- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Image-based Quantification of Direct Cell-to-cell Transmission of Bovine Viral Diarrhea Virus
Published: Vol 9, Iss 15, Aug 5, 2019 DOI: 10.21769/BioProtoc.3319 Views: 5321
Reviewed by: Vamseedhar RayaproluVaibhav B. ShahShweta Kailasan

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
General Maintenance and Reactivation of iSLK Cell Lines
Ariana C. Calderón-Zavala [...] Ekaterina E. Heldwein
Jun 5, 2025 1876 Views
Inducible HIV-1 Reservoir Reduction Assay (HIVRRA), a Fast and Sensitive Assay to Test Cytotoxicity and Potency of Cure Strategies to Reduce the Replication-Competent HIV-1 Reservoir in Ex Vivo PBMCs
Jade Jansen [...] Neeltje A. Kootstra
Jul 20, 2025 2457 Views
Assembly and Mutagenesis of Human Coronavirus OC43 Genomes in Yeast via Transformation-Associated Recombination
Brett A. Duguay and Craig McCormick
Aug 20, 2025 3033 Views
Abstract
Different viruses rely on direct cell-to-cell transmission to propagate infection within the infected host. Measuring this mode of transmission in cultured cells is often complicated by the contribution of cell free viruses to spread, and the difficulty to distinguish between primary infected cells that produce the virus and neighboring cells that are the target of spreading. Here, we present a protocol to quantify cell-to-cell transmission of the model pestivirus bovine viral diarrhea virus that is based on the co-culture of producer cells that are infected with a reporter virus expressing mCherry and target cells that stably express GFP. Spread of cell-free viruses is blocked by the presence of a neutralizing antibody in the cell culture medium, and cell-associated transmission is unequivocally quantified by numbering cells that are positive for both GFP and mCherry using automated analysis of fluorescence microscopy images.
Keywords: PestivirusBackground
The spread of virus infections relies both on the dissemination of free virus particles and direct cell-to-cell transmission. In an infected host, cell-associated transmission overcomes multiple barriers imposed to diffusion of free viruses, such as epithelial and mucosal barriers, or antibody neutralization (Sattentau, 2008; Mothes et al., 2010). In vitro, spread of infection in the presence of virus neutralizing antibodies serves as a model to study direct cell-to-cell transmission. Virus spread from a producer cell to a target cell is usually studied in the presence of neutralizing antibodies. Different protocols have been previously devised that measure foci size in non-dividing cell cultures (Barretto and Uprichard, 2014), that use a viral infection-activated split-intein-mediated reporter system to light the expression of fluorescent reporters of contrasting colors in co-cultured producer and target cells (Zhao et al., 2018), or that require immunofluorescence staining of viral antigens in co-cultures where target cells express a fluorescent protein (Yang et al., 2015). In addition, luciferase-based assays have been developed to measure cell-associated spread of HIV in co-cultures of cells expressing an HIV reporter genome and target CD4+ T cells (Zhong et al., 2013; Agosto et al., 2014). We have recently shown that bovine viral diarrhea virus (a pestivirus in the family Flaviviridae) spreads by cell-to-cell transmission in cell culture using an approach that relies on the use of a co-culture of producer cells infected with a reporter bovine viral diarrhea virus expressing mCherry fluorescent protein, and target cells expressing GFP (Merwaiss et al., 2019). In this setup, numbering cells positive for both mCherry and GFP scores virus transmission from producer to target cells. Quantification is performed by automated image analysis of fluorescence microscopy images, and spreading is expressed as the percentage of (mCherry and GFP positive cells)/(GFP positive cells). The protocol provides an unambiguous method to quantify virus spread and has the main advantages of not requiring immunofluorescence staining, and of relying only on the use of any susceptible cell line. In addition, it can be adapted to fully automated image acquisition and analysis, live cell imaging or flow cytometry setups.
Materials and Reagents
- 24-well tissue culture plates (catalog number: BF-TCP011024)
- 35-mm tissue culture plate (catalog number: TCD010035)
- 100 mm tissue culture dish (catalog number: JBF-TCD010100)
- Glass coverslips 12 mm Ø (Marienfeld, catalog number 0111520)
- MDBK cells (ATCC, catalog number: CCL-22)
- MDBK-GFP cells
Note: MDBK cells stably expressing GFP are generated by transduction with a lentivirus carrying a GFP transgene under the control of the CMV promoter. In our hands, the transduction efficiency of MDBK cells is around 5-10% and the population is enriched in GFP expressing cells by fluorescence activated cell sorting (Merwaiss et al., 2019). - ncpBVDV/mCherry-E2 (generated using a reverse genetic system as previously described in Merwaiss et al., 2019)
- Dulbecco’s Modified Eagle Medium (DMEM) with High Glucose (Gibco, catalog number: 12800082)
- Fetal Bovine Serum (PAA, catalog number: A15-101)
- Horse Serum heat inactivated, New Zealand origin (Gibco, catalog number: 26050088)
- Penicillin, Streptomycin (10,000 U/ml) (Gibco, catalog number: 15140122)
- DPBS (Gibco, catalog number: 21600069)
- Trypsin-EDTA 10x (Gibco, catalog number: 15400-054)
- DAPI (Molecular Probes, catalog number: D1306)
- FluoroGuard antifade reagent (Bio-Rad, catalog number: 170-3140)
- Mouse anti BVDV E2 monoclonal antibody (DMAB28412; Creative Diagnostics)
- Mouse anti BVDV E2 polyclonal serum and total IgG fraction (Merwaiss et al., 2019)
- 4% solution of paraformaldehyde [PFA, (Sigma, catalog number: 158127)] in PBS (CSH Protocols, 2006)
- Complete medium (see Recipes)
- Infection medium (see Recipes)
Equipment
- 37 °C, 5% CO2 cell culture incubator (Thermo Scientific Form Series II Water Jacketed, model: 3131)
- Nikon Eclipse 80i Fluorescence microscope equipped with a DS-Qi1Mc camera
Software
- Fiji (Schindelin et al., 2012)
Procedure
- Infection of MDBK cells with non cytopathic (ncp) BVDV/mCherry-E2 and establishment of persistently infected cells
- Maintain MDBK cells in complete medium splitting 1:10 every 3 to 4 days.
- Lift cells with trypsin and seed 3 x 105 cells in a 35-mm tissue culture plate. To lift cells, wash the monolayer with PBS and add 0.3 ml of trypsin 0.05%/EDTA 0.02%, incubate for 3-5 min at 37 °C, add 1 ml complete medium, and lift cells by pipetting up and down.
- Allow cells to grow overnight at 37 °C in a 5% CO2 cell culture incubator.
- Dilute ncpBVDV/mCherry-E2 in 1 ml of infection medium.
Note: The virus stock is recovered from RNA transfection of ncpBVDV/mCherry-E2 into MDBK cells. Usually, we check the expression of mCherry 3-4 days after transfection and harvest cells and supernatant by scrapping. After three cycles of freeze-thawing, the cell lysate is clarified by centrifugation at 500 x g, and the supernatant containing the virus stock aliquoted and stored at -80 °C. To establish cells persistently infected with the ncp virus, we prepare a 1:5 dilution of the stock in Infection medium. Spinoculation (see Supplemental Protocol Procedure A) can be used to favor infection. The establishment of persistently infected cells is independent of the multiplicity of infection, since cells are passaged until infection is detected in 80-100% of the cell monolayer. - Add the 1 ml virus inoculum to MDBK cell culture and incubate for 2 h at 37 °C in a 5% CO2 cell culture incubator to allow entry.
- Remove the inoculum and add 2 ml of complete medium.
- Incubate at 37 °C in 5% CO2 cell culture incubator until cells reach 100% confluency (usually 2-3 days).
- Lift cells with trypsin and plate into a 100 mm tissue culture dish.
- Split cells every 3-4 days and check the proportion of infected cells by scoring expression of mCherry through direct visualization of the plate under an inverted fluorescence microscope. Usually after 2 or 3 passages, 80-100% of the cells should be positive for mCherry expression.
- Co-culture of persistently infected cells and MDBK-GFP cells (Figure 1) (Merwaiss et al., 2019)
- Lift cells with trypsin and count.
Note: In this step, cell washing and trypsin treatment remove any infectious extracellular virus. - Combine persistently infected and GFP cells at a 1:4 ratio in complete medium at a final density of 2 x 105 cells/ml and add virus neutralizing antibodies at the appropriate dilution. Prepare a suspension of cells without neutralizing antibodies for the control.
Note: The dilution of virus neutralizing antibodies to be used in the spreading assay is determined in a classical virus neutralization assay (Merwaiss et al., 2019). We usually use 10-fold above the 50% neutralizing dose to completely block the spread of cell free viruses. In particular, we used in our assays a 1:1000 dilution of commercial mouse anti BVDV E2 monoclonal antibody, 1:500 dilution of mouse anti BVDV E2 polyclonal serum and 1:100 dilution of total IgG fraction. - Plate 500 μl of the cell mix (approximately 105 cells per well) on 12 mm glass coverslips into 24-well plates and allow infection to proceed for 3 days in 5% CO2 cell culture incubator.
Note: Three days after infection, no free infectious virus is recovered from the supernatant of co-cultures that were incubated in the presence of virus neutralizing antibodies as tested by reinfection of a fresh monolayer of MDBK cells.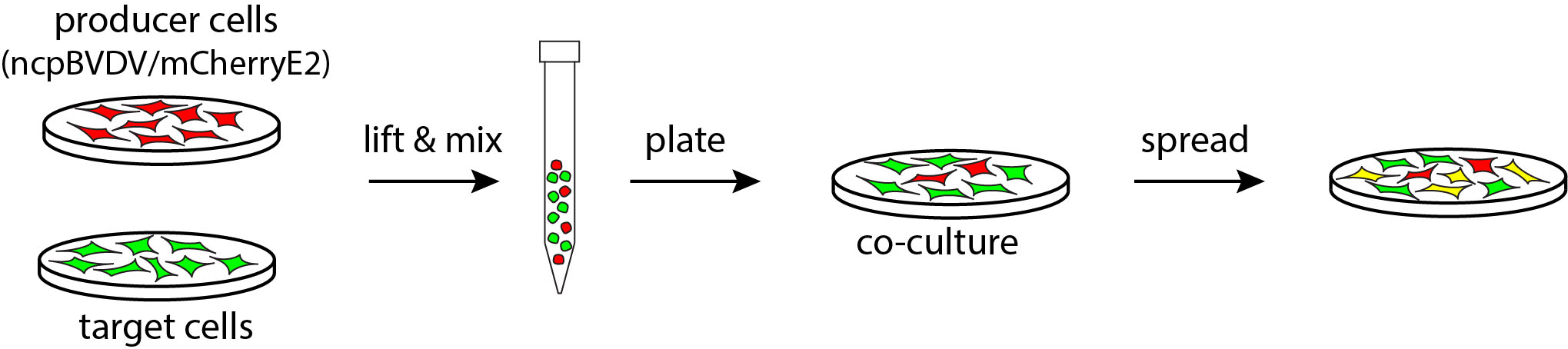
Figure 1. Schematic representation of the experimental setup used to assess cell-to-cell transmission using persistently infected producer cells. Producer cells persistently infected with ncpBVDV/mCherry-E2 (red) are co-cultured with non-infected target cells expressing GFP (green). Spreading from producer cells to target cells is unequivocally scored by the expression of both GFP and mCherry (yellow).
- Lift cells with trypsin and count.
- Visualizing BVDV cell-to-cell spread
- Wash the cells once with PBS and fix by adding 400 μl of PFA 4% in PBS. Incubate for 15-20 min at room temperature.
- Remove the PFA solution and wash twice with PBS.
- To stain cell nuclei, add 300 μl of a 1:1,000 dilution of DAPI in PBS and incubate for 10 min.
- Wash twice with PBS and mount on glass slides on a drop of Fluoroguard antifade.
- Air dry slides and capture images in a fluorescence microscope.
- Evaluate the requirement of host factors in target cells
The protocol can be used to test for the requirement of host factors for cell-to-cell transmission by using target MDBK cells stably expressing GFP fusions of dominant negative versions of proteins involved in the attachment and internalization of the virus. For example, we generated in our study MDBK cells expressing dominant negative Eps15 fused to GFP to evaluate the dependence on clathrin-mediated endocytosis for BVDV spread into target cells (Merwaiss et al., 2019). Factors with bona fide loss of function mutations such as Caveolin, Dynamin, Rab5, and Rab7 can be evaluated using this setup. This protocol is also compatible with target cells depleted of host factors.
Supplemental protocol: Quantification of cell-to-cell spread using cytopathic BVDV/mCherry-E2
Materials and Reagents
The same as in the Main Protocol with the following addition: cpBVDV/mCherry-E2 generated using a reverse genetic system as previously described (Merwaiss et al., 2019).
Equipment
The same as in the Main Protocol with the following addition: Thermo Scientific SL 16 benchtop centrifuge with M-20 microplate swinging bucket rotor.
Software
The same as in the Main Protocol.
Procedure
- Infection of MDBK cells with cytopathic BVDV (Figure 2)
- Seed 3 x 104 cells per well on 12 mm glass coverslips in 500 μl of complete medium in 24-well plates and allow to grow overnight in 5% CO2 cell culture incubator.
- On the following day, prepare a dilution of cpBVDV/mCherry-E2 to 105 TCID50/ml in infection medium.
- Wash cells and add 200 μl of virus dilution per well.
- Spinoculate the cells by centrifuging the plate at 500 x g for 10 min.
- Move the plate to 37 °C in a 5% CO2 cell culture incubator for 4 h. At this stage, it is not necessary to replace the virus containing infection medium with fresh medium.
- Co-culture with MDBK-GFP cells (Figure 2)
- Lift MDBK-GFP cells with trypsin and count.
- Prepare a suspension of cells in complete medium at a density of 2 x 105 cells/ml and add an appropriate dilution of virus neutralizing antibodies. Keep 1 ml of the suspension without neutralizing antibodies for the control.
- Thoroughly, wash infected cells with PBS and plate 500 μl of MDBK-GFP cell suspension on top of the adhered infected cells per well.
- Incubate for two days at 37 °C in 5% CO2 cell culture incubator.
Note: Due to faster replication kinetics of the cytopathic biotype of BVDV compared to the non-cytopathic biotype, cell spread is assessed 2 days after infection. Usually, extended cytopathic effect is observed in the cell monolayer 3 days in assays using cpBVDV. - Proceed to fixation and DAPI staining as described for Procedure C in the Main Protocol.
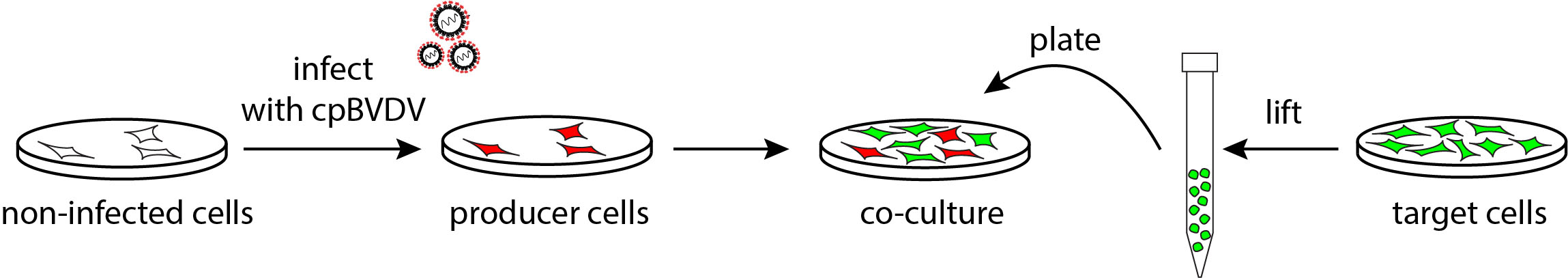
Figure 2. Schematic representation of the experimental setup used to assess cell-to-cell transmission of cpBVDV. To set up the co-culture, producer cells are first infected with cpBVDV/mCherry-E2 (red), and next non-infected target cells expressing GFP (green) are plated on to the infected cell monolayer.
Data analysis
- We generate data from three independent experiments. For each experiment, we capture ten images with a 20x objective.
- Spreading is scored for each image in Fiji using a script that performs the following steps (Figure 3):
- Manually set threshold for the red channel.
Note: Appropriate threshold setting is critical to obtain reliable results. The threshold is adjusted for each channel, and we use the same threshold across the images in the experiment. The decision on the suitability of the threshold used on a set of images is made after visual inspection of 3-4 random images for false positives. False positive cells (target cells that are not infected) should not exceed 10% of the double positive cells in a given image. A non-spreading phenotype is usually an excellent control to test if threshold settings are adequate. We allow a maximum of 3% double positive cells (false positive) for non-spreading phenotypes. For instance, in our study cell-to-cell transmission is completely blocked to non-susceptible CRIB cells or in the presence of soluble BVDV E2 envelope protein. - Create a mask.
- Process the mask with Dilate tool (Process > Binary > Dilate, iterations = 5, counts = 3).
- Save mask as “mCherry”.
- Manually set the threshold for the green channel.
- Create a mask.
- Save mask as “GFP”.
- Manually set the threshold for the blue channel.
- Create a mask.
- Process the mask with Watershed tool (Process > Binary > Watershed).
- Save mask as “DAPI”.
- Use logical operators (Process > Image Calculator) to generate a new mask for the nuclei of mCherry positive cells using mCherry AND DAPI masks as source images.
- Count mCherry positive cells using Analyze Particles tools (Analyze > Analyze Particles, size = 50-infinity).
- Use logical operators (Process > Image Calculator) to generate a new mask for the nuclei of GFP positive cells using GFP AND DAPI masks as source images.
- Count GFP positive cells using Analyze Particles tools (Analyze > Analyze Particles, size = 50-infinity).
- Use logical operators (Process > Image Calculator) to generate a result mask of infected GFP positive cells using GFP nuclei AND mCherry nuclei masks as source images.
- Count infected GFP positive cells using Analyze particles tools (Analyze > Analyze Particles, size = 50-infinity).
- Calculate the percentage of spread as (# infected GFP positive cells)/(# GFP positive cells).
- Manually set threshold for the red channel.
- We usually count a total number of 1,000 GFP positive target cells for each condition in 8 to 10 images in one experiment (see Appendix).
- The mean % of spread is calculated for each experiment, and the results are represented as the mean and the standard deviation of three independent experiments.
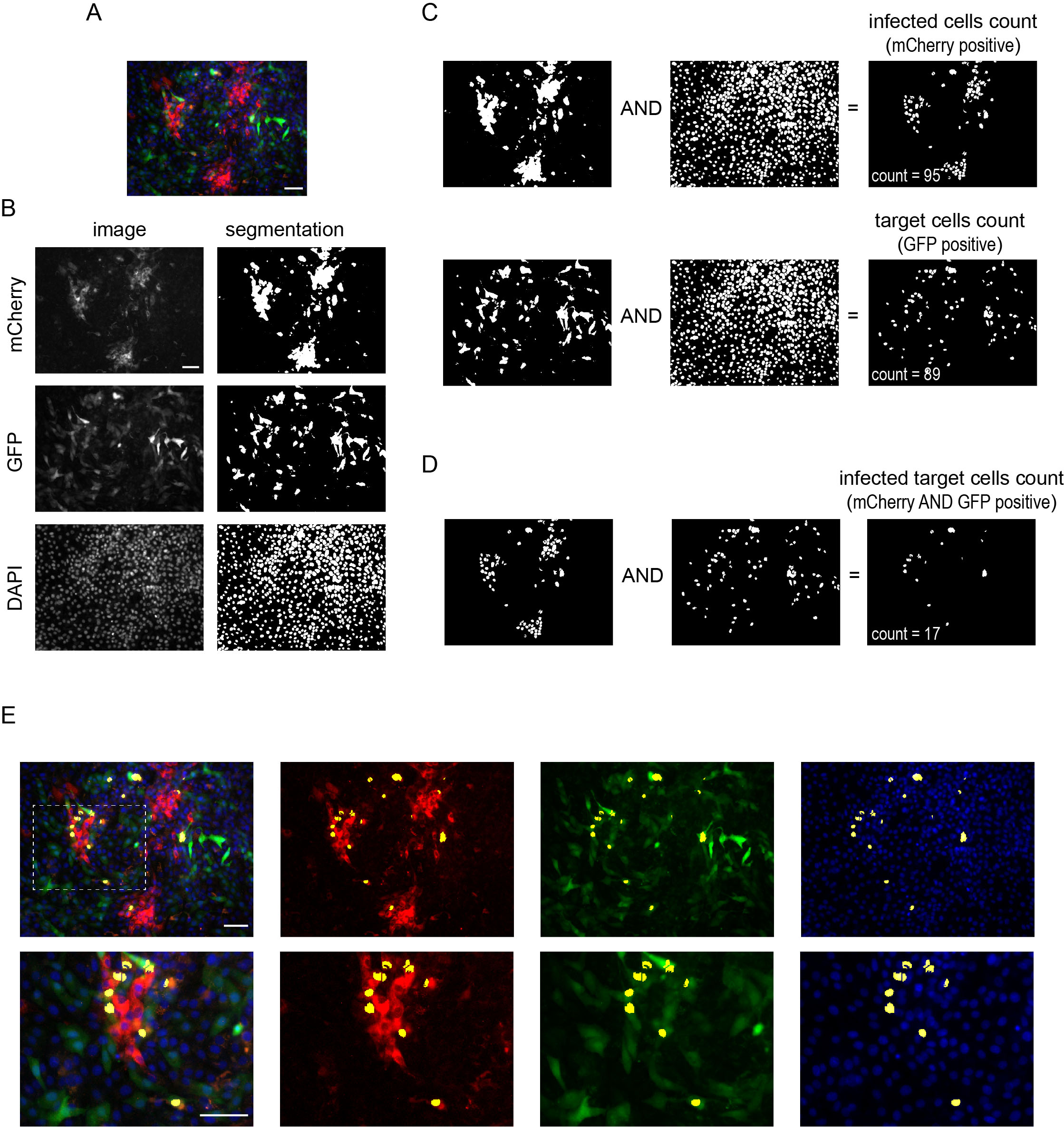
Figure 3. Automated image analysis of BVDV spread. A. Representative image of the spread of BVDV from cells persistently infected with the reporter virus expressing mCherry to target cells expressing GFP. B-D. Step-by-step image analysis. B. Left, gray scale images of mCherry (red), GFP (green), and DAPI (blue) channels. Right, image segmentation. C. Numbering mCherry and GFP positive cells using the masks generated in panel B and logical operators. D. Numbering of spreading from donor to target cells. E. Overlay of the result mask (infected target cells, yellow) with merged image in A and split channels. Bottom, close up of the boxed region in the top row. Scale bars = 20 μm.
Recipes
- Complete medium
DMEM
100 units/ml penicillin and 100 mg/ml streptomycin
10% fetal bovine serum - Infection medium
DMEM
100 units/ml penicillin and 100 mg/ml streptomycin
2% horse serum
Acknowledgments
Maria José Pascual assisted in the production of immune sera against E2. This work was supported by a National Agency for the Promotion of Science and Technology (ANPCyT) grant (PICT2014-2213) to D.E.A.
Competing interests
The authors declare no financial or non-financial competing interests.
References
- Agosto, L. M., Zhong, P., Munro, J. and Mothes, W. (2014). Highly active antiretroviral therapies are effective against HIV-1 cell-to-cell transmission. PLoS Pathog 10(2): e1003982.
- Barretto, N. and Uprichard, S. L. (2014). Hepatitis C virus cell-to-cell spread assay. Bio-protocol 4(24): e1365.
- Merwaiss, F., Czibener, C. and Alvarez, D. E. (2019). Cell-to-cell transmission is the main mechanism supporting bovine viral diarrhea virus spread in cell culture. J Virol 93(3). doi: 10.1128/JVI.01776-18.
- Mothes, W., Sherer, N. M., Jin, J. and Zhong, P. (2010). Virus cell-to-cell transmission. J Virol 84(17): 8360-8368.
- CSH Protocols. (2006). Paraformaldehyde in PBS. Cold Spring Harbor Protocols 2006(1): pdb.rec9959.
- Sattentau, Q. (2008). Avoiding the void: cell-to-cell spread of human viruses. Nat Rev Microbiol 6(11): 815-826.
- Schindelin, J., Arganda-Carreras, I., Frise, E., Kaynig, V., Longair, M., Pietzsch, T., Preibisch, S., Rueden, C., Saalfeld, S., Schmid, B., Tinevez, J. Y., White, D. J., Hartenstein, V., Eliceiri, K., Tomancak, P. and Cardona, A. (2012). Fiji: an open-source platform for biological-image analysis. Nat Methods 9(7): 676-682.
- Yang, C. F., Tu, C. H., Lo, Y. P., Cheng, C. C. and Chen, W. J. (2015). Involvement of tetraspanin C189 in cell-to-cell spreading of the dengue virus in C6/36 cells. PLOS Neglected Tropical Diseases 9(7): e0003885.
- Zhao, F., Zhao, T., Deng, L., Lv, D., Zhang, X., Pan, X., Xu, J. and Long, G. (2018). HCV reporter system (viral infection-activated split-intein-mediated reporter system) for testing virus cell-to-cell transmission ex-vivo. Bio-protocol 8(15): e2949.
- Zhong, P., Agosto, L. M., Ilinskaya, A., Dorjbal, B., Truong, R., Derse, D., Uchil, P. D., Heidecker, G. and Mothes, W. (2013). Cell-to-cell transmission can overcome multiple donor and target cell barriers imposed on cell-free HIV. PLoS One 8(1): e53138.
Article Information
Copyright
© 2019 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Merwaiss, F. and Alvarez, D. E. (2019). Image-based Quantification of Direct Cell-to-cell Transmission of Bovine Viral Diarrhea Virus. Bio-protocol 9(15): e3319. DOI: 10.21769/BioProtoc.3319.
Category
Microbiology > Microbe-host interactions > Virus
Cell Biology > Cell imaging > Fluorescence
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link








