- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Isolating Multiple Extracellular Vesicles Subsets, Including Exosomes and Membrane Vesicles, from Bovine Milk Using Sodium Citrate and Differential Ultracentrifugation
Published: Vol 10, Iss 11, Jun 5, 2020 DOI: 10.21769/BioProtoc.3636 Views: 6666
Reviewed by: Karem A CourtAlexandros C KokotosAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
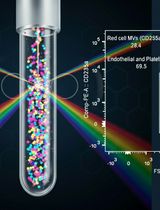
Protocol for the Isolation and Analysis of Extracellular Vesicles From Peripheral Blood: Red Cell, Endothelial, and Platelet-Derived Extracellular Vesicles
Bhawani Yasassri Alvitigala [...] Lallindra Viranjan Gooneratne
Nov 5, 2025 1502 Views

Optimized Secretome Sample Preparation From High Volume Cell Culture Media for LC–MS/MS Proteomic Analysis
Basil Baby Mattamana [...] Peter Allen Faull
Dec 20, 2025 1338 Views
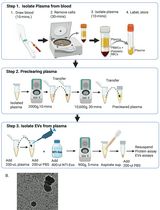
A Rapid and High-Recovery Extracellular Vesicle (EVs) Isolation Technique from Blood Samples
Aryan N. Sankpal and Narendra V. Sankpal
Mar 5, 2026 72 Views
Abstract
Milk is a complex fluid that contains various types of proteins and extracellular vesicles (EVs). Some proteins can mingle with EVs, and interfere with their isolation. Among these proteins, caseins form micelles of a size comparable to milk EVs, and can thus be co-isolated with EVs. Preliminary steps that affect milk are crucial for EV isolation and impact the purity and abundance of isolated EVs. In the course of our previous works on cow’s milk EVs, we found that sodium citrate (1% final), which is a biocompatible reagent capable of breaking down casein micelles into 40-nm monomers, allowed the isolation of high quantities of EVs with low coprecipitation of caseins or other contaminating proteins. Using this protocol, we successfully separated different EV subsets, characterized in depth their morphology, protein content and small RNA enrichment patterns. We were also able to describe their biological function in a mouse model of intestinal inflammation. We, hereby, detail the differential ultracentrifugation procedure that leads to high quantify, medium specificity, isolation of different milk EV subsets from the same sample. More specifically, we highlight the use of sodium citrate as a standardized approach to isolate and study milk EVs and its potential for isolation techniques other than differential ultracentrifugation.
Keywords: Extracellular VesiclesBackground
In our previous publications (Benmoussa et al., 2016, 2017, 2019b and 2019c; Benmoussa and Provost, 2019), we highlighted that bovine milk is a complex fluid containing a myriad of extracellular vesicles (EVs) subsets. Among these, exosomes are ~100-nm vesicles released when multivesicular bodies (MVB) fuse with the cell membrane. When subjected to ultracentrifugation, these sediment at centrifugation speeds equal or higher than 100,000 x g (P100K, where P stands for pellet) (Pieters et al., 2015). Other non-exosome EV subsets are found in milk and are comparable to exosomes, in shape and size, but sediments at a lower speed (e.g., 12,000 x g, P35K; 35,000 x g, P35K; 70,000 x g, P100K). These are thought to be originating from budding of the cell membrane and contain specific proteins and microRNAs different from exosomes content (Benmoussa et al., 2016, 2017, 2019b and 2019c; Benmoussa and Provost, 2019).
For a long time, separating these different milk EVs subsets remained challenging because of their morphological and chemical similarities. Moreover, protocols used in milk EV studies were often centered on the isolation of milk exosomes specifically, leading to the discarding of the biologically active non-exosome milk EVs (Benmoussa et al., 2019a, 2019b and 2019c).
When using differential ultracentrifugation to isolate milk EVs, the first steps are often impaired by the co-precipitation of milk proteins with EVs, which form a dense jelly at the bottom of the tubes (Zonneveld et al., 2014). Such jelly is formed when caseins are subjected to high mechanical pressures (Famelart et al., 1998; Zonneveld et al., 2014). Because this jelly is so dense, and practically impossible to resuspend, it is not clear whether some EV subsets are trapped within its matrix and some protocols recommend discarding this casein-rich jelly.
Certain reports suggested the use of density gradients or cushions to keep the EVs from mixing with the casein jelly (Zonneveld et al., 2014). Others suggested avoiding the formation of such jelly by discarding caseins before subjecting milk “serum” or whey to differential ultracentrifugation. To this end, milk is often mixed with acids (Somiya et al., 2018) or with cold EDTA to precipitate the caseins prior to milk EVs sedimentation (Wolf et al., 2015). However, such preprocessing of biological fluids have an immense impact on the isolation, quality and yield of extracellular vesicles (EVs) (Zonneveld et al., 2014). It also requires to discard low speed-pelleting EVs (P12K or P35K). In addition, there is a lack of information about the effect of acidification, or casein precipitation, on milk EVs, and the possibility that some EVs might be lost along the process.
This is especially important because there are multiple EV subsets in milk (Benmoussa et al., 2017 and 2019b) and because milk whey has a different microRNA and protein content than milk (Benmoussa et al., 2019b; Benmoussa and Provost, 2019), which, as microRNA are found within EVs, suggests the loss of certain EVs during casein precipitation. This is also supported by the discovery of microRNAs in milk-derived casein-rich products, like cheese (Benmoussa and Provost, 2019). Therefore, the biological activity of the milk EVs, and the bioactive molecules they transport, depend highly on the reagents/preprocessing steps and isolation protocols this fluid is exposed or subjected to (Zonneveld et al., 2014).
When we encountered this issue, we experimented different venues to avoid precipitating milk caseins while ensuring the separation of different structurally conserved, and functional, milk EV subsets. In milk, casein proteins are arranged in the form of micelles that are 70 to 200 nm in diameter, which is close to the size of certain EV subsets (Blans et al., 2017). Knowing that calcium is important for maintaining these micelles (de Kort et al., 2009, 2011 and 2012; Kort et al., 2012), we supposed that calcium chelation would prevent the formation of casein superstructures that hamper EV isolation by ultracentrifugation.
Several calcium-chelating agents have been previously investigated for their ability to disrupt casein micelles, including disodium uridine monophosphate (Na2UMP), disodium phosphate (Na2HPO4), trisodium citrate (referred to simply as "sodium citrate"), sodium phytate, sodium hexametaphosphate (SHMP) or ethylenediaminetetraacetic acid (EDTA) (Ward et al., 1997; de Kort et al., 2009, 2011 and 2012; Kort et al., 2012). These chelating agents interact with colloidal calcium phosphate (CCP) leading to the breaking of casein micelles into small monomers. This process augments the stability of milk during heating and change its physical properties, often reducing its viscosity (de Kort et al., 2009, 2011 and 2012; Kort et al., 2012).
Among these, sodium citrate is a biocompatible compound known for 100+ years for its ability to prevent casein curdling in the stomach of infants (Poynton, 1904). It is widely used to preserve biological fluids, like blood (Janse van Rensburg and van der Merwe, 2017), as a food additive and as the major form of rehydration salt recommended by the World Health Organization (Pizarro et al., 1986; Banipal et al., 2016). Notably, it was previously used to disrupt casein micelles to facilitate the isolation of milk fat globules (MFGs) by diafiltration and prevented pore clogging (Phan et al., 2014). Interestingly, sodium citrate specifically impacts milk protein gel formation upon high mechanical pressure, although the underlying mechanisms remain unclear, except for its dependence on the solution’s pH (Famelart et al., 1998).
In the course of our experimentations with sodium citrate, we found that pre-treating milk with sodium citrate (1% final) was a cost-effective way to avoid casein jellification and isolation of high quantities of different milk EVs using differential ultracentrifugation. We hereby detail the methods underlying this method that led to the discovery (Benmoussa et al., 2016 and 2017) and characterization (Benmoussa et al., 2017, 2019b and 2019c) of different EVs subsets in milk, keeping them functional and able to transfer their content to human cells (Benmoussa et al., 2019c). EVs, including exosomes, isolated by this methodology were able to modulate intestinal inflammation during experimental colitis (Benmoussa et al., 2019a).
Materials and Reagents
- Sterilized glass bottle (Pyrex, Corning Life-Science, catalog number: 1395-1L) or sterile plastic bottles for mixing milk and sodium citrate (Corning, catalog number: 431533)
- Falcon tubes 50 ml (any product, as long as sterile and pyrogen-free, e.g., Corning, catalog number: 352070, Fisher Scientific, catalog number: 14-432-22)
- 1.7 ml snap cap tubes (any product as long as sterile and pyrogen-free, e.g., Corning, Costar via Sigma-Aldrich, catalog number: CLS3620)
- 0.22 µm membrane microfilters (Corning, Sigma-Aldrich, catalog number: CLS431224)
- Serological pipets different volumes (any product as long as sterile and pyrogen-free, e.g., Sigma-Aldrich, catalog number: SIAL1485)
- Filtered 1 ml pipet tips (any product as long as sterile, pyrogen free and filtered, e.g., Thermo Scientific, catalog number: 94052410)
- Nitrile disposable gloves (any type)
- Sterile wipes/gauze (any as long as sterile single-packed, e.g., VWR, catalog number: CA95041-740)
- Optional: EDTA. Can be bought as ready-to-use solution (Sigma-Millipore, catalog number: 324506) or homemade from powdered EDTA
- Milk
Use commercially available milk bought on the day of the experiment or raw cow milk collected and stored in “sterile conditions” (as sterile as possible using sterilized bottles and keeping the bottles closed until getting under a biological hood). In our work we used mostly pasteurized commercial ultrafiltered skim milk (Lactantia PureFilter, bought in a local grocery store) as it avoids creaming steps that could lead to variability. We recommend working with a pool of three milks with different preemption dates. Milk must be stored at 4 °C.
Note: If working with raw milk, use pool from different cows to minimize interindividual differences. Noncommercial raw or pasteurized fresh cow milk can be frozen at -80 °C to avoid bacterial contamination/proliferation but note such storage might impact EVs content. If freezing milk, make sure it is previously skimmed and decellularized to minimize EVs formations due to milk fat globule or cell destruction upon freeze/thawing cycles. Freeze milk in small aliquots (50 ml) so, when thawing, it reaches melting point quicker and limit EVs destruction. - Bradford's reagent (Sigma-Aldrich, catalog number: B6916)
- BCA Protein Assay Kit (Millipore, catalog number: 71285-M)
- Milli-Q water or any filtered ultra-pure water (e.g., Millipore, water SystemClear Sorting & Filtering, catalog number: ZRXQ003WW)
- NaCl (preferably low in endotoxins, e.g., Millipore-Sigma, SAFC, catalog number: 1.16224)
- KCl (e.g., Sigma-Aldrich, catalog number: P3911)
- Na2HPO4 (e.g., Sigma-Aldrich, catalog number: NIST2186II)
- KH2PO4 (e.g., Sigma-Aldrich, catalog number: NIST200B)
- HCl (e.g., Sigma-Aldrich, catalog number: 320331)
- EDTA (e.g., Sigma-Aldrich, catalog number: EDS-500G)
- NaOH solution in Milli-Q water (e.g., Sigma-Aldrich, catalog number: S8045)
- Sodium citrate dihydrate (Sigma-Aldrich, catalog number: W302600)
- Phosphate buffered saline
Either bought as commercially available as filtered sterile phosphate buffered saline (PBS) pH 7.4 (Sigma-Aldrich, catalog number: P5493) or homemade. - 2% sodium citrate solution (1 L) (see Recipes)
- Phosphate buffer saline (PBS) 10x (1 L) (see Recipes)
- EDTA 0.5 M recipe (100 ml) (see Recipes)
Equipment
- Tubes for ultracentrifugation compatible with rotor bellow (36 ml, Beckman, catalog number: 355631 or 344058 or other, see rotor compatibility)
- Ultracentrifuge Sorvall WX ultracentrifuge (or one of Thermo ScientificTM SorvallTM WX ultraCentrifuges, catalog numbers: 75000100, 7500090, 7500080)
- Rotor SureSpin 630 swinging bucket Rotor (Thermo Scientific, catalog number: 79368)
Note: You can use another centrifuge and another rotor but make sure to get comparable material. Note that it is not recommended to use fixed-angle rotors as they will induce higher EV degradation. - Pipetting device (any type, e.g., Drummond Scientific Portable Pipet-Aid XP via Mandel, catalog number: DRU-4-000-101)
- Orbital shaker (any type, e.g., Bel-ArtTM SP SciencewareTM SpindriveTM Orbital Shaker Platform, via Fisher-scientific, catalog number: 1451176)
- Rotating mixer/tube revolver (any type, e.g., Thermo-scientific, catalog number: 88881001)
- High precision balance (any type, e.g., Sartorius, catalog number: PRACTUM224-1S)
- Tube rack holder different sizes (any product)
- Laminar flow hood (if working on sterile conditions) (any product)
- Micropipettes, different volumes (any type as long as they are precise and pipetting is smooth enough to avoid too fast ejection of suspension liquid, e.g., PIPETMAN L P1000L, 100-1,000 µl, Metal Ejector, Gilson, catalog number: FA10006M)
- -80 °C freezer
- Magnetic stirrer
Procedure
Note: All procedure should be performed at 4 °C.
- Start ultracentrifuge and set device temperature at 4 °C.
- Mix 125 ml of skimmed milk and 2% sodium citrate 1:1 in a sterile glass or plastic bottle.
Note: Pour sodium citrate first then milk to accelerate casein micelles breaking. - Keep on a rocker for 15 min at 4 °C. Make sure milk is gently mixing (1/4th max speed).
Note: This can be done in cold room or by putting ice on the rocker and milk on the ice. - After 15 min, milk should clarify (get translucid with an aspect comparable to blood plasma) and ready for loading into ultracentrifugation tubes (Figure 1).

Figure 1. Mixing skimmed milk with 2% sodium citrate 1:1 leads to milk clarification after 15 min incubation on a rocker at 4 °C
Notes:- Adding calcium carbonate reverses the clarification confirming the involvement of calcium chelation in the process.
- If the milk you use is not clarified, you can increase sodium citrate concentration to 3% or 4%.
- Fill the ultra-clear centrifuge tubes (Figure 2) completely with milk-sodium citrate mix.
Note: Make sure the tubes are filled completely or they might break upon centrifugation. - Put tubes in rotor tube holders and use sterile pipet to equilibrate mass between opposed tubes (Figure 2) by transferring milk-sodium-citrate mix between the opposed tubes (keeping comparable added mass of the tube, milk-citrate solution, tube holder and cap).

Figure 2. Ultraclear thick ultracentrifugation tubes should be loaded into the tube holder before mass equilibration for ultracentrifugation - Close caps and place them in the socket they are meant for (indicated by the number on the tube holder and the rotor, Figure 3).

Figure 3. Ensure loading SureSpin tubes in the right socket to avoid equilibrium troubles - Spin EVs are the desired speed. In our protocol we isolated low-speed sedimenting EVs at 35,000 x g (35K) for 2 h, at 4 °C. We used Sorvall WX TL-100 ultracentrifuge’s automated calculations of the K factors and set acceleration at A = 9 and break at D = 9.
Note: If working with another ultracentrifuge, make sure to translate the g-force speed into the proper rotation per minute (RPM) considering rotor angle and k factor. If wishing to use different speeds, refer to the rotor and centrifuge manufacturer recommendations. If you work with raw milk, you can decellularize the liquid with two centrifugations at 1,000 x g for 10 min and 4,500 x g for 30 min, at 4 °C. Afterwards, you can proceed as described above. - After 2 h, EVs should have sedimented forming a translucid gelatinous pellet on the bottom of the tube (Figure 4C).
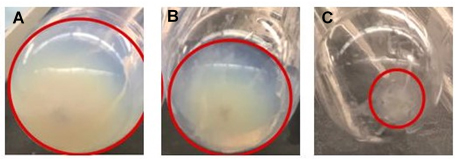
Figure 4. Non-diluted and PBS-diluted milk led to the formation of casein jelly after 35,000 x g ultracentrifugation. Dilution with sodium citrate leads to small EV-rich pellet with very little casein contamination. A. 35,000 x g pellet from non-treated milk. B. 35,000 x g pellet from diluted milk 1:1 in PBS. C. 35,000 x g pellet from diluted milk 1:1 in 2% sodium citrate contains very little contaminating casein.
Note: The pellet on the right contains P35K EVs. It might not be visible prior to discarding the supernatant and suspension. It is well fixed to the bottom of the tube so there is little risk to lose it in the next steps. - Transfer the supernatant into new clean/sterile ultracentrifugation tubes.
Note: This can be done using a serological pipet and pipetting device or directly by pouring the supernatant into a new tube, this depending on the desired sterility levels. - Equilibrate the supernatant tubes and centrifuge at 70,000 x g (70K) for 1 h, at 4 °C following the same procedure as previously.
- During the 70K ultracentrifugation, revert the remaining liquid free 35K pellet-containing tubes on a tube holder with sterile gauze/paper on the bottom of it, with the opening of the tubes touching the gauze/paper to allow remaining liquid to be discarded by capillarity within the gauze/paper (Figure 5).

Figure 5. Reverting tubes over a paper sheet allows remaining liquid to be discarded by capilarity
Note: This is important to avoid contamination with proteins from the supernatant and to keep precise suspension volumes. You can use pipet and tips to discard any remaining liquid or sterile gauze/paper inside the tube. - Suspend all the pellets (6 tubes) in the same 1 ml sterile filtered PBS solution.
Notes:- The pellet is gelatinous and not easy to suspend. Avoid breaking it with the tips. Use pipet and 1 ml filtered tip to drop PBS on it and use PBS to gently “erode” the pellet starting from its sides and getting gently to the center. Try to be quick and efficient to avoid drying of the pellets but avoid rough mixing or you will end up with a foam limiting pellet suspension.
- You can also pour 100 μl PBS on the tubes you are not suspending while using 500 μl PBS to suspend the first tube. Transfer the 500 μl to the second tube having thus 600 μl for the second pellet, and so on. Depending on your application, you can include 0.5% EDTA to the PBS to help with the suspension with caution if planning on doing qPCR or for functional studies.
- Transfer suspended P35K EVs into a 1.7 ml Eppendorf tube and put on a rotating mixer at 4 °C at least for 24 h to allow full suspension of P35K EVs. EVs can then be stored at -80 °C for few days. However, this might lead to a certain level of degradation and EVs are thus better to be used on the day of preparation. EVs can also be filtered through 0.22 µm membrane microfilters to ensure their sterility, and to break down EV aggregates, before using them for any application.
- After P70K ultracentrifugation, proceed as previously for P35K EVs, transfer the supernatant in new ultracentrifugation tubes and subject these tubes to a 100,000 x g ultracentrifugation (100K) at 4 °C for at least 1 h (the longer, the higher yield of exosomes will be reached) (Figure 6).
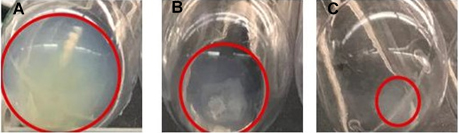
Figure 6. Non-diluted and PBS-diluted milk led to the formation of casein jelly after 70,000 x g ultracentrifugation. Dilution with sodium citrate leads to small EV-rich pellet with very little casein contamination. A. 70,000 x g pellet from non-treated milk. B. 70,000 x g pellet from diluted milk 1:1 in PBS. C. 70,000 x g pellet from diluted milk 1:1 in 2% sodium citrate.
Note: Discard 70K pellet or keep it if interested by its content (Figure 6C). If so, proceed to its suspension as for P35K EVs. - After P100K ultracentrifugation, transfer the 100K supernatant to new ultracentrifugation tubes and subject to 100,000 x g ultracentrifugation for 18 h at 4 °C if you wish to obtain an EV-free supernatant to use as a control.
- Suspend 100K pellet (Figure 7C) the same way as for previous pellets and store EVs the same way.
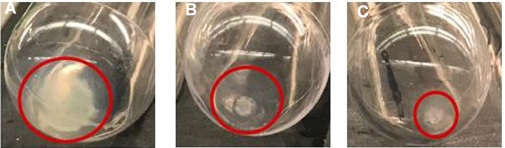
Figure 7. Non-diluted and PBS-diluted milk led to the formation of casein jelly after 100,000 x g ultracentrifugation. Dilution with sodium citrate leads to small EV-rich pellet with very little casein contamination. A. 100,000 x g pellet from non-treated milk. B. 100,000 x g pellet from diluted milk 1:1 in PBS. C. 100,000 x g pellet from diluted milk 1:1 in 2% sodium citrate.
Data analysis
This protocol does not generate data but functional EVs. The quality and content of these can be assessed through different approaches.
- EV concentration
EVs content of the final pellets can be indirectly quantified by assessing their protein content using Bradford's reagent (Sigma-Aldrich) or BCA Protein Assay Kit (Millipore), by isolating total RNA and determining its concentration (NanoDrop, Bioanalyzer, Qbit, etc.) or more precisely by directly quantifying EVs using high sensitivity flow cytometry (Benmoussa et al., 2017; Morales-Kastresana and Jones, 2017) or nanoparticle tracking analysis (NTA) (Gardiner et al., 2013). - EV characteristics and quality
Cow’s milk EVs characteristics can be routinely checked using densitometry, dynamic light scattering (DLS) and electron microscopy as previously described (Benmoussa et al., 2017). - EV characterization
We provided in our previous publications methodologies to analyze milk EV’s proteins (Benmoussa et al., 2019), small RNAs (Benmoussa et al., 2019c), resistance to digestion and bioaccessibility (Benmoussa et al., 2016) and bioactivity (Benmoussa et al., 2019a). These methodologies include EV density analysis, exploration of protein content by western blot, proteins profile by liquid-chromatography tandem mass spectrometry, small RNA and mRNA profiling by RNA microarray or next generation sequencing, TIM-1 in vitro digestion, dual luciferase assays, etc. - EV functional activity
Depending on the expected outcomes, different approaches might be suitable, and we would recommend to anyone exploring the field of milk EVs to read and follow MISEV2018 guidelines (Thery et al., 2018). For those exploring milk EVs biological activity on inflammation when delivered orally, we would recommend exploring, along with MISEV2018, the report associated to this protocol (Benmoussa et al., 2019a).
Notes
Through the setup of this protocol, we discovered that addition of sodium citrate (used at a final concentration of 1%) made cow’s milk translucid, comparable to plasma in appearance and viscosity. More importantly, it prevented the co-isolation of milk caseins and other milk proteins with different milk EV subsets (Famelart et al., 1998; Kort, 2012; Benmoussa et al., 2017), which confirmed the importance of calcium in casein gel formation upon ultracentrifugation. It is likely that our discovery of different EV subset in cow’s milk was made possible by the change in milk viscosity induced by sodium citrate (Benmoussa et al., 2017). This change in viscosity would also explain the relative purity of the milk EVs we isolated in our previous work, as it may prevent sedimentation of certain proteins or aggregates at the considered speeds (Benmoussa et al., 2016, 2017 and 2019b).
Importantly, we diluted milk in a 1:1 ratio with a 2% sodium citrate solution, rather than dissolving citrate crystals in milk, which would have changed the biological properties (e.g., viscosity) of milk even more and possibly impeding the isolation of certain EVs (Momen-Heravi et al., 2012).
Also, in our hands, EDTA used at a concentration sufficient to chelate all calcium in milk, did not have the same effect on milk as sodium citrate, and actually causes casein precipitation (Wolf et al., 2015). Also, in our previous works, we found that acidified milk or preprocessing with EDTA led to changes in microRNA content in EVs and exosomes in comparison to whole milk (Benmoussa et al., 2019c). EDTA may differ from the other chelators discussed above in that it may not provide the sodium that links to caseins to form soluble sodium caseinates (de Kort et al., 2009, 2011 and 2012; Kort et al., 2012).
This protocol has shown robust reproducibility over 5 years of use within the hands of more than 7 manipulators. It worked with different bovine cow milks (raw and commercial pasteurized milk) and would be believed to function with milk from different species.
It is, however, of importance to note that differential ultracentrifugation is a long process that, in certain conditions, can have deleterious effects on EVs quality (Thery et al., 2018). Filtration steps might also discard some EVs of importance. Therefore, one should always characterize the full content of milk EVs before discarding potentially important populations and compare the effects of the EVs before and after filtering. It might also be important to consider other approaches to isolate milk EVs and for which sodium citrate is compatible. As it is known to break casein micelles, which are roughly the size of EVs (~200 nm), sodium citrate might provide the means to isolate milk EVs through tangential filtration or continuous diafiltration (Phan et al., 2014; Busatto et al., 2018), and help with the use of size-exclusion chromatography, by relieving the need to discard caseins (Blans et al., 2017). It may even be considered for isolating EVs from industrial scale volumes of milk, in a continuous way, using simulated moving bed size-exclusion chromatography (Satzer et al., 2014). It is also important to note that the use of sodium citrate might not be compatible with other approaches or milks from other species. Validation should be done prior to any long-term / large-scale implementation of this methodology.
In any case, and independently of the chosen protocol, filling the 2018 MISEV guideline checklist and attaching it to the chosen protocol is highly recommended to ensure replicability and proper reporting in milk EVs studies.
Recipes
- 2% sodium citrate solution (1 L)
- Dilute 20 g of sodium citrate dihydrate in 1 L of autoclaved water filtered through 0.22 µm membrane microfilters or in autoclaved Milli-Q water
- Stir for 10 min using magnetic stirrer
- Filter through 0.22 µm membrane microfilters
- Phosphate buffer saline (PBS) 10x (1 L)
In 800 ml of Milli-Q water- Add 8 g of NaCl (preferably low in endotoxins, e.g., Millipore-Sigma, SAFC)
- Add 0.2 g of KCl
- Add 1.44 g of Na2HPO4
- Add 0.24 g of KH2PO4
- Adjust the pH to 7.4 with HCl
- Complete with Milli-Q water to a total volume of 1 L
- Filter the solution through 0.22 µm membrane microfilters
Note: Solution can be autoclaved to ensure further sterility. - Dilute 10x stock solution to 1x and before using it, filter the diluted PBS through 0.22 µm membrane microfilters
Note: Stock solution can be stored at room temperature. Diluted solutions should be stored in sterile conditions at 4 °C to avoid contamination for few weeks.
- EDTA 0.5 M recipe (100 ml)
In 80 ml of Milli-Q water:- Add 18.6 g of EDTA
- Stir using magnetic stirrer for 10 min
- Adjust pH to 8.0 using NaOH in Milli-Q water solution
- Adjust to 100 ml
- Complete with Milli-Q water to a total volume of 100 ml and filter it through 0.22 µm membrane microfilters
Acknowledgments
This work was supported by the Canadian Institutes of Health Research (CIHR) Grants No. IG1-134171, MOP-137081 (through the Institute of Genetics) and PJT-165806 (to P.P.). A.B. (No. 262093) received a PhD studentship award from the FRQ-S. Authors wish to acknowledge Patricia Savard for her help in solving casein jelly formation. This protocol was previously used in our reported works (Benmoussa et al., 2016, 2017, 2019a, 2019b and 2019c).
Competing interests
The author(s) declare no competing interests and, as we worked with commercially available cow milk, the authors were under no requirement of ethical committee approval.
References
- Banipal, T. S., Kaur, H., Kaur, A. and Banipal, P. K. (2016). Effect of tartarate and citrate based food additives on the micellar properties of sodium dodecylsulfate for prospective use as food emulsifier. Food Chem 190: 599-606.
- Benmoussa, A., Diallo, I., Salem, M., Michel, S., Gilbert, C., Sevigny, J. and Provost, P. (2019a). Concentrates of two subsets of extracellular vesicles from cow's milk modulate symptoms and inflammation in experimental colitis. Sci Rep 9(1): 14661.
- Benmoussa, A., Gotti, C., Bourassa, S., Gilbert, C. and Provost, P. (2019b). Identification of protein markers for extracellular vesicle (EV) subsets in cow's milk. J Proteomics 192: 78-88.
- Benmoussa, A., Laugier, J., Beauparlant, C. J., Lambert, M., Droit, A. and Provost, P. (2019c). Complexity of the microRNA transcriptome of cow milk and milk-derived extracellular vesicles isolated via differential ultracentrifugation. J Dairy Sci 103(1): 16-29.
- Benmoussa, A., Lee, C. H., Laffont, B., Savard, P., Laugier, J., Boilard, E., Gilbert, C., Fliss, I. and Provost, P. (2016). Commercial dairy cow milk micrornas resist digestion under simulated gastrointestinal tract conditions. J Nutr 146(11): 2206-2215.
- Benmoussa, A., Ly, S., Shan, S. T., Laugier, J., Boilard, E., Gilbert, C. and Provost, P. (2017). A subset of extracellular vesicles carries the bulk of microRNAs in commercial dairy cow's milk. J Extracell Vesicles 6(1): 1401897.
- Benmoussa, A. and Provost, P. (2019). Milk MicroRNAs in health and disease. In: Comprehensive Reviews in Food Science and Food Safety. In: Benmoussa, A. and Provost, P. (Eds.). DOI: 10.1111/1541-4337.12424.
- Blans, K., Hansen, M. S., Sorensen, L. V., Hvam, M. L., Howard, K. A., Moller, A., Wiking, L., Larsen, L. B. and Rasmussen, J. T. (2017). Pellet-free isolation of human and bovine milk extracellular vesicles by size-exclusion chromatography. J Extracell Vesicles 6(1): 1294340.
- Busatto, S., Vilanilam, G., Ticer, T., Lin, W. L., Dickson, D. W., Shapiro, S., Bergese, P. and Wolfram, J. (2018). Tangential flow filtration for highly efficient concentration of extracellular vesicles from large volumes of fluid. Cells 7(12).
- de Kort, E., Minor, M., Snoeren, T., van Hooijdonk, T. and van der Linden, E. (2012). Effect of calcium chelators on heat coagulation and heat-induced changes of concentrated micellar casein solutions: The role of calcium-ion activity and micellar integrity. Int Dairy J 26(2): 112-119
- Famelart, M. H., Chapron, L., Piot, M., Brulé, G. and Durier, C. (1998). High pressure-induced gel formation of milk and whey concentrates. J Food Eng 36(2): 149-164.
- Gardiner, C., Ferreira, Y. J., Dragovic, R. A., Redman, C. W. G. and Sargent, I. L. (2013). Extracellular vesicle sizing and enumeration by nanoparticle tracking analysis. J Extracell Vesicles 2(1): 19671.
- Janse van Rensburg, W. J. and van der Merwe, P. (2017). Comparison of commercially available blood collection tubes containing sodium citrate and hirudin in platelet aggregation testing. Med Sci Monit Basic Res 23: 264-269.
- Kort, E.J.P. de. (2012). Influence of calcium chelators on concentrated micellar casein solutions: from micellar structure to viscosity and heat stability. In: Kort, E. J. P. de. (Ed). ISBN: 978-94-6173-237-8.
- de Kort, E., Minor, M., Snoeren, T., Hooijdonk, T. and Linden, E. (2011). Effect of calcium chelators on physical changes in casein micelles in concentrated micellar casein solutions. Int Dairy J 21(12): 907-913.
- de Kort, E., Minor, M., Snoeren, T. H. M., Hooijdonk, T. and Linden, E. (2009). Calcium-binding capacity of organic and inorganic ortho- and polyphosphates. Dairy Sci Technol 89(3-4): 283-299.
- Momen-Heravi, F., Balaj, L., Alian, S., Trachtenberg, A. J., Hochberg, F. H., Skog, J. and Kuo, W. P. (2012). Impact of biofluid viscosity on size and sedimentation efficiency of the isolated microvesicles. Front Physiol 3: 162-162.
- Morales-Kastresana, A. and Jones, J. C. (2017). Flow cytometric analysis of extracellular vesicles. Methods Mol Biol 1545: 215-225.
- Phan, T. T., Le, T. T., Van der Meeren, P. and Dewettinck, K. (2014). Comparison of emulsifying properties of milk fat globule membrane materials isolated from different dairy by-products. J Dairy Sci 97(8): 4799-4810.
- Pieters, B. C. H., Arntz, O. J., Bennink, M. B., Broeren, M. G. A., van Caam, A. P. M., Koenders, M. I., van Lent, P. L. E. M., van den Berg, W. B., de Vries, M., van der Kraan, P. M. and van de Loo, F. A. J. (2015). Commercial cow milk contains physically stable extracellular vesicles expressing immunoregulatory TGF-β. PLOS ONE 10(3): e0121123.
- Pizarro, D., Posada, G., Segreda, O. and Mata, L. (1986). Comparison of the efficacy of 2 oral rehydration solutions: the conventional solution recommended by WHO containing sodium bicarbonate and another containing sodium citrate. Bol Med Hosp Infant Mex 43(7): 402-406.
- Poynton. (1904). An Address ON THE VALUE OF THE ADDITION OF CITRATE OF SODA TO COW'S MILK IN INFANT FEEDING. The Lancet 164(4224): 433-436.
- Satzer, P., Wellhoefer, M. and Jungbauer, A. (2014). Continuous separation of protein loaded nanoparticles by simulated moving bed chromatography. J Chromatogr A 1349(100): 44-49.
- Somiya, M., Yoshioka, Y. and Ochiya, T. (2018). Biocompatibility of highly purified bovine milk-derived extracellular vesicles. J Extracell Vesicles 7(1): 1440132.
- Thery, C., Witwer, K. W., Aikawa, E., Alcaraz, M. J., Anderson, J. D., Andriantsitohaina, R., Antoniou, A., Arab, T., Archer, F., Atkin-Smith, G. K., Ayre, D. C., Bach, J. M., Bachurski, D., Baharvand, H., Balaj, L., Baldacchino, S., Bauer, N. N., Baxter, A. A., Bebawy, M., Beckham, C., Bedina Zavec, A., Benmoussa, A., Berardi, A. C., Bergese, P., Bielska, E., Blenkiron, C., Bobis-Wozowicz, S., Boilard, E., Boireau, W., Bongiovanni, A., Borras, F. E., Bosch, S., Boulanger, C. M., Breakefield, X., Breglio, A. M., Brennan, M. A., Brigstock, D. R., Brisson, A., Broekman, M. L., Bromberg, J. F., Bryl-Gorecka, P., Buch, S., Buck, A. H., Burger, D., Busatto, S., Buschmann, D., Bussolati, B., Buzas, E. I., Byrd, J. B., Camussi, G., Carter, D. R., Caruso, S., Chamley, L. W., Chang, Y. T., Chen, C., Chen, S., Cheng, L., Chin, A. R., Clayton, A., Clerici, S. P., Cocks, A., Cocucci, E., Coffey, R. J., Cordeiro-da-Silva, A., Couch, Y., Coumans, F. A., Coyle, B., Crescitelli, R., Criado, M. F., D'Souza-Schorey, C., Das, S., Datta Chaudhuri, A., de Candia, P., De Santana, E. F., De Wever, O., Del Portillo, H. A., Demaret, T., Deville, S., Devitt, A., Dhondt, B., Di Vizio, D., Dieterich, L. C., Dolo, V., Dominguez Rubio, A. P., Dominici, M., Dourado, M. R., Driedonks, T. A., Duarte, F. V., Duncan, H. M., Eichenberger, R. M., Ekstrom, K., El Andaloussi, S., Elie-Caille, C., Erdbrugger, U., Falcon-Perez, J. M., Fatima, F., Fish, J. E., Flores-Bellver, M., Forsonits, A., Frelet-Barrand, A., Fricke, F., Fuhrmann, G., Gabrielsson, S., Gamez-Valero, A., Gardiner, C., Gartner, K., Gaudin, R., Gho, Y. S., Giebel, B., Gilbert, C., Gimona, M., Giusti, I., Goberdhan, D. C., Gorgens, A., Gorski, S. M., Greening, D. W., Gross, J. C., Gualerzi, A., Gupta, G. N., Gustafson, D., Handberg, A., Haraszti, R. A., Harrison, P., Hegyesi, H., Hendrix, A., Hill, A. F., Hochberg, F. H., Hoffmann, K. F., Holder, B., Holthofer, H., Hosseinkhani, B., Hu, G., Huang, Y., Huber, V., Hunt, S., Ibrahim, A. G., Ikezu, T., Inal, J. M., Isin, M., Ivanova, A., Jackson, H. K., Jacobsen, S., Jay, S. M., Jayachandran, M., Jenster, G., Jiang, L., Johnson, S. M., Jones, J. C., Jong, A., Jovanovic-Talisman, T., Jung, S., Kalluri, R., Kano, S. I., Kaur, S., Kawamura, Y., Keller, E. T., Khamari, D., Khomyakova, E., Khvorova, A., Kierulf, P., Kim, K. P., Kislinger, T., Klingeborn, M., Klinke, D. J., 2nd, Kornek, M., Kosanovic, M. M., Kovacs, A. F., Kramer-Albers, E. M., Krasemann, S., Krause, M., Kurochkin, I. V., Kusuma, G. D., Kuypers, S., Laitinen, S., Langevin, S. M., Languino, L. R., Lannigan, J., Lasser, C., Laurent, L. C., Lavieu, G., Lazaro-Ibanez, E., Le Lay, S., Lee, M. S., Lee, Y. X. F., Lemos, D. S., Lenassi, M., Leszczynska, A., Li, I. T., Liao, K., Libregts, S. F., Ligeti, E., Lim, R., Lim, S. K., Line, A., Linnemannstons, K., Llorente, A., Lombard, C. A., Lorenowicz, M. J., Lorincz, A. M., Lotvall, J., Lovett, J., Lowry, M. C., Loyer, X., Lu, Q., Lukomska, B., Lunavat, T. R., Maas, S. L., Malhi, H., Marcilla, A., Mariani, J., Mariscal, J., Martens-Uzunova, E. S., Martin-Jaular, L., Martinez, M. C., Martins, V. R., Mathieu, M., Mathivanan, S., Maugeri, M., McGinnis, L. K., McVey, M. J., Meckes, D. G., Jr., Meehan, K. L., Mertens, I., Minciacchi, V. R., Moller, A., Moller Jorgensen, M., Morales-Kastresana, A., Morhayim, J., Mullier, F., Muraca, M., Musante, L., Mussack, V., Muth, D. C., Myburgh, K. H., Najrana, T., Nawaz, M., Nazarenko, I., Nejsum, P., Neri, C., Neri, T., Nieuwland, R., Nimrichter, L., Nolan, J. P., Nolte-'t Hoen, E. N., Noren Hooten, N., O'Driscoll, L., O'Grady, T., O'Loghlen, A., Ochiya, T., Olivier, M., Ortiz, A., Ortiz, L. A., Osteikoetxea, X., Ostergaard, O., Ostrowski, M., Park, J., Pegtel, D. M., Peinado, H., Perut, F., Pfaffl, M. W., Phinney, D. G., Pieters, B. C., Pink, R. C., Pisetsky, D. S., Pogge von Strandmann, E., Polakovicova, I., Poon, I. K., Powell, B. H., Prada, I., Pulliam, L., Quesenberry, P., Radeghieri, A., Raffai, R. L., Raimondo, S., Rak, J., Ramirez, M. I., Raposo, G., Rayyan, M. S., Regev-Rudzki, N., Ricklefs, F. L., Robbins, P. D., Roberts, D. D., Rodrigues, S. C., Rohde, E., Rome, S., Rouschop, K. M., Rughetti, A., Russell, A. E., Saa, P., Sahoo, S., Salas-Huenuleo, E., Sanchez, C., Saugstad, J. A., Saul, M. J., Schiffelers, R. M., Schneider, R., Schoyen, T. H., Scott, A., Shahaj, E., Sharma, S., Shatnyeva, O., Shekari, F., Shelke, G. V., Shetty, A. K., Shiba, K., Siljander, P. R., Silva, A. M., Skowronek, A., Snyder, O. L., 2nd, Soares, R. P., Sodar, B. W., Soekmadji, C., Sotillo, J., Stahl, P. D., Stoorvogel, W., Stott, S. L., Strasser, E. F., Swift, S., Tahara, H., Tewari, M., Timms, K., Tiwari, S., Tixeira, R., Tkach, M., Toh, W. S., Tomasini, R., Torrecilhas, A. C., Tosar, J. P., Toxavidis, V., Urbanelli, L., Vader, P., van Balkom, B. W., van der Grein, S. G., Van Deun, J., van Herwijnen, M. J., Van Keuren-Jensen, K., van Niel, G., van Royen, M. E., van Wijnen, A. J., Vasconcelos, M. H., Vechetti, I. J., Jr., Veit, T. D., Vella, L. J., Velot, E., Verweij, F. J., Vestad, B., Vinas, J. L., Visnovitz, T., Vukman, K. V., Wahlgren, J., Watson, D. C., Wauben, M. H., Weaver, A., Webber, J. P., Weber, V., Wehman, A. M., Weiss, D. J., Welsh, J. A., Wendt, S., Wheelock, A. M., Wiener, Z., Witte, L., Wolfram, J., Xagorari, A., Xander, P., Xu, J., Yan, X., Yanez-Mo, M., Yin, H., Yuana, Y., Zappulli, V., Zarubova, J., Zekas, V., Zhang, J. Y., Zhao, Z., Zheng, L., Zheutlin, A. R., Zickler, A. M., Zimmermann, P., Zivkovic, A. M., Zocco, D. and Zuba-Surma, E. K. (2018). Minimal information for studies of extracellular vesicles 2018 (MISEV2018): a position statement of the International Society for Extracellular Vesicles and update of the MISEV2014 guidelines. J Extracell Vesicles 7(1): 1535750.
- Ward, B. R., Goddard, S. J., Augustin, M.-A. and McKinnon, I. R. (1997). EDTA-induced dissociation of casein micelles and its effect on foaming properties of milk. J Dairy Res 64(4): 495-504.
- Wolf, T., Baier, S. R. and Zempleni, J. (2015). The intestinal transport of bovine milk exosomes is mediated by endocytosis in human colon carcinoma Caco-2 cells and rat small intestinal IEC-6 Cells. J Nutr 145(10): 2201-2206.
- Zonneveld, M. I., Brisson, A. R., van Herwijnen, M. J. C., Tan, S., van de Lest, C. H. A., Redegeld, F. A., Garssen, J., Wauben, M. H. M. and Nolte-'t Hoen, E. N. M. (2014). Recovery of extracellular vesicles from human breast milk is influenced by sample collection and vesicle isolation procedures. J Extracell Vesicles 3.
Article Information
Copyright
© 2020 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Benmoussa, A., Michel, S., Gilbert, C. and Provost, P. (2020). Isolating Multiple Extracellular Vesicles Subsets, Including Exosomes and Membrane Vesicles, from Bovine Milk Using Sodium Citrate and Differential Ultracentrifugation. Bio-protocol 10(11): e3636. DOI: 10.21769/BioProtoc.3636.
Category
Biochemistry > Protein > Isolation and purification
Cell Biology > Organelle isolation > Extracellular vesicle
Cell Biology > Organelle isolation > Exosomes
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










