- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Measuring Protein Half-life in Arabidopsis thaliana
Published: Vol 9, Iss 15, Aug 5, 2019 DOI: 10.21769/BioProtoc.3318 Views: 5702
Reviewed by: ANU PRIYA MINHASRunlai HangAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
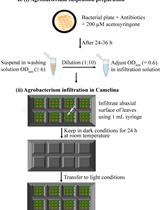
Agrobacterium-Mediated Transient Gene Expression Optimized for the Bioenergy Crop Camelina sativa
Pawan Kumar [...] Jean T. Greenberg
Apr 5, 2024 2114 Views

Identification of S-locus F-box Protein Sequences in Diploid Potato, Solanum okadae, via Degenerate PCR
Amar Hundare [...] Timothy P. Robbins
Jun 5, 2025 1994 Views

Quantitative Analysis of the Arabidopsis Leaf Secretory Proteome via TMT-Based Mass Spectrometry
Sakharam Waghmare [...] Rucha Karnik
Nov 20, 2025 2019 Views
Abstract
Post-translational modifications play important roles in controlling protein function and can lead to altered protein stability. Protein stability can be determined after treatment with the protein synthesis inhibitor Cycloheximide. Cycloheximide is a translational inhibitor that inhibits protein synthesis via cytoplasmic ribosomes. Here we describe how to measure the stability of MYC2 in the context of regulation by FERONIA receptor kinase. First, we describe how to measure MYC2 stability in wild-type and feronia mutant; then we describe similar assays in transgenic plants expressing MYC2-FLAG and MYC2A12-FLAG (12 FERONIA phosphorylation sites are mutated to Alanine and the mutant protein is stabilized). MYC2 can be induced by mechanical touch, which can be a confounding factor in protein level measurement. In this protocol, we take that into consideration and try to achieve more accurate measurement.
Keywords: FERONIABackground
We have shown that FERONIA receptor kinase can phosphorylate and destabilize MYC2, a major transcription factor in Jasmonic Acid (JA) signaling, which positively contributes to plant immunity (Guo et al., 2018). Under normal growth conditions, MYC2 is present at very low levels. Overnight treatment with MG132 allows MYC2 to accumulate to a higher level before Cycloheximide treatment (Jung et al., 2015; Jeong et al., 2017), which proves to be important for detection of such unstable proteins. Additionally, MYC2 can be induced by mechanical touch, which can lead to inconsistency in the protein level measurement. Here we provide a detailed protocol to help determine the half-life of proteins like MYC2. We illustrate how this approach can be used with both the endogenous protein and epitope-tagged proteins in transgenic plants. This protocol provides some general guidelines for determining the levels of proteins that exist at low levels and are sensitive to handling (e.g., mechanical touch).
Materials and Reagents
Note: For a complete list of reagents, please refer to Guo et al. (2018).
- Pipette tips (MidSci, catalog number: AVR96 for 200 μl tips, AVR96-12 for 1.2 ml tips)
- Micropore tape (3M, catalog number: 1530-0)
- Kimwipes (Fisher Scientific, catalog number: 06-666A)
- Plastic blue pestle (Fisher Scientific, catalog number: 12-141-368)
- 24-well culture plate (Fisher Scientific, catalog number: 07-200-686)
- Nitrocellulose membrane (Bio-Rad, catalog number: 1620112)
- Cycloheximide (Sigma-Aldrich, catalog number: C4859)
- MG132 (Sigma-Aldrich, catalog number: M7449)
- Rabbit polyclonal anti-MYC2 (Guo et al., 2018)
- Rabbit polyclonal anti-FLAG (Sigma-Aldrich, catalog number: F7425; RRID: AB_439687)
- Goat anti-rabbit IgG-HRP secondary antibody (Thermo Fisher, catalog number: 31460)
- Ponceau S (Sigma-Aldrich, catalog number: P7170-1L)
- Liquid nitrogen (Chemstore on campus)
- DMSO (Fisher Scientific, catalog number: D159-4)
- Linsmaier and Skoog packet (CaissonLabs, catalog number: LSP03-1LT)
- Sucrose (Fisher Scientific, catalog number: BP220-10)
- Tris base (Fisher Scientific, catalog number: BP152-10)
- SDS (Fisher Scientific, catalog number: BP166-5)
- Glycerol (Fisher Scientific, catalog number: BP229-1)
- Bromophenol blue (Fisher Scientific, catalog number: BP115-25)
- 2-Mercaptoethanol (Fisher Scientific, catalog number: BP176-100)
- ½ MS liquid medium (see Recipes)
- 2x SDS sample buffer (see Recipes)
Equipment
- Sterilized forceps
- Sartorius Entris Analytical Balance
- P1000 pipettor
- Black & Decker Electric drill
- Sterile culture hood
- Microcentrifuge
- Mini-Protein Tetra Cell (Bio-Rad, catalog number: 1658001EDU)
- Trans-Blot Turbo system (Bio-Rad, catalog number: 1704155)
Software
- ImageJ (https://imagej.net/Fiji), a friendly tutorial provided on the website for new users
Procedure
- For MYC2 half-life measurement in WT and fer mutant
Note: Steps A1 to A4 are carried out in a sterile culture hood.- Sterilize and germinate seeds on ½ MS plates for 7 days, vertically with constant light at 100 μmol m-2 s-1, 22 °C.
- Add 1 ml of liquid ½ MS medium to each well of a 24-well plate, 5 wells for each genotype.
- With a pair of sterilized forceps, gently transfer 50 mg of whole seedlings to the wells.
Note: The number of seedlings for 50 mg can be predetermined by weighing the seedlings. - Add 5 μl of 10 mM MG132 to each well. The final concentration of MG132 is 50 μM.
- Seal the plate with Micropore tape and incubate at room temperature with gentle rocking at 40 RPM, for 16 h.
- Gently remove the liquid from the well with a P1000 pipettor and wash the seedlings with 1 ml fresh ½ MS liquid medium. Repeat the process for a total of 5 washes.
- Gently add 1 ml fresh ½ MS liquid medium to each well, add Cycloheximide to 4 wells, and add the same volume of 100% DMSO to 1 well as control for each genotype, the final concentration of Cycloheximide is 200 μM.
- Collect the seedlings at 30-min intervals by gently picking out the seedlings with a pair of forceps. Dab dry on Kimwipes. Gently put the seedlings into a microtube and freeze in liquid nitrogen.
- For Western blotting to estimate the amount of MYC2 protein, add 150 μl of 2x SDS sample buffer to 50 mg of frozen seedlings and grind the tissue for about 30 s with a brief stop every 10 s, with a blue pestle mounted to an electric drill machine.
- Vortex and boil the ground samples for 5 min and cool down on ice for 1 min.
- Spin the cooled samples for 5 min at full speed in a microcentrifuge and load 20 μl to SDS-PAGE gel. The Mini-Protein Tetra Cell and Trans-Blot Turbo system are used for protein gel electrophoresis and transfer.
- Conduct Western blotting with proper primary and secondary antibodies to detect protein levels.
- For the MYC2-FLAG and MYC2A12-FLAG half-life in transgenic plants
- Sterilize and germinate seeds on Gentamycin containing plates (75 μg/ml) for 7 days.
- Add 1 ml fresh ½ MS liquid medium to each well of a 24-well plate, 5 wells per genotype.
- With a pair of forceps, gently transfer 50 mg of seedlings to the wells. Leave the plate for 2 h to minimize the effect of mechanical touch.
- Add 1 ml ½ MS liquid containing 400 μM Cycloheximide to 4 wells and 1 ml ½ MS liquid containing the same volume of DMSO to 1 well. Gently rock the plate a few times to mix. The final concentration of Cycloheximide is 200 μM.
- Then follow Steps A8-A12 described above.
Data analysis
After Western blotting, scan the film using a flat-bed scanner. Stain the membrane using Ponceau S and scan the stained membrane (Alternatively, this step can be done right after transfer. The membrane is then washed with distilled water before blotting). Use ImageJ to measure the intensity of the protein bands of interest and the intensity of Rubisco from the staining. Calculate the ratios of the protein of interest over the corresponding Rubisco. The protein half-life is the time when the ratio decreases by half. For an example, see Figures 3C and 3D in Guo et al. (2018). Alternatively, a housekeeping protein whose level doesn’t change dramatically due to Cycloheximide treatment can be monitored by western blotting and used as a reference instead of Rubisco.
Recipes
- ½ MS liquid medium
In 2 L of Millipore water, dissolve one Linsmaier and Skoog packet, then add 20 g of sucrose till it dissolves completely; autoclave at 121 °C and 15 psi for 15 min. Cool it down before use - 2x SDS sample buffer
To make 50 ml of 2x SDS buffer, mix 10 ml of 0.5 M Tris pH 6.8, 10 ml of 100% glycerol, 10 ml of 20% SDS, 0.02% (w/v) Bromophenol blue, and 700 μl of 2-Mercaptoethanol, bring the volume up to 50 ml with water
Acknowledgments
The authors thank Dr. Trevor Nolan for editing the manuscript. The research was supported by Signature Research Initiative of the College of Liberal Arts and Sciences and Plant Sciences Institute at Iowa State University.
Competing interests
Hongqing Guo and Yanhai Yin are co-inventors on the patent titled "Modulation of receptor-like kinases for promotion of plant growth", US9512440B2.
References
- Guo, H., Nolan, T. M., Song, G., Liu, S., Xie, Z., Chen, J., Schnable, P. S., Walley, J. W. and Yin, Y. (2018). FERONIA receptor kinase contributes to plant immunity by suppressing jasmonic acid signaling in Arabidopsis thaliana. Curr Biol 28(20): 3316-3324. e6.
- Jeong, J. S., Jung, C., Seo, J. S., Kim, J. K. and Chua, N. H. (2017). The deubiquitinating enzymes UBP12 and UBP13 positively regulate MYC2 levels in jasmonate responses. Plant Cell 29(6): 1406-1424.
- Jung, C., Zhao, P., Seo, J. S., Mitsuda, N., Deng, S. and Chua, N. H. (2015). PLANT U-BOX PROTEIN10 regulates MYC2 stability in Arabidopsis. Plant Cell 27(7): 2016-2031.
Article Information
Copyright
© 2019 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Guo, H. and Yin, Y. (2019). Measuring Protein Half-life in Arabidopsis thaliana. Bio-protocol 9(15): e3318. DOI: 10.21769/BioProtoc.3318.
Category
Plant Science > Plant molecular biology > Protein
Biochemistry > Protein > Stability
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link











