- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
In vivo Quantification of Alkanes in Escherichia coli
Published: Vol 10, Iss 8, Apr 20, 2020 DOI: 10.21769/BioProtoc.3593 Views: 4818
Reviewed by: Juan Facundo Rodriguez AyalaJose Antonio Reyes-DariasAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Prokaryotic Expression and Purification of the hSox2-HMG Domain
Lijie Yang [...] Jingjun Hong
Aug 20, 2025 2472 Views
An Optimized Enzyme-Coupled Spectrophotometric Method for Measuring Pyruvate Kinase Kinetics
Saurabh Upadhyay
Aug 20, 2025 2524 Views
On-Column Dual-Gradient Refolding for Efficient Recovery of Insoluble Affinity-Tagged Recombinant Proteins
Anna Vlaskina [...] Maxim Patrushev
Feb 5, 2026 211 Views
Abstract
Microbial production of alkanes employing synthetic biology tools has gained tremendous attention owing to the high energy density and similarity of alkanes to existing petroleum fuels. One of the most commonly studied pathways includes the production of alkanes by AAR (acyl-ACP (acyl carrier protein) reductase)-ADO (aldehyde deformylating oxygenase) pathway. Here, the intermediates of fatty acid synthesis pathway are used as substrate by the AAR enzyme to make fatty aldehyde, which is then deformylated by ADO to make linear chain alkane. However, the variation in substrate availability to the first enzyme of the pathway, i.e., AAR, via fatty acid synthesis pathway and low turnover of the ADO enzyme make calculation of yields and titers under in vivo conditions extremely difficult. In vivo assay employing external addition of defined substrates for ADO enzyme into the medium helps to monitor the influx of substrate hence providing a more accurate measurement of the product yields. In this protocol, we include a detailed guide for implementing the in vivo assay for monitoring alkane production in E. coli.
Background
Research on alkane production using engineered microbes has gained significant popularity as it provides an attractive alternative to reduce dependence on fossil fuels while mitigating the climate change effects (Lee et al., 2008; Knothe, 2010; Lu, 2010; Schirmer et al., 2010; Tan et al., 2011). Various pathways have been uncovered or artificially assembled for the production of alka(e)nes in microbes (Schirmer et al., 2010; Mendez-Perez et al., 2011; Rude et al., 2011; Akhtar et al., 2013; Howard et al., 2013; Rui et al., 2014). However the highest reported titres for alkane production so far have been in E. coli using the AAR-ADO pathway (Figure 1) (Fatma et al., 2018). AAR catalyzes the reduction of fatty acyl-ACP or fatty acyl-CoA into fatty aldehydes using NADPH which is subsequently converted into alkanes by ADO (Marsh et al., 2013). E. coli is the most widely used host for heterologous production of biofuel candidates due to a broader knowledge of its cellular metabolic network as compared to other hosts. Heterologous co-expression of cyanobacterial AAR and ADO in E. coli results in production and secretion of alkanes (Schirmer et al., 2010). However, quantification of alkanes by the ADO enzyme and determination of the in vivo efficacy of the ADO enzyme cannot be determined due to the variation in the substrate availability by the first enzyme AAR of the pathway. Reports on differences in the solubility of AAR and hence its activity (Kudo et al., 2016) indicate that using AAR as a source of substrate for measuring in vivo activity of ADO is not a suitable approach. Hence, we developed an in vivo enzyme assay, which involves addition of the substrate aldehyde exogenously to the medium that is taken up by the growing cells and gets converted to alkane by the heterologously expressed intracellular ADO enzyme, thus giving a more reliable and accurate measurement of its in vivo efficacy.
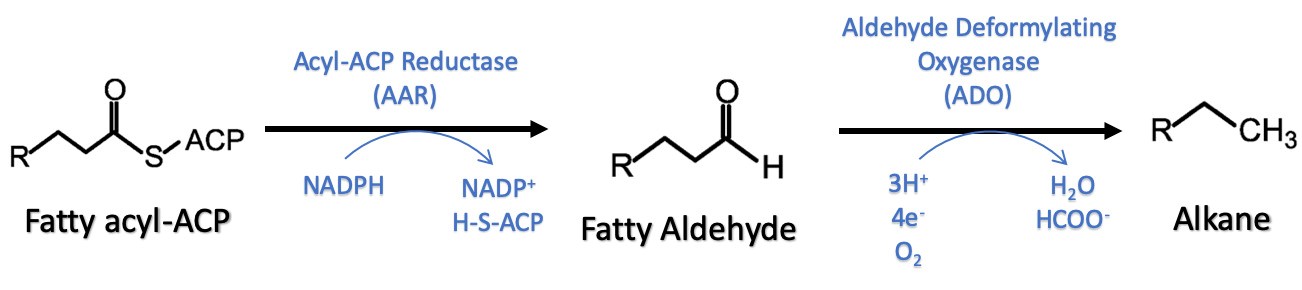
Figure 1. Schematic representation of AAR-ADO pathway for alkane production
Materials and Reagents
- 2 ml micro-centrifuge tubes (Tarson, catalog number: 500020 )
- Culture tubes (55 ml, Borosil, catalog number: 9820U08 )
- GC vials (Agilent, catalog number: 5190-2280 )
- Parafilm (Sigma, catalog number: P7793 )
- 0.2 μm filter (mdi Membrane Technologies, catalog number: SYNN0301MNXX104 )
- E. coli DH5α strain overexpressing ADO enzyme cloned in pQE30 vector (strain source: Invitrogen, vector source: Qiagen)
- Hexadecanal (TCI America, catalog number: H1296 )
- Octadecene (TCI America, catalog number: S0348 )
- Sodium phosphate dibasic (Na2HPO4) (Merck, catalog number: 255793 )
- Potassium dihydrogen phosphate (KH2PO4) (Sigma, catalog number: P5655 )
- Sodium chloride (NaCl) (Sigma, catalog number: S9888 )
- Ammonium chloride (NH4Cl) (Sigma, catalog number: 213330 )
- Magnesium sulphate heptahydrate (MgSO4·7H2O) (Sigma, catalog number: 230391 )
- Calcium chloride (CaCl2) (Sigma, catalog number: C4901 )
- Ferric chloride hexahydrate (FeCl3·6H2O) (Sigma, catalog number: 236489 )
- Zinc chloride (ZnCl2) (Sigma, catalog number: 746355 )
- Sodium molybdate dihydrate (Na2MoO4·2H2O) (Sigma, catalog number: 331058 )
- Copper sulphate (CuSO4) (Sigma, catalog number: C1297 )
- Boric acid (H3BO3) (Sigma, catalog number: B7901 )
- Thiamine hydrochloride (Sigma, catalog number: T4625 )
- Glucose (Sigma, catalog number: G8270 )
- Bis Tris (Amresco, catalog number: 0715-250G )
- Triton X-100 (Amresco, catalog number: M143-1L )
- Ethyl acetate (Merck, catalog number: 1096232500 )
- IPTG (Sigma, catalog number: I6758 )
- LB broth (HiMedia, catalog number: M1245 )
- Ampicillin (Sigma, catalog number: A9393 )
- M9 salts (see Recipes)
- M9 modified medium (see Recipes)
Equipment
- Table top centrifuge (Eppendorf, models: 5418R and 5810R)
- Incubator shaker (Kuhner, model: ISF-1 )
- GC FID (Agilent, model: 7890 A System , equipped with HP-5 column with catalog no. 19091J-413 )
- Spectrophotometer (GE Healthcare, model: UltrospecTM 2100)
- Vortex (Sigma, catalog number: Z258423-1EA )
- Autoclave (Natsteel, model: 24 SR )
Procedure
- In vivo culture
- Inoculate E. coli strains harbouring the plasmids carrying ADO gene (Figure 2) from plates streaked and incubated overnight at 37 °C (plates originally streaked from glycerol stocks) in 5 ml LB broth medium with 5 μl of appropriate antibiotics (ampicillin–100 μg/ml used in this study) in 55 ml culture tubes (1 ml of Ampicillin stock of 100 mg/ml was prepared by adding 100 mg of Ampicillin powder in 1 ml of water followed by filter sterilization).
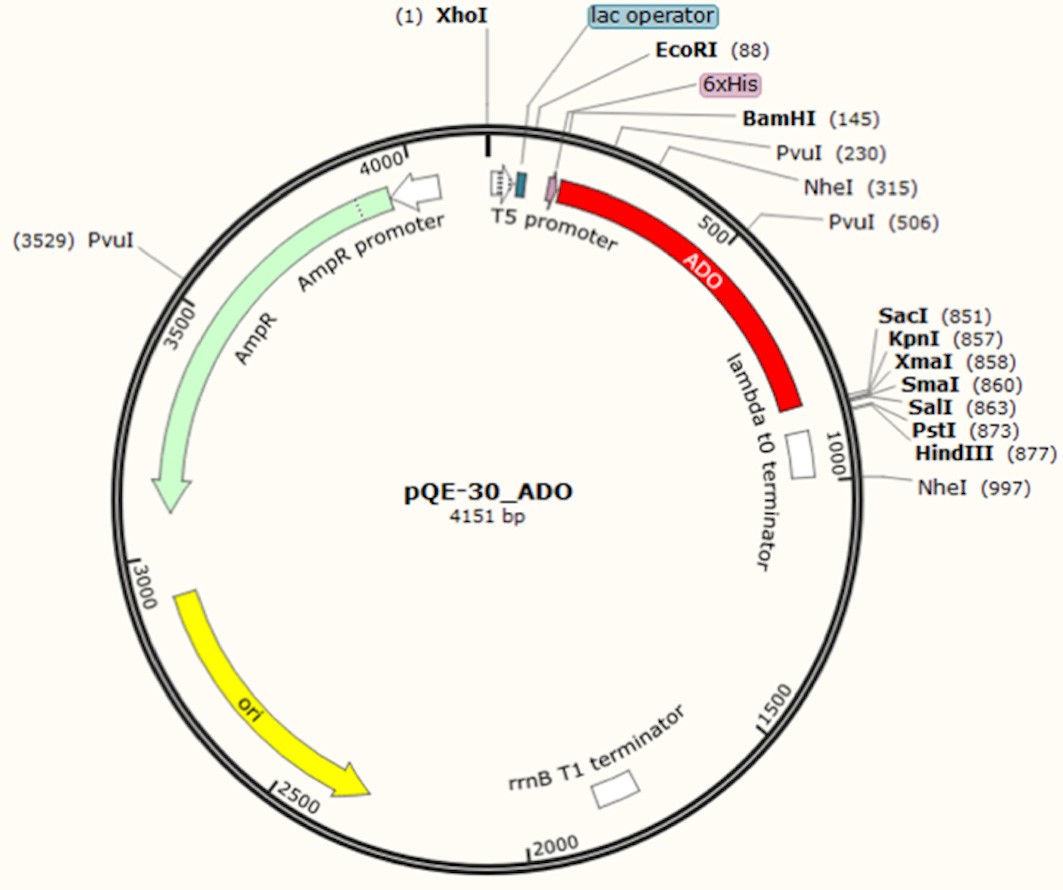
Figure 2. Schematic representation of the plasmid involved in alkane pathway construction. Plasmid vector map of pQE_ADO, carrying ado gene in pQE30 plasmid at the restriction sites BamHI/SacI. - Incubate the samples at 37 °C overnight in an incubator shaker set at 180 rpm.
- Use 50 μl (OD600 ranging from 3-4) of the primary inoculum grown overnight to inoculate 3 ml secondary culture having M9 modified medium (Recipe 1) supplemented with 2% glucose, 1 mg/L thiamine and 100 μg/ml ampicillin.
- Induce the culture with 0.01 mM IPTG (3 μl of 10 mM IPTG to 3 ml culture) at the time of secondary inoculation in M9 medium as the plasmid pQE30 carrying ADO gene is inducible by IPTG.
- Add 100 mg/L hexadecanal (dissolved in absolute ethanol) exogenously to the culture medium at the time of secondary culture inoculation and seal the culture tubes with parafilm to prevent any loss of substrate or product due to evaporation (15 μl of 20 mg/ml stock added to 3 ml culture).
- Incubate the culture tubes at 30 °C (or higher to test for thermostable enzymes) in an incubator shaker set to 120 rpm for 48 h.
- Inoculate E. coli strains harbouring the plasmids carrying ADO gene (Figure 2) from plates streaked and incubated overnight at 37 °C (plates originally streaked from glycerol stocks) in 5 ml LB broth medium with 5 μl of appropriate antibiotics (ampicillin–100 μg/ml used in this study) in 55 ml culture tubes (1 ml of Ampicillin stock of 100 mg/ml was prepared by adding 100 mg of Ampicillin powder in 1 ml of water followed by filter sterilization).
- Hydrocarbon extraction
- Measure the optical density at 600 nm of the samples to estimate the growth before proceeding with extraction of alkanes.
- Take equal volumes (0.75 ml) of sample and ethyl acetate (containing 10 mg/L Octadecene added as internal standard–from a 100 ml stock of ethyl acetate containing 1.265 μl of 99% 1-Octadecene) in a 2 ml microcentrifuge tube.
- Vortex the samples for 20 min and centrifuge at 15,700 x g for 3 min at room temperature.
- Collect the upper fraction and aliquot it into a fresh GC (gas chromatography) vial.
- Analyze the samples using gas chromatography (GC).
- Hydrocarbon analysis
- Set the hydrocarbon analysis method on the gas chromatography (GC) system equipped with HP-5 column of 30 m length, 0.32 mm internal diameter and 0.25 mm film thickness. The FID detector is used for analysis. The hydrocarbon analysis method is to be set as follows. The oven temperature programme is set as: initial 100 °C held for 3 min, increase temperature to 250 °C at the rate of 10 °C min-1 and held at 250 °C for additional 10 min. The total run-time of the programme will be 28 min. The inlet and detector temperature are to be maintained at 150 °C and 280 °C, respectively.
- Run pentadecane, hexadecanal, 1-octadecene, and hexadecanol standards of known quantity (Figure 3) prior to sample analysis and use the area under curve to calculate the concentration of metabolites in the test samples.
- Run the test samples in GC machine (Figure 4) and quantify the pentadecane content with respect to its standard.
Data analysis
Representative data
For detailed data analysis, please refer to the article of Shakeel et al. (2018). The representative GC chromatograms for all the standards are shown in Figure 3. The area under curve of known quantities of these standards are used to quantify metabolites in the test samples. The representative GC chromatograms for the typical in vivo samples containing alkane are shown in Figure 4. The area under curve of each metabolites is used to calculate their concentrations by comparing it with the area under curve of corresponding standards of known quantities.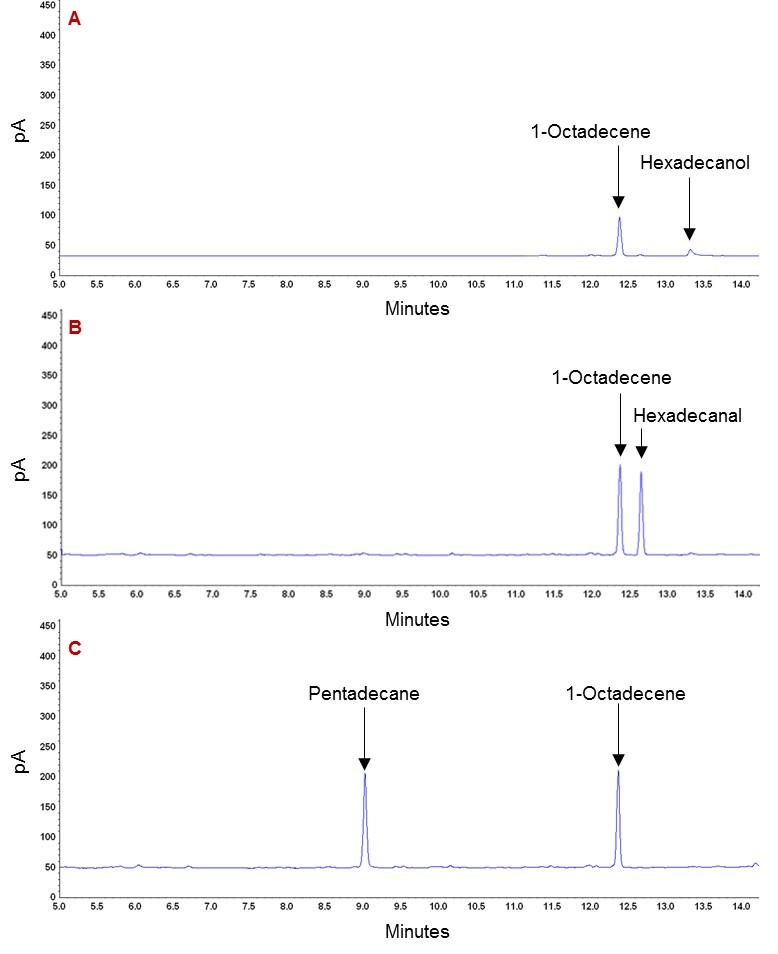
Figure 3. GC chromatograms of the standard metabolites. A. Chromatogram of Hexadecanol and 1-Octadecene. B. Chromatogram of Hexadecanal and 1-Octadecene. C. Chromatogram of Pentadecane and 1-Octadecene. The 1-Octadecene was used as an internal standard for checking extraction efficiency, while other metabolites are either substrate (Hexadecanal) or products (Hexadecanol and Pentadecane) observed after the hydrocarbon extraction.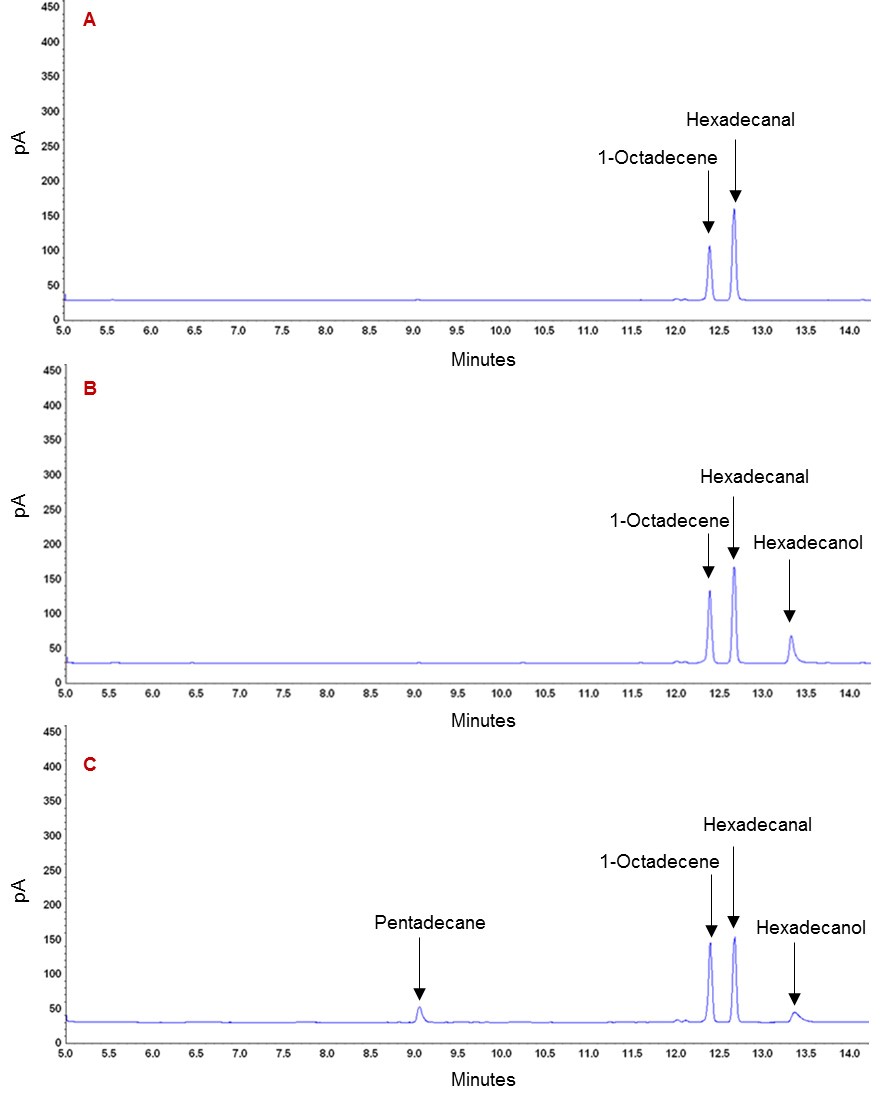
Figure 4. Functionality of ADO protein in recombinant E. coli after addition of exogenous precursor i.e. hexadecanal. A. GC chromatogram of control sample containing hexadecanal and internal standard. B. GC chromatogram of E. coli expressing empty pQE30 plasmid, showing peak of hexadecanal and hexadecanol. C. GC chromatogram of E. coli expressing pQE_ADO, showing formation of end metabolites i.e., pentadecane and hexadecanol.
Recipes
- M9 modified medium
Components of M9 modified medium are prepared as follows:- M9 salts
For 100 ml M9 salts (10x)
11.32 g Na2HPO4
3 g KH2PO4
1 g NaCl
2 g NH4Cl
Volume made up to 100 ml by adding MQ water and filter sterilized using 0.2 μm filter - Trace element recipe
1x Trace element composition:
0.25 g/L MgSO4·7H2O
11 mg/L CaCl2
18.71 mg/L FeCl3·6H2O
1.311 mg/L ZnCl2·4H2O
2 mg/L Na2MoO4·2H2O
1.9 mg/L CuSO4
0.5 mg/L H3BO3 (Boric acid)
1 mg/L Thiamine
All the above components are prepared individually as 100x stock in MQ water and filter sterilized using 0.2 μm filter - 1x Additive composition
200 mM Bis Tris (5x stock prepared)
0.1% Triton X-100 (10x stock prepared)
2% Glucose (10x stock prepared)
All the above components are prepared individually as in MQ water and filter sterilized using 0.2 μm filter
The 1x constituents from (a), (b) and (c) are mixed sequentially and volume is made up to 100 ml with MQ water autoclaved at 121 °C for 20 min. - M9 salts
Acknowledgments
The research was funded by the grants from the Department of Biotechnology, Government of India to SSY with Grant no. BT/PB/Center/03/2011 and BT/IN/INDO-UK/SuBB/21/SSY/2013. The protocol was adapted from the Fatma et al., 2016 where the method was used to screen the E. coli endogenous aldehyde reductase responsible for the conversion of long chain aldehyde to alcohol, and from the Shakeel et al., 2018 where the method was used to screen the thermostable ADO.
Competing interests
The authors declare that they have no conflicts of interest with the contents of this article.
References
- Akhtar, M.K., Turner, N.J. and Jones, P.R. (2013). Carboxylic acid reductase is a versatile enzyme for the conversion of fatty acids into fuels and chemical commodities. Proc Natl Acad Sci 110(1): 87-92.
- Fatma, Z., Jawed, K., Mattam, A. J. and Yazdani, S. S. (2016). Identification of long chain specific aldehyde reductase and its use in enhanced fatty alcohol production in E. coli. Metab Eng 37: 35-45.
- Fatma, Z., Hartman, H., Poolman, M. G., Fell, D. A., Srivastava, S., Shakeel, T. and Yazdani, S. S. (2018). Model-assisted metabolic engineering of Escherichia coli for long chain alkane and alcohol production. Metab Eng 46: 1-12.
- Howard, T. P., Middelhaufe, S., Moore, K., Edner, C., Kolak, D. M., Taylor, G. N., Parker, D. A., Lee, R., Smirnoff, N., Aves, S. J. and Love, J. (2013). Synthesis of customized petroleum-replica fuel molecules by targeted modification of free fatty acid pools in Escherichia coli. Proc Natl Acad Sci U S A 110(19): 7636-7641.
- Knothe, G. (2010). Biodiesel and renewable diesel: a comparison. Progress in Energy and Combustion. Science 36: 364-373.
- Kudo, H., Nawa, R., Hayashi, Y. and Arai, M. (2016). Comparison of aldehyde-producing activities of cyanobacterial acyl-(acyl carrier protein) reductases. Biotechnol Biofuels 9: 234.
- Lee, S. K., Chou, H., Ham, T. S., Lee, T. S. and Keasling, J. D. (2008). Metabolic engineering of microorganisms for biofuels production: from bugs to synthetic biology to fuels. Curr opin Biotech. 19: 556-563.
- Lu, X. (2010). A perspective: photosynthetic production of fatty acid-based biofuels in genetically engineered cyanobacteria. Biotechnol Adv 28: 742-746.
- Marsh, E. N. and Waugh, M. W. (2013). Aldehyde decarbonylases: enigmatic enzymes of hydrocarbon biosynthesis. ACS Catal 3(11). Doi: 10.1021/cs400637t.
- Mendez-Perez, D., Begemann, M. B. and Pfleger, B. F. (2011). Modular synthase-encoding gene involved in α-olefin biosynthesis in Synechococcus sp. strain PCC 7002. Appl Environ Microbiol 77(12):4264-4267.
- Rude, M.A., Baron, T.S., Brubaker, S., Alibhai, M., Del Cardayre, S.B. and Schirmer, A. (2011). Terminal olefin (1-alkene) biosynthesis by a novel P450 fatty acid decarboxylase from Jeotgalicoccus species. Appl Environ Microbiol 77(5): 1718-1727.
- Rui, Z., Li, X., Zhu, X., Liu, J., Domigan, B., Barr, I., Cate, J. H. and Zhang, W. (2014). Microbial biosynthesis of medium-chain 1-alkenes by a nonheme iron oxidase. Proc Natl Acad Sci U S A 111(51): 18237-18242.
- Schirmer, A., Rude, M. A., Li, X., Popova, E. and Del Cardayre, S. B. (2010). Microbial biosynthesis of alkanes. Science 329: 559-562.
- Shakeel, T., Gupta, M., Fatma, Z., Kumar, R., Kumar, R., Singh, R., Sharma, M., Jade, D., Gupta, D., Fatma, T., Yazdani, S. S. (2018). A consensus-guided approach yields a heat-stable alkane-producing enzyme and identifies residues promoting thermostability. J Biol Chem 293(24): 9148-9161.
- Tan, X., Yao, L., Gao, Q., Wang, W., Qi, F. and Lu, X. (2011). Photosynthesis driven conversion of carbon dioxide to fatty alcohols and hydrocarbons in cyanobacteria. Metab Eng 13: 169-176.
Article Information
Copyright
© 2020 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Shakeel, T., Fatma, Z. and Yazdani, S. S. (2020). In vivo Quantification of Alkanes in Escherichia coli. Bio-protocol 10(8): e3593. DOI: 10.21769/BioProtoc.3593.
- Shakeel, T., Gupta, M., Fatma, Z., Kumar, R., Kumar, R., Singh, R., Sharma, M., Jade, D., Gupta, D., Fatma, T., Yazdani, S. S. (2018). A consensus-guided approach yields a heat-stable alkane-producing enzyme and identifies residues promoting thermostability. J Biol Chem 293(24): 9148-9161.
Category
Microbiology > Heterologous expression system > Escherichia coli
Biochemistry > Other compound > Alkane
Biochemistry > Protein > Activity
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link