- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Candida albicans Mitochondrial Protein Import Assay
Published: Vol 3, Iss 4, Feb 20, 2013 DOI: 10.21769/BioProtoc.331 Views: 10510

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Halo Assay for Toxic Peptides and Other Compounds in Microorganisms
Houjian Cai [...] Jeffrey M. Becker
Nov 20, 2016 9296 Views
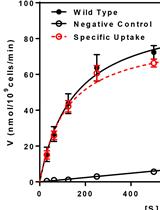
Uptake Assay for Radiolabeled Peptides in Yeast
Melinda Hauser [...] Jeffrey M. Becker
Nov 20, 2016 9222 Views
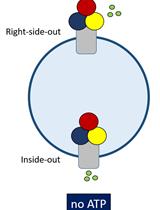
Polyamine Transport Assay Using Reconstituted Yeast Membranes
Sarah van Veen [...] Peter Vangheluwe
Jan 20, 2021 4522 Views
Abstract
We have established the use of Candida albicans as a new model system to study mitochondrial biogenesis. This dimorphic yeast provides an excellent system to investigate the coordination of mitochondrial biogenesis with other cellular networks including cellular metabolism and the cell cycle. Unlike the model lab yeast Saccharomyces cerevisiae, C. albicans is not subject to the Crabtree effect and hence grows aerobically in glucose when oxygen is present. Therefore the control of mitochondrial biogenesis in C. albicans is more typical of eukaryotic cells. C. albicans has a fully sequenced genome and there are many published tools for genetic manipulation facilitating Systems Biology approaches. The isolation of mitochondria (as described in the protocol: Preparation of Mitochondria from Candida albicans) (Hewitt et al., 2013) produces a more simplified system that can be interrogated using the standard tools of molecular biology. In addition, the import of radiolabelled proteins described in this protocol is a sensitive technique that can be used to determine details of kinetics and interactions of imported proteins.
Keywords: MitochondriaMaterials and Reagents
- Rabbit Reticulocyte Lysate System, Nuclease Treated (Promega Corporation, catalog number: L4960 )
- 35S labeled cysteine and methionine (PerkinElmer, catalog number: 072007MC )
- SUPERase-In (Life Technologies, Ambion®, catalog number: AM2696 )
- Mitochondria
- Sorbitol
- Valinomycin
- Oligomycin
- Antimycin
- Soybean Trypsin Inhibitor (SBTI) (Sigma-Aldrich)
- Amino acids
- Coomassie Brilliant Blue G250
- Digitonin (Calbiochem, catalog number: CAS 11024-24-1 )
- RNA for in vitro translation (see Recipes)
- Import buffer (IB) (see Recipes)
- Blue native lysis buffer (see Recipes)
- 10x BN (Blue Native) loading dye (see Recipes)
- RNA (see Recipes)
- ATP (see Recipes)
- NADH (see Recipes)
- AVO (see Recipes)
- Proteinase K (PK) (see Recipes)
- Apyrase (see Recipes)
- Phenylmethanesulfonylfluoride (PMSF) (see Recipes)
- Trypsin (see Recipes)
Equipment
- Water bath with temperature control
- Centrifuge
- Filter – 0.45 μm Acrodisc syringe filter (Pall Life Sciences)
Procedure
- Translation Reaction (30 °C for 50 min) – 15 μl reaction, scale as needed.
- 10 μl rabbit reticulocyte lysate.
- 1 μl amino acids mix (1 mM but no cysteine or methionine).
- 1 μl RNA
- 1 μl 35S labeled cysteine and methionine
- 1 μl SUPERase-In (add a maximum of 2.5 μl).
- 10 μl rabbit reticulocyte lysate.
- Mitochondria Preparation
- 100 μl of Import buffer (IB) per 50 μg of mitochondria
- Standard conditions for 100 μl IB:
- ATP (0.4 M) 1 μl
- NADH (0.5 M) 0.4 μl
- Mix by inversion and leave at import temperature (25 °C standard) for 5 min to equilibrate.
- Add 5 μl translation reaction per 100 μl import (reaction mix can be diluted by addition of water and 2.4 M sorbitol to a final sorbitol concentration of 0.6 M so up to 15 μl of this solution can be added to 100 μl of import buffer which can help increase signal if translation efficiency is low) and incubate at import temperature removing samples at suitable time points as described in part 3.
- ATP (0.4 M) 1 μl
- Optional additions:
- AVO mix of mitochondrial inhibitors for dissipation of membrane potential (1 μl per 100 μl).
- Apyrase- 2 μl per 100 μl IB – treat lysate and mitochondria for 10 min at 25 °C.
- AVO mix of mitochondrial inhibitors for dissipation of membrane potential (1 μl per 100 μl).
- 100 μl of Import buffer (IB) per 50 μg of mitochondria
- Stopper solutions
- 100 μl cold IB per 100 μl import in standard conditions
- Additions include the following but concentrations may need to be optimized:
- Trypsin: 1 μl per 100 μl of stopper solution
- SBTI: 4 μl per 200 μl of mix
- PK: 2 μl per 100 μl of stopper solution
- PMSF: 1 mM final (vortex when added)
- PK/trypsin containing solutions need 20 min before addition of inhibitor
- Trypsin: 1 μl per 100 μl of stopper solution
- At each time point remove 95 μl if adding 5 μl translation reaction/100 μL IB or 105 μl if adding 15 μl/100 μl IB.
- 100 μl cold IB per 100 μl import in standard conditions
- Sample Treatment
- Once inhibitor added to all samples spin at 4 °C/8,000 x g/10 min.
- Remove and discard supernatant.
- Add 100 μl cold IB (do not resuspend pellet – add protease inhibitor if protease used in stopper solution).
- Spin at 4 °C/8,000 x g/5 min.
- Remove and discard supernatant.
(For SDS resuspend pellet in appropriate amounts of water and SDS loading dye)
For blue native PAGE: - Resuspend pellet in 50 μl cold blue native lysis buffer (see Recipes) per 2 μl of mitochondria.
- Leave samples on ice for 15 min with gentle vortex every 5 min.
(FOR ANTIBODY SHIFT: Add 2 μl antibody and leave on ice for 1 h – gentle vortex every 20 min – then continue as per method below) - Spin samples at 4 °C/14,000 x g/10 min (or 15,000 if available).
- Remove 45 μl supernatant to new tube with 5 μl blue Native loading dye.
- Once inhibitor added to all samples spin at 4 °C/8,000 x g/10 min.
Recipes
- RNA from in vitro transcription reaction using 5 μg linear plasmid DNA (or 1.5 μg of PCR product with SP6 promoter)
- Import buffer (IB)
0.6 M sorbitol
50 mM HEPES (adjust to pH7.4 with KOH)
2 mM KPi (pH 7.4 made by mixing 1 M KH2PO4 with 1 M K2HPO4)
25 mM KCl
10 mM MgCl2
0.5 mM EDTA
1 mM DTT stored at RT (room temperature) - ATP
Aliquots of 0.4 M stored at -20 °C (pH to 7) - NADH
Aliquots of 0.5 M stored at -20 °C - AVO
100x stock frozen in EtOH & foiled (1 μM valinomycin, 20 μM oligomycin, 8 μM antimycin) stored at -20 °C - PK (Proteinase K)
5 mg/ml aliquots in IB stored at -20 °C - PMSF
0.2 M in EtOH stored at RT for up to 1 month - Trypsin
Use 10 mg/ml stock (to give 0.1 mg/ml final) for 20 min (stored at -20 °C) - SBTI
50 mg/ml (for 5 min on ice) (stored at -20 °C) - Apyrase
500 units/ml in 50% glycerol stored at -20 °C - 10x BN (Blue Native) loading dye
0.5 M aminocaproic acid
0.1 M Bis Tris pH 7.0
10% Coomassie Brilliant Blue G250 – filter and store at room temperature - Blue native lysis buffer
20 mM Tris-HCl
0.1 mM EDTA
50 mM NaCl
10% glycerol (store at room temperature)
Add PMSF and 5% w/v digitonin (Calbiochem – heat to dissolve and store at 4 °C) to give a final concentration of 1% digitonin and 1 mM PMSF just before use.
Acknowledgments
The protocol was adapted from: Hewitt et al. (2012). The work in the Traven lab and Lithgow lab on Candida albicans mitochondria is supported by a project grant from the Australian National Health and Medical Research Council (APP1023973). V.H. is the recipient of an Australian Postgraduate Award.
References
- Chan, N. C. and T. Lithgow (2008). The peripheral membrane subunits of the SAM complex function codependently in mitochondrial outer membrane biogenesis. Mol Biol Cell 19(1): 126-136.
- Hewitt, V. L., Heinz, E., Shingu-Vazquez, M., Qu, Y., Jelicic, B., Lo, T. L., Beilharz, T. H., Dumsday, G., Gabriel, K., Traven, A. and Lithgow, T. (2012). A model system for mitochondrial biogenesis reveals evolutionary rewiring of protein import and membrane assembly pathways. Proc Natl Acad Sci U S A 109(49): E3358-3366.
- Hewitt, V., Lithgow, T. and Traven, A. (2013). Preparation of Mitochondria from Candida albicans. Bio-protocol 3(4): e330.
Article Information
Copyright
© 2013 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Hewitt, V. L., Lithgow, T. and Traven, A. (2013). Candida albicans Mitochondrial Protein Import Assay. Bio-protocol 3(4): e331. DOI: 10.21769/BioProtoc.331.
Category
Cell Biology > Cell-based analysis > Transport
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









