- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
MAMP-triggered Medium Alkalinization of Plant Cell Cultures
Published: Vol 10, Iss 8, Apr 20, 2020 DOI: 10.21769/BioProtoc.3588 Views: 4994
Reviewed by: Zhibing LaiYingnan HouShweta Panchal

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Measurement of H+ Flux in Rice by Non-invasive Micro-test Technology
Xi Zhang [...] Li-Ping Yin
Dec 5, 2015 10439 Views

Quantitative Estimation of Auxin, Siderophore, and Hydrogen Cyanide Production in Halo and Drought-Tolerant Bacterial Isolates for Cucumber Growth
Zeinab Fotoohiyan and Ali Salehi Sardoei
Oct 5, 2025 1382 Views
Reproducible Emu-Based Workflow for High-Fidelity Soil and Plant Microbiome Profiling on HPC Clusters
Henrique M. Dias [...] Christopher Graham
Jan 20, 2026 501 Views
Abstract
Plants recognize a wide variety of microbial molecules to detect and respond to potential invaders. Recognition of Microbe-Associated Molecular Patterns (MAMPs) by cell surface receptors initiate a cascade of biochemical responses that include, among others, ion fluxes across the plasma membrane. A consequence of such event is a decrease in the concentration of extracellular H+ ions, which can be experimentally detected in plant cell suspensions as a shift in the pH of the medium. Thus, similarly to reactive oxygen species (ROS) accumulation, phosphorylation of MAP kinases and induction of defense-related genes, MAMP-induced medium alkalinization can be used as a proxy for the activation of plant immune responses. Here, we describe a detailed protocol for the measurement of medium alkalinization of tobacco BY-2 cell suspensions upon treatment with two different MAMPs: chitohexamers derived from fungal cell walls (NAG6; N-acetylglucosamine) and the flagellin epitope flg22, found in the bacterial flagellum. This method provides a reliable and fast platform to access MAMP-Triggered Immunity (MTI) in tobacco cell suspensions and can be easily adapted to other plant species as well as to other MAMPs.
Keywords: Plant immunityBackground
Throughout evolution, plants developed the ability to detect microbe-derived molecules and mount immune responses that seize detrimental interactions (Boutrot and Zipfel, 2017). The microbial-associated molecular patterns (MAMPs) that induce such immune responses are often broadly conserved structural components of microbes, such as chitin from fungi and flagellin from bacteria (Cook et al., 2015). Recognition of MAMPs by extracellular plant receptors leads to MAMP-triggered immunity (MTI), which contributes to halt microbial invaders and maintain plant health (Böhm et al., 2014).
At the molecular level, MAMP perception is followed by an orchestrated set of biochemical events. Firstly, within seconds to few minutes, influx of Ca2+ and H+ ions occurs, leading to membrane depolarization and extracellular alkalinization (Ranf et al., 2011). Subsequently, production of reactive oxygen species (ROS) is observed, which may locally block pathogen growth and mediate downstream signaling (Bigeard et al., 2015). Phosphorylation of MAP kinases occurs within minutes after MAMP perception and promotes transcriptional activation of defense-related genes, many of which encode antimicrobial proteins or enzymes involved in the synthesis of hormones and secondary metabolites (Bigeard et al., 2015). Lastly, accumulation of antimicrobial compounds and callose deposition are observed, further hampering microbial growth and spread.
Mechanistic understanding of the plant immune system and its activity in plant-microbe interactions has advanced through methods that monitor many of the biochemical events underlying MTI. The most widely used experimental approaches are based on detection of ROS burst, MAP kinase activation, induction of defense-related marker genes and medium alkalinization of cell suspensions (Flury et al., 2013; Bisceglia et al., 2015; Liu et al., 2018). ROS measurement is typically performed by means of luminol-based chemiluminesce assays. MAP kinase activation is detected by western blots using specific antibodies. Transcriptional reprogramming is usually accessed by monitoring the expression levels of one or a few marker genes using real-time PCR. Medium alkalinization assays are based on recording the pH changes in MAMP-treated plant cell suspensions over time using a pH sensor for small volumes. Compared to the other methods, medium alkalinization assays are less laborious and require low cost laboratory equipment. Both model plant species and crops have been studied using medium alkalinization assays (Nühse et al., 2000; Moroz et al., 2017).
Materials and Reagents
- 12-well cell culture plate (Greiner Bio-One, CELLSTAR®, catalog number: 665180 )
- Serological pipette without tip 10 ml (SARSTEDT, catalog number: 86.1688.010 )
- P200 pipette tips
- 20 ml syringe (VWR, catalog number: 613-2046 )
- 0.22 µm syringe filter (GE Healthcare, WhatmanTM Uniflo, catalog number: 9915-2502 )
- Hexa-N-acetylchitohexaose (two manufacturers are recommended: Santa Cruz Biotechnology, catalog number: sc-222018 ; Isosep, catalog number: 56/11-0010 ). Store at -20 °C
Note: Chitoheptaose (NAG7) and chitooctaose (NAG8) or even chitin (Sigma, catalog number: C7170) may be used instead of chitohexaose. - Flagellin 22 peptide (flg22: QRLSTGSRINSAKDDAAGLQIA) (Genscript, catalog number: RP19986 ). Store at -20 °C
- Gamborg’s vitamin Solution 1,000x (sterile) (Sigma-Aldrich, catalog number: G1019-50ML ). Store at 4 °C
Note: Before using, inspect for the presence of precipitates. If present, warm the solution up to 42 °C and vortex until all precipitates dissolve. - Murashige and Skoog Basal salt mixture (PhytoTechnology Laboratories, catalog number: M524 ). Store at 4 °C
- Sucrose (Sigma-Aldrich, catalog number: S5391 )
- Myo-inositol (Sigma-Aldrich, catalog number: I7508 )
- Thiamine hydrochloride (Sigma-Aldrich, catalog number: T1270 ). Store at 4 °C
- KH2PO4 (Sigma-Aldrich, catalog number: P5655 )
- Tryptone (Oxoid, catalog number: LP0042 )
- 2,4-Dichlorophenoxyacetic acid (Sigma-Aldrich, catalog number: D7299 )
- 2-(N-Morpholino)ethanosulfonic acid (MES) hydrate (Sigma-Aldrich, catalog number: M2933 )
- Kinetin (Sigma-Aldrich, catalog number: K0753 )
- Chitohexamers (NAG6) solution (see Recipes)
- Flg22 solution (see Recipes)
- Kinetin stock solution (1,000x) (see Recipes)
- 2,4-Dichlorophenoxyacetic acid (2,4-D) stock solution (1,000x) (see Recipes)
- KH2PO4 stock solution (see Recipes)
- Thiamine/myo-inositol stock solution (see Recipes)
- Tryptone stock solution (10%) (see Recipes)
- MES/Phosphate-buffered culture medium (see Recipes)
- Subculturing medium (see Recipes)
Equipment
- 125 ml Erlenmeyer flasks (PyrexTM, catalog number: 15685767 )
- 250 ml Erlenmeyer flasks (PyrexTM, catalog number: 11902619 )
- P200 adjustable pipette
- Flow hood (Thermo Scientific, model: 1300 Series Class II , catalog number: 1323TS )
- Autoclave (VWR, catalog number: 481-0666 )
- pH meter (Mettler Toledo, Seven Compact S220 Basic, catalog number: 30019028 ). Alternative models may be used as long as the pH resolution is at least 0.001
- pH sensor InLab Micro for small volumes (Mettler Toledo, catalog number: 51343160 )
- Orbital shaker (LaboTech, model: RS150 )
Software
- Microsoft Excel
- EasyDirectTM pH Software (Mettler Toledo, 30323214)
Procedure
- Cell suspension preparation and maintenance
The protocol described here uses Nicotiana tabacum cultivar Bright Yellow 2 (BY-2) cell suspensions, which are established from in vitro-induced calli (Figure 1A). The BY-2 cell line was produced from tobacco seedlings by Kato et al. in 1972 (Kato et al.,1972) and has since been propagated and shared among plant scientists (Nagata et al., 1992). BY-2 cells are currently used as models in a number of experimental systems and are available from various research groups.- To generate a suspension of plant cells, transfer approximately 1 ml of calli to a sterile 125 ml Erlenmeyer flask containing 20 ml of MES/Phosphate-buffered culture medium (Recipe 8) using a sterile spatula or inoculation loop (in the flow hood). Be gentle to avoid unnecessary mechanical stress to the calli. In order to prevent contamination, work carefully and orderly at the flow hood. Incubate at 27 °C in the dark and under gentle agitation (100-120 rpm) in an orbital shaker. The calli will release small cell aggregates that will eventually result in a cell suspension.
- After 10-14 days, transfer 5 ml of the resulting cell suspension using a 10 ml tipless serological pipette to a sterile 250 ml Erlenmeyer flask containing 45 ml of subculturing medium (Recipe 9). Incubate at 27 °C in the dark and under gentle agitation (100-120 rpm) in an orbital shaker.
- After 7 days, repeat the subculturing as described in Step A2. At this point, the newly established cell suspension should be similar to the one shown in Figures 1B-1C. Inspect an aliquot of the suspension under the microscope to ensure that it is free from contaminants.
- For regular maintenance of BY-2 cell suspensions, subculture the cells weekly as described in Step A2. It is advisable to routinely keep two or three independent flasks of cell suspensions to prevent culture loss due to contamination.
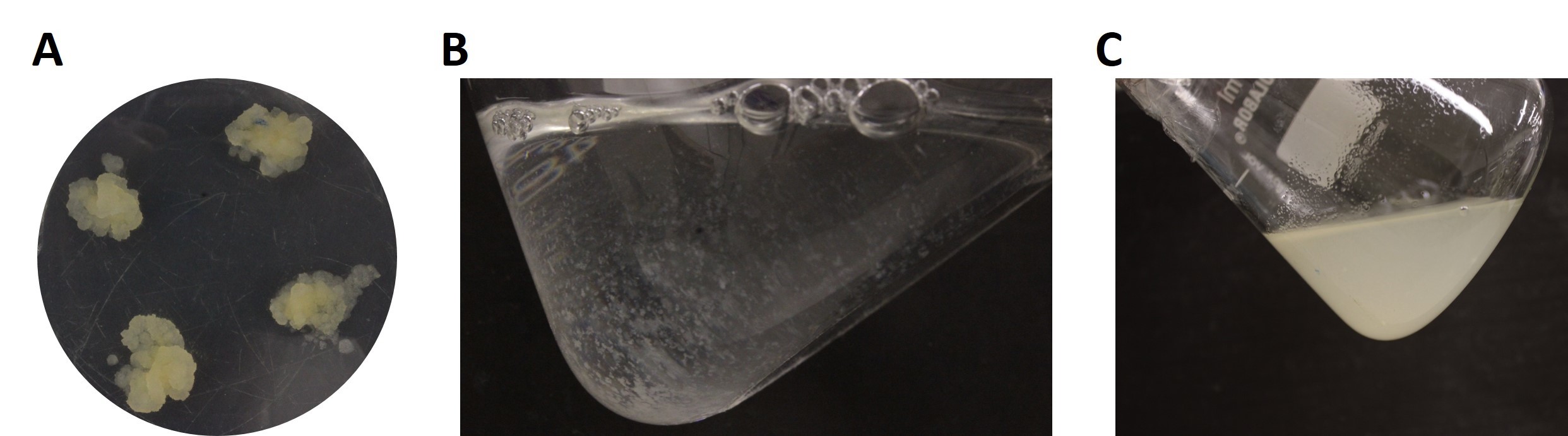

Figure 1. Nicotiana tabacum cv. Bright Yellow 2 (BY-2) cells growing as calli and cell suspensions. A. Calli growing on solid MES/Phosphate-buffered culture medium. B. Cell suspension just after subculturing. C. A 7-day-old cell suspension. Note the pale-yellow color indicating saturation.
- MAMP treatment and measurement of medium alkalinization
Once established cell suspensions are available (as described in the previous section), MAMP-triggered medium alkalinization can be accessed. Aliquots of cell suspension are individually treated with MAMPs and probed for response (Figure 2).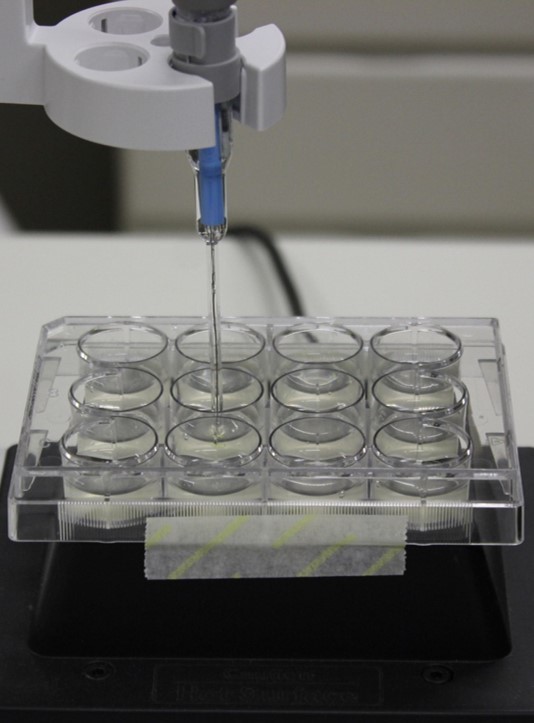
Figure 2. Experimental setup for medium alkalinization measurement. A 12-well microplate containing 2.5 ml of cell suspensions in each well is kept under gentle agitation on an orbital shaker. The pH sensor is inserted so that the probe is completely submerged and does not touch the walls.- Carefully transfer a 3- to 5-day-old cell suspension to a 12-well plate (2.5 ml per well) using a 10 ml tipless serological pipette. The cell cultures to be used should be similar to the ones shown in Figure 1C, with a pale-yellow coloration. Dark yellow or brownish colored suspensions are typically stressed and should not be used.
Note: One key factor that affects the medium alkalinization assay is the age of the cell suspension. In our experiments with BY-2 cells, we used 3- to 5-day-old suspensions. However, the optimum age may vary from lab to lab and therefore needs to be determined experimentally. - Incubate the plate for at least 2 h and up to 4 h at room temperature and under gentle agitation in an orbital shaker (typically 100-200 rpm; the speed should be just enough to keep the cells in suspension). This step allows the cells to recover from stresses caused by the transference to the plate.
- Before starting the measurements, adjust the pH meter settings appropriately: measurement resolution of 0.001, endpoint format set as manual and timed interval readings of 3 s. Calibrate the sensor according to the manufacturer’s recommendations using calibration standard solutions.
- Open the EasyDirectTM pH Software on a computer connected to the pH meter and set the export option to Excel. An Excel sheet will automatically be opened.
- Keeping the plate under agitation on the orbital shaker (100-200 rpm), insert the sensor in one well, carefully adjusting the position to ensure that the sensor is sufficiently inserted (colored tip submerged) and is not touching the walls or the bottom of the well.
- Press the Start button on the pH meter. The instrument will start recording the pH values in the Excel sheet.
- Once the measurements become stable (same value for at least 12 s), add 25 μl of 1 µM NAG6 (Recipe 1) (final concentration: 10 nM) or 25 μl of 10 µM flg22 (Recipe 2) (final concentration: 100 nM) to the well. To avoid artefactual changes in pH, equilibrate the elicitors to room temperature for 10 min before adding to the cell suspension. Make sure that each plate includes at least one well treated with elicitor solvent (H2O) as non-elicited control.
- Record the pH values for 10 min for NAG6 and 60 min for flg22.
- After the measurement is complete, remove the sensor from the well, wash thoroughly with ultrapure water and dry using tissue paper.
- Repeat Steps B5-B9 until all samples included in the experiment have been tested.
- Carefully transfer a 3- to 5-day-old cell suspension to a 12-well plate (2.5 ml per well) using a 10 ml tipless serological pipette. The cell cultures to be used should be similar to the ones shown in Figure 1C, with a pale-yellow coloration. Dark yellow or brownish colored suspensions are typically stressed and should not be used.
Data analysis
The EasyDirectTM pH Software features an export tool that records pH measurements directly into Excel sheets. Therefore, once all the raw data from an experiment is collected, the variation in pH (ΔpH) over time for each tested condition can be calculated by subtracting the pH value measured at time 0 (i.e., the pH of the medium before MAMP treatment) from the pH at each subsequent time point (i.e., the pH of the medium after MAMP treatment). Line plots depicting the variation in pH over time (Figure 3A) or bar plots showing the maximum ΔpH value (Figure 3B) for each treatment may be plotted to visualize the results. Standard deviation should be presented to indicate variation among replicates. It is desirable to have at least 3 biological replicates for each condition tested.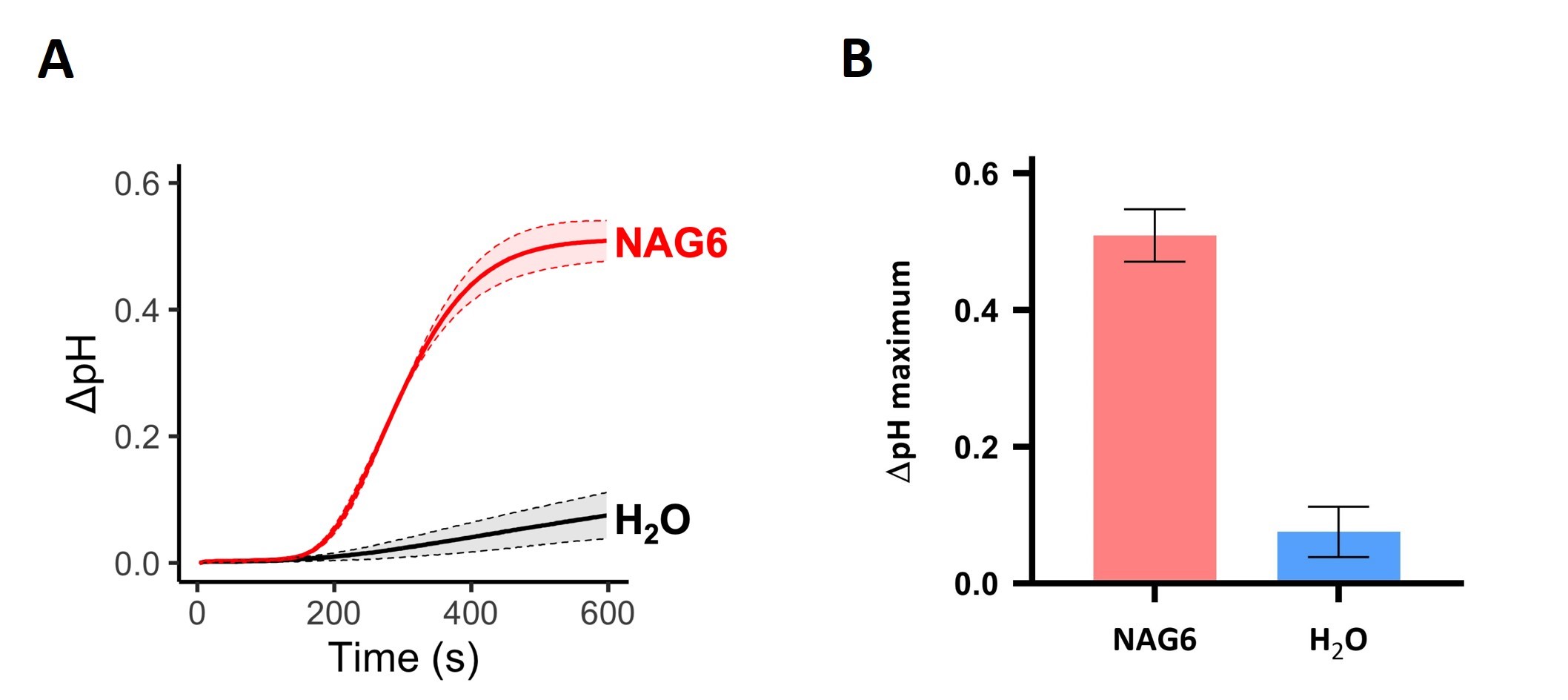
Figure 3. NAG6-induced medium alkalinization of tobacco BY-2 cell suspensions. The pH of cell suspensions treated with 10 nM NAG6 or ultrapure water (mock) was recorded for 10 min after the treatment. A. Variation in pH (ΔpH) over time (in seconds) shown in a line plot. B. Maximum variation in pH plotted as a bar graph. Shades in the line graph and error bars in the bar plot indicate standard deviation of three biological replicates.
Notes
- As an alternative to the tipless serological pipette, it is possible to cut the end of a regular serological pipette or even from a P5000 tip and use it for handling of the cell suspensions.
Recipes
- Chitohexamers (NAG6) solution
Dissolve in sterile ultrapure water to 1 µM (1.24 µg/ml)
Store at -20 °C - Flg22 solution
Dissolve in sterile ultrapure water to 10 µM (22.7 µg/ml)
Store at -20 °C - Kinetin stock solution (1,000x)
- Dissolve 0.1 mg/ml kinetin in 1/10 volume (100 µl per ml) of 1 M NaOH aqueous solution
- Once the powder is dissolved, complete the final volume with ultrapure water
- Filter sterilize and store at -20 °C
- Dissolve 0.1 mg/ml kinetin in 1/10 volume (100 µl per ml) of 1 M NaOH aqueous solution
- 2,4-Dichlorophenoxyacetic acid (2,4-D) stock solution (1,000x)
Dissolve 1 mg/ml 2,4-D in absolute ethanol
Store at -20 °C - KH2PO4 stock solution
- Dissolve 60 mg/ml KH2PO4 in deionized water
- Prepare 10 ml aliquots and store at -20 °C
- Dissolve 60 mg/ml KH2PO4 in deionized water
- Thiamine/myo-inositol stock solution
- Dissolve 0.1 mg/ml thiamine hydrochloride and 10 mg/ml myo-inositol in deionized water
- Prepare 3 ml aliquots and store at -20 °C
- Dissolve 0.1 mg/ml thiamine hydrochloride and 10 mg/ml myo-inositol in deionized water
- Tryptone stock solution (10%)
- Dissolve 10 g/L tryptone in deionized water
- Sterilize by autoclavation or filtration through a 0.22 µm syringe filter
- Store at room temperature
- Dissolve 10 g/L tryptone in deionized water
- MES/Phosphate-buffered culture medium
Composition:
0.5 g/L 2-(N-Morpholino)ethanosulfonic acid (MES) hydrate
30 g/L sucrose
1x Murashige and Skoog basal salts
100 mg/L myo-inositol
1 mg/L Thiamine hydrochloride (vitamin B1)
0.18 g/L KH2PO4
0.22 mg/L (0.1 µM) 2,4-Dichlorophenoxyacetic acid
1% tryptone
pH 5.7
Preparation (1 L):- Dissolve 4.3 g of Murashige and Skoog basal salt mixture, 30 g sucrose and 0.5 g MES hydrate in 900 ml of deionized water
- Adjust the pH to 5.7 using a 1 M KOH aqueous solution
- Add 3 ml of KH2PO4 stock solution, 10 ml of Thiamine/myo-inositol stock solution and 220 µl of 2,4-D stock solution
- Complete the final volume to 1 L with deionized water
- Transfer 18 ml aliquots to 125 ml Erlenmeyer flasks
- Autoclave at 121 °C and 15 psi for 20 min
- After equilibration at room temperature, add 2 ml of tryptone stock solution. The medium is ready to be used
- Dissolve 4.3 g of Murashige and Skoog basal salt mixture, 30 g sucrose and 0.5 g MES hydrate in 900 ml of deionized water
- Subculturing medium
Composition:
1x Murashige and Skoog basal salts
30 g/L sucrose
1x Gamborg’s vitamins
1 mg/L (4.5 µM) 2,4-Dichlorophenoxyacetic acid
0.1 mg/L (0.46 µM) kinetin
pH 5.7
Preparation (1 L):- Dissolve 4.3 g of Murashige and Skoog basal salt mixture and 30 g sucrose in 900 ml of deionized water
- Adjust the pH to 5.7 using a 1 M KOH aqueous solution
- Complete the final volume with deionized water
- Transfer 45 ml aliquots to 250 ml Erlenmeyer flasks
- Autoclave at 121 °C and 15 psi for 20 min
- After equilibration at room temperature, transfer the autoclaved flasks to a flow hood and add 45 µl of 1,000x Gamborg’s vitamin solution (sterile), 45 µl of 2,4-Dichlorophenoxyacetic acid stock solution and 45 µl of kinetin stock solution. The medium is ready to be used
- Dissolve 4.3 g of Murashige and Skoog basal salt mixture and 30 g sucrose in 900 ml of deionized water
Acknowledgments
This protocol describes in detail the procedure to access MAMP-triggered medium alkalinization of cell suspensions used in the research report by Fiorin et al. (2018). This research was supported by the São Paulo Research Foundation (FAPESP) through fellowships 09/51018-1, 09/50119-9, 11/23315-1, 13/09878-9 and 14/06181-0.
Competing interests
The authors declare no conflict of interest.
References
- Bigeard, J., Colcombet, J. and Hirt, H. (2015). Signaling mechanisms in pattern-triggered immunity (PTI). Mol Plant 8(4): 521-539.
- Bisceglia, N. G., Gravino, M. and Savatin, D. V. (2015). Luminol-based assay for detection of immunity elicitor-induced hydrogen peroxide production in Arabidopsis thaliana leaves. Bio-protocol 5(24): e1685.
- Böhm, H., Albert, I., Fan, L., Reinhard, A. and Nürnberger, T. (2014). Immune receptor complexes at the plant cell surface. Curr Opin Plant Biol 20: 47-54.
- Boutrot, F. and Zipfel, C. (2017). Function, discovery, and exploitation of plant pattern recognition receptors for broad-spectrum disease resistance. Annu Rev Phytopathol 55(1): 257-286.
- Cook, D. E., Mesarich, C. H. and Thomma, B. P. H. J. (2015). Understanding plant immunity as a surveillance system to detect invasion. Annu Rev Phytopathol 53(1): 541-563.
- Fiorin, G. L., Sanchéz-Vallet, A., Thomazella, D. P. de T., do Prado, P. F. V., do Nascimento, L. C., Figueira, A. V. and de O. Teixeira, P. J. P. L. (2018). Suppression of plant immunity by fungal chitinase-like effectors. Current Biology 28(18): 3023-3030.
- Flury, P., Klauser, D., Boller, T. and Bartels, S. (2013). MAPK phosphorylation assay with leaf disks of Arabidopsis. Bio-protocol 3(19): e929.
- Kato, K., Matsumoto, T., Koiwai, A., Mizusaki, S., Nishida, K., Noguchi, M. and Tamaki, E. (1972). Liquid suspension culture of tobacco cells. In: Ferment Technology Today. Terui, G. (Ed.) Society of Fermentation Technology, Osaka, pp 689-695.
- Liu, F., Xu, Y., Wang, Y. and Wang, Y. (2018). Real-time PCR analysis of PAMP-induced marker gene expression in Nicotiana benthamiana. Bio-protocol 8(19): e3031.
- Moroz, N., Fritch, K. R., Marcec, M. J., Tripathi, D., Smertenko, A. and Tanaka, K. (2017). Extracellular alkalinization as a defense response in potato cells. Front Plant Sci 8:32.
- Nagata, T., Nemoto, Y. and Hasezawa, S. (1992). Tobacco BY-2 cell line as the “HeLa” cell in the cell biology of higher plants. Int Rev Cytol 132: 1-30.
- Nühse, T. S., Peck, S. C., Hirt, H. and Boller, T. (2000). Microbial elicitors induce activation and dual phosphorylation of the Arabidopsis thaliana MAPK 6. J Biol Chem 275(11): 7521-7526.
- Ranf, S., Eschen-Lippold, L., Pecher, P., Lee, J. and Scheel, D. (2011). Interplay between calcium signalling and early signalling elements during defence responses to microbe- or damage-associated molecular patterns. Plant J 68(1): 100-113.
Article Information
Copyright
© 2020 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Fiorin, G. L., Sánchez-Vallet, A., Thomma, B. P. H. J., Pereira, G. A. G. and Teixeira, P. J. (2020). MAMP-triggered Medium Alkalinization of Plant Cell Cultures. Bio-protocol 10(8): e3588. DOI: 10.21769/BioProtoc.3588.
Category
Plant Science > Plant immunity > Host-microbe interactions
Plant Science > Plant biochemistry > Other compound
Cell Biology > Cell-based analysis > Ion analysis > Hydrogen
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link












