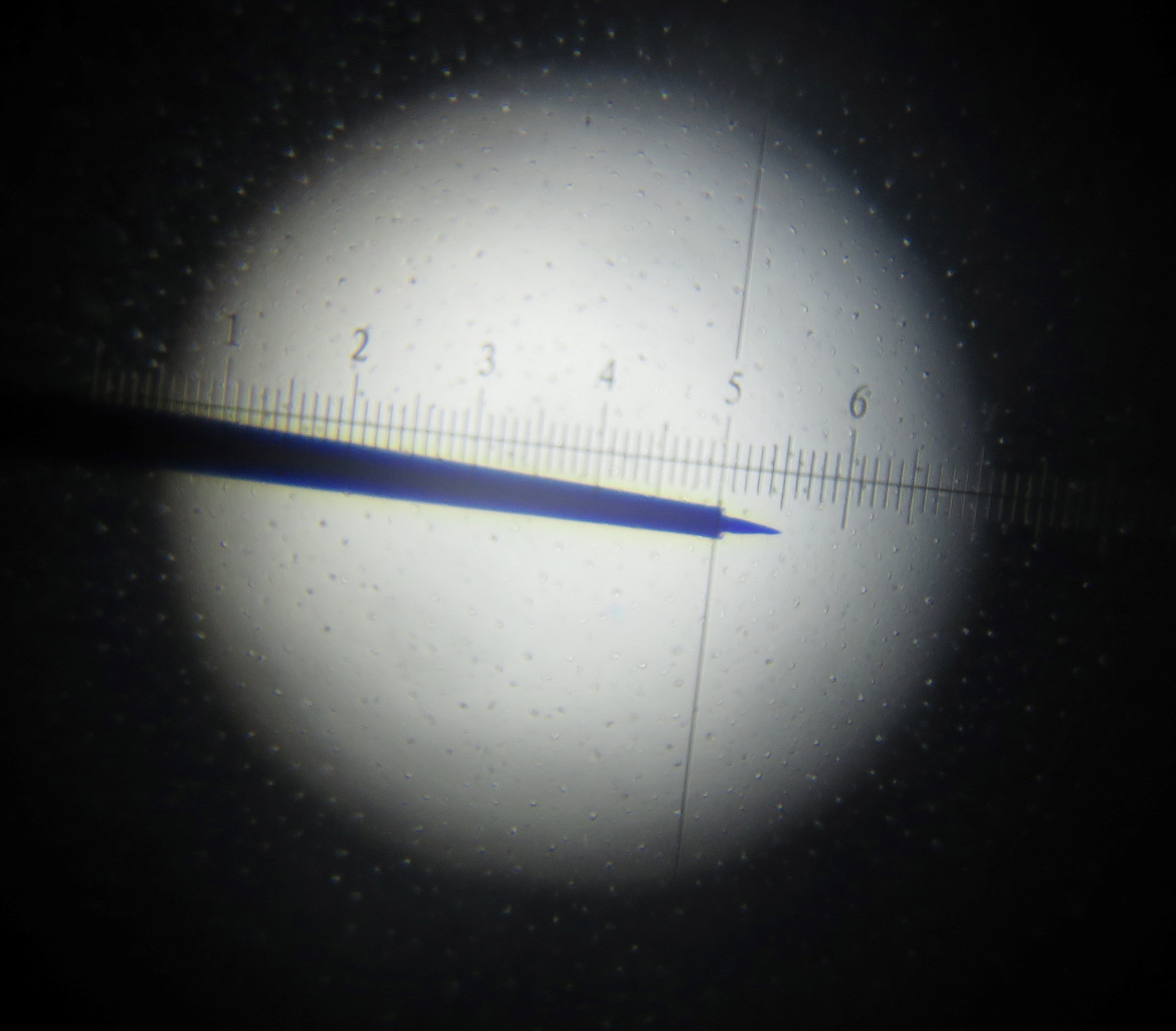
Figure 1. Photograph of tungsten-in-glass microelectrode. 1 grid mark is 2.5 µm Image shows electrode tip length of of ~10 µm, and tip diameter of ~1 µm. - Animals
These procedures were developed using 16 anesthetized adult cats of both sexes. The following procedures can be applied to the visual cortex of other species as well. - Ketamine hydrochloride
- 1% atropine sulfate eyedrops
- Agar
- Barbiturate
Equipment
- Neural data acquisition system (Blackrock Microsystems Inc., Cerebus, Cerebus 32)
- Video camera (Andor Technology, Andor, model: iXon DU897 )
- Lenses (Nikon, NIKKOR, D = 50 mm, f = 1.2)
- Visual stimulus Generator (Cambridge Research Systems Ltd. ViSaGe MKII)
- Stereotaxic apparatus (Narishige, SN-3N)
Software
- Matlab (Mathworks, ww2.mathworks.cn), and the matlab code as the supplemental file
Procedure
- Animal Preparation Procedures
Detailed descriptions of procedures for animal surgery and anesthesia can be found in previous studies (Song and Li, 2008).- Anesthetize cats before surgery with ketamine hydrochloride (30 mg/kg), and then perform endotracheal intubation and venous cannulations. After surgery, the animal was placed in a stereotaxic frame for performing a craniotomy and conducting neurophysiological procedures. During recording, maintain anesthesia (urethane 20 mg/kg/h), paralysis (gallamine triethiodide 10 mg/kg/h), and blood sugar and fluids (glucose 200 mg/kg/h in Ringer’s solution 3 ml/kg/h i.v.). Monitor continuously heart rate, electrocardiography, electroencephalography (EEG), end-expiratory CO2, and rectal temperature. Anesthesia is considered to be sufficient when the EEG indicates a stable sleep-like state marked by sleep spindles. Reflexes, including cornea, eyelid, and withdrawal reflexes are tested at regular intervals.
- The eyes are prepared for vision experiments: Retract nictitating membranes and dilate the pupils using 1% atropine sulfate eyedrops. Apply contact lenses and use additional corrective lenses to achieve focus of the retina on a computer monitor in front of the animal.
- Perform a craniotomy and durotomy above area 17 (V1) and cement a stainless steel chamber on the surrounding skull. To dampen cortical pulsations, either seal the chamber with a cover glass and fill with silicone oil, or cover cortex with 4% agar and a cover glass.
- At the end of the experiment, sacrifice the animal by an overdose of barbiturate administered intravenously (dosage, 5 ml, 6% barbiturate).
- Optical Imaging Procedures
- Position camera and lens over the optical window. Crude positioning of the camera was enabled using a homemade camera arm to which a micromanipulator with 100 millimeter of freedom (manually adjust x, y, z position). Fine positioning of the camera was achieved by electrically adjusting the x, y, z position (10 millimeter freedom) and the declination angle α (45 degree freedom) with a micromanipulator platform. The optical instrument is a 14-bit video camera (Andor Technology, Northern Ireland) equipped with two front-to-front connected 50 mm Nikon lenses.
- Illluminate the cortical surface as evenly as possible using green light (546 nm) illumination. Obtain a reference image of the blood vessel pattern (see Figure 2A).
- Focus the camera ~400 μm below the surface of the cortex and collect optical images using red light illumination (605 nm, a wavelength optimal for hemodynamic oximetry signals).
- Obtain orientation preference maps. The intrinsic signals were recorded in response to binocularly viewed full-screen, high-contrast (100%) sinewave gratings. The main set of stimuli included drifting gratings, presented at 8 equally spaced orientations, in both directions (16 conditions). Each stimulus was presented 20 times for 9 s followed by an inter stimulus interval of 16 s. The visual stimuli were generated by a Cambridge Systems VSG graphics board. Stimuli were presented on a high-resolution monitor screen (40 × 30 cm) at a 100 Hz vertical refresh rate. The screen background was maintained at the identical mean luminance as the stimulus patches (10 cd/m2). The monitor was placed 57cm from the animal’s eyes.
- Generate colored-coded orientation preference maps (Figure 2B) via pixel-by-pixel vector summation of the 8 orientations (Bonhoeffer and Grinvald, 1991). In some experiments, drifting gratings were presented at 16 equally spaced orientations (at 11.25° intervals).
- Carefully choose only highly accurate and reproducible maps. That is, first generate maps from different subsets of trials. Then select only maps in which there is no displacement (shifted by no more than 13-26 μm) of pinwheel centers. These are deemed acceptable (Mariño et al., 2005).
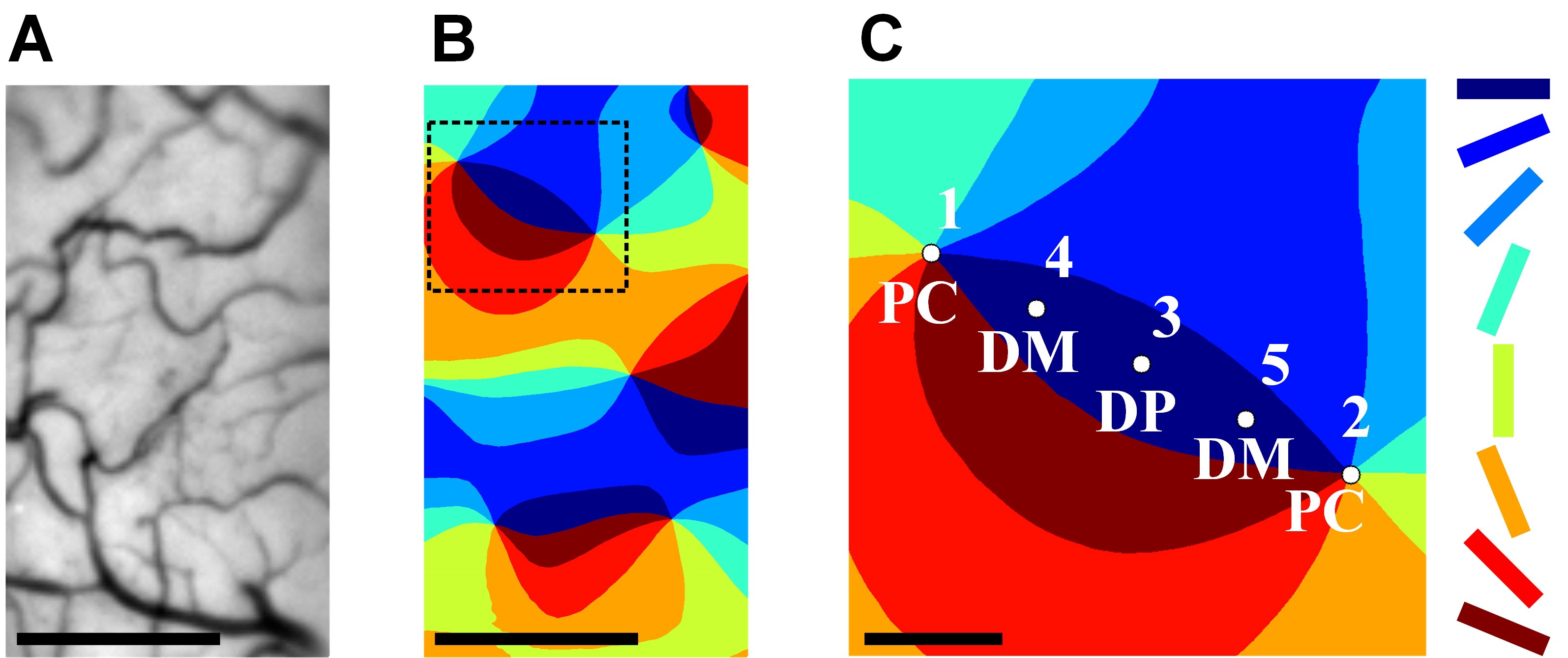
Figure 2. Highly accurate and quantitative determination of PC locations. A. Vascular pattern of the cortical surface. B. The color-coded orientation map of the cortex in (A). C. Two neighboring pinwheel centers (PC, locations 1 and 2) and three evenly spaced intervening points (DP, domain point, location 3; DM, domain midway, locations 4 and 5) in the orientation domain. The eight stimulus orientations are color coded at 22.5° intervals. Scale bars in (A) and (B): 500 μm. Scale bar in (C): 100 μm.
- Electrophysiological Recording Procedures
- Single unit isolation
- Insert and advance the electrode through the cortex, and record the signals using the Cerebus System.
- Spike signals were band-pass filtered at 250-7,500 Hz and sampled at 30 kHz. Only well-isolated cells satisfying the strict criteria for single-unit recordings (fixed shape of the action potential and the absence of spikes during the absolute refractory period) are selected and saved for further analyses.
- Insert and advance the electrode through the cortex, and record the signals using the Cerebus System.
- Single unit functional characterization
- All cells recorded were located in the area of the cortex representing the central 10° of the visual field. When the single-cell action potentials were isolated, the preferred orientation, SF, and TF of each cell were determined. Each cell was stimulated monocularly through the dominant eye, with the no dominant eye occluded. To locate the center of the CRF, a narrow rectangular sine wave grating patch (0.5° to 1.0° wide, 3.0° to 5.0° long, 100% contrast) was moved at successive positions along axes perpendicular or parallel to the optimal orientation of the cell, and the response to its drift was measured.
- The grating was set at the optimal orientation and SF and drifted in the preferred direction at the optimal speed for the recorded cells. The peak of the response profiles for both axes was defined as the center of the CRF. We determined the size of the CRF by performing an occlusion test, in which a mask consisting of a circular blank patch and centered on the CRF was gradually increased in size on a background drifting grating (Song and Li, 2008).
- All cells recorded were located in the area of the cortex representing the central 10° of the visual field. When the single-cell action potentials were isolated, the preferred orientation, SF, and TF of each cell were determined. Each cell was stimulated monocularly through the dominant eye, with the no dominant eye occluded. To locate the center of the CRF, a narrow rectangular sine wave grating patch (0.5° to 1.0° wide, 3.0° to 5.0° long, 100% contrast) was moved at successive positions along axes perpendicular or parallel to the optimal orientation of the cell, and the response to its drift was measured.
- LFP recording
- Obtain the orientation tuning of LFP signals.
- Present visual stimuli for LFP recordings (10° sine wave grating at 50% contrast), and pseudorandom sequences of gratings of varying orientation and spatial phase, each flashed for 32 ms.
- To analyze stimulus-evoked LFP responses, filter the recordings between 3 Hz and 100 Hz, and compute z scores by averaging responses across trials (for more detail, please see Figure 1 in Katzner et al. [2009]).
- Obtain the orientation tuning of LFP signals.
- Single unit isolation
- Procedure for Determining Pinwheel Centers Accurately
The following 3 steps ensure high accuracy of targeting PCs. (1) Identifying preliminary PC locations based on blood vessel maps. (2) Ensuring perpendicularity of electrodes. And (3) Selecting sites based on ‘non-sine-like’ LFPs.- Preliminarily identify pinwheel centers based on surface blood vessel pattern aligned to the orientation map (see Figures 2A and 2B).
- Make electrode penetrations perpendicular to the cortical surface. To ensure that the electrode penetration is perpendicular to the cortical surface, extra effort must be expended (Figures 3A-3C). First, to ensure the camera’s angle is exactly perpendicular to the plane of the cortex, image a large field of view (e.g., 1 cm) with a narrow depth of field (~50 μm, front-to-front lens with f1.2 [Ratzlaff and Grinvald, 1991]). We then obtain α value of the declination angle (α, shown in Figure 3D). Adjusted the declination angle of clamping device (clamp electrode) in Narishige stereotaxic instrument (whose mechanical accuracy is about 10 µm), made it equaling to α value, to ensure the electrode parallels exactly the axis of optical imaging. Using this procedure, we have estimated that the deviation of electrodes from perpendicular is less than 2 deg (shown in Figure 3D).
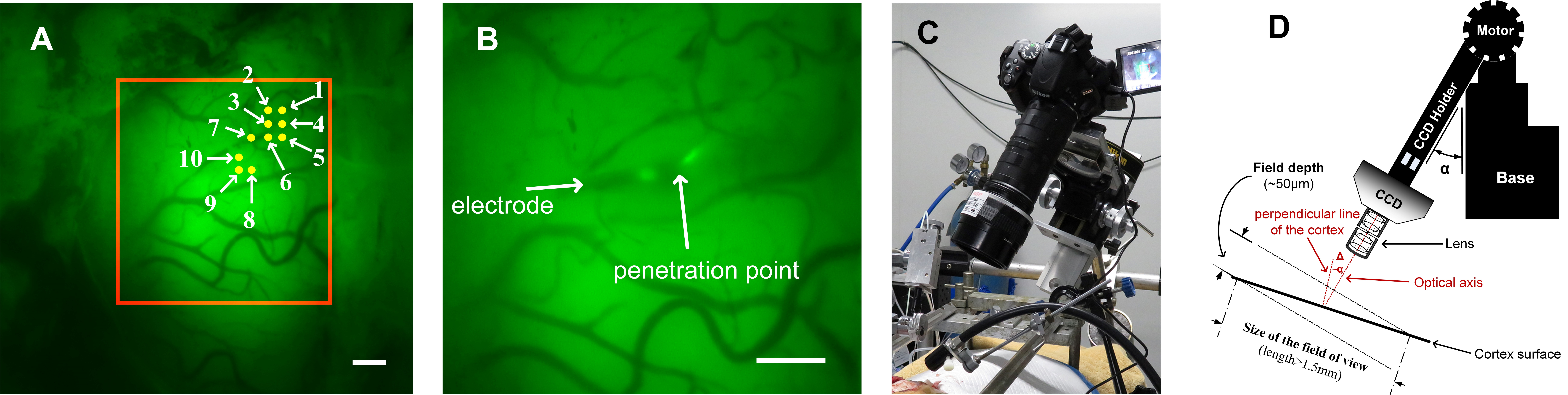
Figure 3. Illustration of electrode deviation from perpendicular. A. The yellow dots are the positions of 10 penetrations. Orange frame is the ROI (region of interest) in B. B. With the fine-tipped electrodes, there is a very little cortical deformation when the electrode is inserted. Scale bars in (A) and (B): 500 μm. C. We designed a lens (105-mm) with a close-up tube producing a working distance of 17 centimeters, which allows enough distance to make electrode penetrations obtaining images. D. The electrode was kept parallel to the optical axis by aligning the electrode at exactly the same angle as the CCD camera holder. The angle between the optical axis and the line perpendicular to the cortex (Δα) is Δα =actan (field_depth/field_length). As our depth of field is about 50 μm and the field of view is several millimeters, then we estimate Δα is less than 2 deg. - Selecting sites based on ‘non-sine-like’ LFPs
- Make a penetration at or near presumed PC within the superficial layers (depth < 600 μm) and record the LFP orientation tuning curve and fit with sinewave function. Then calculate the non-sine-like index (NSI) of this sinewave. See Data analysis. Repeat this for several penetrations within the very local region (about 200-500 μm diameter area around the preliminary PC) until a non-sine-like LFP curve is obtained (NSI > 0.10). Explanation: Based on our recent work (Li et al., 2019), we demonstrated that true PCs have ‘non-sine like’ LFPs (NSI > 0.1), while locations away from PCs exhibit ‘sine-like’ LFPs (Li et al., 2019). Figures 4A and 4B shows a ‘false’ PC site. The recorded LFP orientation tuning curve (4A) is sine-like (NSI = 0) and several single unit recordings in this penetration (4B) have low orientation scatter (MOS = 18°). Figures 4C and 4D shows a ‘true’ PC site. The recorded LFP orientation tuning curve (4C) is non–sine-like (NSI = 0.34) and several single unit recordings (4D) have high orientation scatter (MOS = 65°), a consistent marker of PCs (Maldonado et al., 1997). Across 26 sites, we showed there is a strong positive relationship between NSI and MOS (shown in the Figure 1F in reference [Li et al., 2019]; when the NSI > 0.1, the MOS > 60°). Thus an LFP of NSI > 0.1 provides precise localization of the pinwheel center.
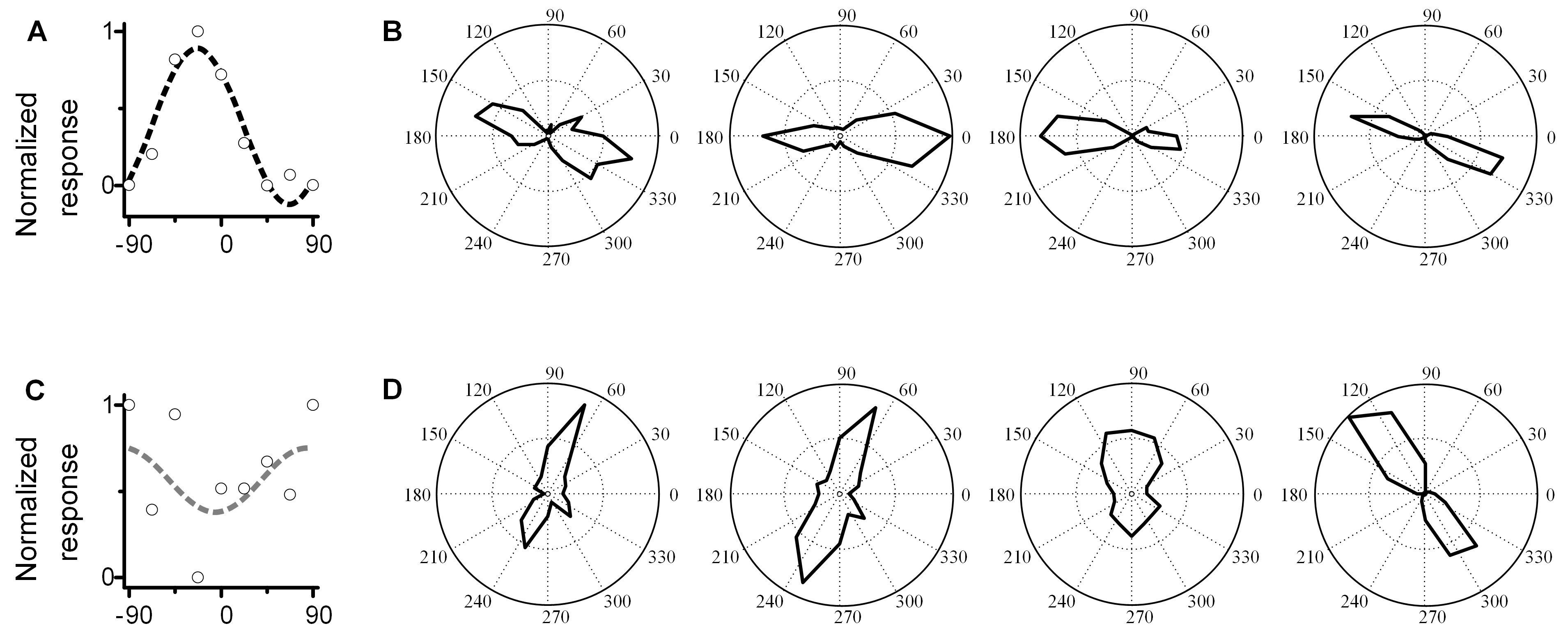
Figure 4. Comparison of LFP and single unit recordings in true PC site and false PC site. A. Examples of the sine-like LFP orientation tuning at the false PC site (NSI = 0, highly sine-like). B. Single unit recordings (four neurons) in same penetration as A. The MOS among these cells is 18°. C. An example of the non-sine-like LFP orientation tuning curve at the true PC site (NSI = 0.34). D. Single unit recordings (four neurons) in the same penetration as C. The MOS among these cells is 65°. - Record the position of the non-sine-like site. This is the first PC position (point 1 in Figure 2C).
- Repeat Steps D3a-D3b, and obtain an adjacent PC position (point 2 in Figure 2C).
- Make a penetration at or near presumed PC within the superficial layers (depth < 600 μm) and record the LFP orientation tuning curve and fit with sinewave function. Then calculate the non-sine-like index (NSI) of this sinewave. See Data analysis. Repeat this for several penetrations within the very local region (about 200-500 μm diameter area around the preliminary PC) until a non-sine-like LFP curve is obtained (NSI > 0.10). Explanation: Based on our recent work (Li et al., 2019), we demonstrated that true PCs have ‘non-sine like’ LFPs (NSI > 0.1), while locations away from PCs exhibit ‘sine-like’ LFPs (Li et al., 2019). Figures 4A and 4B shows a ‘false’ PC site. The recorded LFP orientation tuning curve (4A) is sine-like (NSI = 0) and several single unit recordings in this penetration (4B) have low orientation scatter (MOS = 18°). Figures 4C and 4D shows a ‘true’ PC site. The recorded LFP orientation tuning curve (4C) is non–sine-like (NSI = 0.34) and several single unit recordings (4D) have high orientation scatter (MOS = 65°), a consistent marker of PCs (Maldonado et al., 1997). Across 26 sites, we showed there is a strong positive relationship between NSI and MOS (shown in the Figure 1F in reference [Li et al., 2019]; when the NSI > 0.1, the MOS > 60°). Thus an LFP of NSI > 0.1 provides precise localization of the pinwheel center.
- Procedure for Determining DP and DM Positions
- Determine domain point (DP) by calculating the geometric midpoint of the two PCs identified in Step D (see Figure 2C).
- Determine domain midway (DM) by calculating the geometric midpoint of PC and DP (see Figure 2C).
- Determine domain point (DP) by calculating the geometric midpoint of the two PCs identified in Step D (see Figure 2C).
- Proceed with Electrophysiological Study of Orientation Subdomains
Once these key orientation domain anchors (PC, DP, DM) are determined, continue with electrophysiological characterization of orientation subdomains.
Data analysis
- Calculating NSI of an LFP curve
- Place the electrode, record a single unit and locate its receptive field. Determine the orientation tuning curve of the LFP at that site by presenting gratings at 8 orientations (see Figures 4A and 4C). Fit the LFP curve to a sine model. C0 denotes the LFP orientation tuning curve, which consists of K samplepoints, i.e., (x1, C0(x1)), (x2, C0(x2)),…, (xK, C0(xK)). The fitted curve is denoted by C'0.
- Calculate the fitting error(q0).
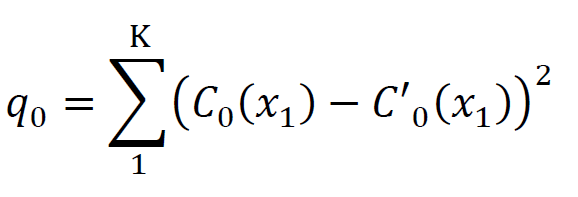
- Get the surrogate curves (C1 to C1000) by randomly shuffled the K sample-points of the LFP curve 1000 times and calutating the fitting errors of the surrogate curves (q1 to q1000).
- Calulate the NSI. The is defined as the p-value. It is ranged from 0 to 1, with 0 as highly sine-like and 1 as highly non-sine-like. The percentage of qj which is smaller than q0.
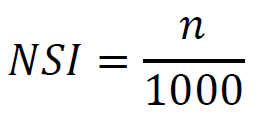
Where n is the number of the qj values which are smaller than q0.
Acknowledgments
This work was supported by the Major State Research Program (2018YFB1305101, 2015AA020515, 2013CB 329401) and the Natural Science Foundation of China (81430010, 91420302, and 31627802).
Competing interests
The authors declare that they have no competing interests.
Ethics
This study was performed in strict accordance with the recommendations contained in the Guide for the Care and Use of Laboratory Animals of the National Institutes of Health and approved by the Committee on the Ethics of Animal Experiments of the Shanghai Institute for Biological Sciences, Chinese Academy of Sciences (permit no. ERSIBS-621001C).
References
- Bonhoeffer, T. and Grinvald, A. (1991). Iso-orientation domains in cat visual cortex are arranged in pinwheel-like patterns. Nature 353(6343): 429-431.
- Das, A. and Gilbert, C. D. (1999). Topography of contextual modulations mediated by short-rangeinteractions in primary visual cortex. Nature 399(17): 655-661.
- Hashemi-Nezhad, M. and Lyon, D. C. (2012). Orientation tuning of the suppressive extraclassical surround depends on intrinsic organization of V1. Cereb Cortex 22(2): 308-326.
- Hubel, D. H. and Wiesel, T. N. (1974). Sequence regularity and geometry of orientation columns in the monkey striate cortex. J Comp Neurol 158(3): 267-293.
- Katzner, S., Nauhaus, I., Benucci, A., Bonin, V., Ringach, D. L. and Carandini, M. (2009). Local origin of field potentials in visual cortex. Neuron 61(1): 35-41.
- Li, C. Y., Xu, X. Z. and Tigwell, D. (1995). A simple and comprehensive method for the construction, repair and recycling of single and double tungsten microelectrodes. J Neurosci Methods 57(2): 217-220.
- Li, M., Song, X. M., Xu, T., Hu, D., Roe, A. W. and Li, C. Y. (2019). Subdomains within orientation columns of primary visual cortex. Science Advances, eaaw0807.
- Maldonado, P. E., Godecke, I., Gray, C. M. and Bonhoeffer, T. (1997). Orientation selectivity in pinwheel centers in cat striate cortex. Science 276(5318): 1551-1555.
- Mariño, J., Schummers, J., Lyon, D. C., Schwabe, L., Beck, O., Wiesing, P., Obermayer, K. and Sur, M. (2005). Invariant computations in local cortical networks with balanced excitation and inhibition. Nat Neurosci 8(2): 194-201.
- Nauhaus, I., Benucci, A., Carandini, M. and Ringach, D. L. (2008). Neuronal selectivity and local map structure in visual cortex. Neuron 57(5): 673-679.
- Nauhaus, I. and Ringach, D. L. (2007). Precise alignment of micromachined electrode arrays with V1 functional maps. J Neurophysiol 97(5): 3781-3789.
- Ohki, K., Chung, S., Kara, P., Hubener, M., Bonhoeffer, T. and Reid, R. C. (2006). Highly ordered arrangement of single neurons in orientation pinwheels. Nature 442(7105): 925-928.
- Ratzlaff, E. H. and Grinvald, A. (1991). A tandem-lens epifluorescence macroscope: hundred-fold brightness advantage for wide-field imaging. J Neurosci Methods 36(2-3): 127-137.
- Song, X. M. and Li, C. Y. (2008). Contrast-dependent and contrast-independent spatial summation of primary visual cortical neurons of the cat. Cereb Cortex 18(2): 331-336.
Article Information
Copyright
© 2020 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Song, X. M., Li, M., Xu, T., Hu, D. and Roe, A. W. (2020). Precise Targeting of Single Microelectrodes to Orientation Pinwheel Centers. Bio-protocol 10(11): e3643. DOI: 10.21769/BioProtoc.3643.
- Li, M., Song, X. M., Xu, T., Hu, D., Roe, A. W. and Li, C. Y. (2019). Subdomains within orientation columns of primary visual cortex. Science Advances, eaaw0807.
Category
Neuroscience > Sensory and motor systems > Visual system
Neuroscience > Sensory and motor systems > Animal model
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









