- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Preparation and Purification of Active Recombinant Human Pancreatic Lipase in Escherichia coli
Published: Vol 9, Iss 13, Jul 5, 2019 DOI: 10.21769/BioProtoc.3286 Views: 5939
Reviewed by: Alessandro DidonnaLouise Jane WalportJane Antony

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Expression and Purification of the Human Voltage-Gated Proton Channel (hHv1)
Emerson M. Carmona [...] Luis G. Cuello
Jun 20, 2025 1986 Views
Prokaryotic Expression and Purification of the hSox2-HMG Domain
Lijie Yang [...] Jingjun Hong
Aug 20, 2025 2367 Views
On-Column Dual-Gradient Refolding for Efficient Recovery of Insoluble Affinity-Tagged Recombinant Proteins
Anna Vlaskina [...] Maxim Patrushev
Feb 5, 2026 39 Views
Abstract
Human pancreatic lipase (HPL) is the main lipolytic enzyme involved in the digestion of dietary fat. An active recombinant human pancreatic lipase (recHPL) was successfully prepared for the first time in an Escherichia coli (E. coli) expression system using a short Strep-tag II (ST II). The recHPL-ST II was solubilized with 8 M urea from the E. coli lysate and purified on a Strep-Tactin-Sepharose column. After refolding by stepwise dialyses against decreasing concentrations of urea in the presence of glycerol and Ca2+ for two days followed by gel filtration FPLC, 1.8-6 mg of active recHPL-ST II was obtained from 1 L of culture. Here we report the expression, purification, and optimized refolding procedures for active recHPL from E. coli, thus establishing it as a suitable system for the production of recHPL of high purity and scaling up.
Keywords: Human pancreatic lipaseBackground
Mammalian lipase, essential for the dietary fat digestion and absorption, can be used for development of an anti-obesity treatment. However, obtaining pure and stable native mammalian lipase has been a challenge because it is often degraded by co-existing proteases in the pancreatic juice. Due to the limited availability of human tissue samples and their potential risk for disease transmission, it may be best to prepare HPL using recombinant expression systems. Several methods have been reported to express recHPL using yeast (Yang and Lowe, 1998), insect (Thirstrup et al., 1993), or mammalian cells (Canalias et al., 1994), which require culturing for at least 5, 6, or 10 days, respectively. The recHPLs obtained using these cells require strict quality control to achieve homogeneity because they are differentially glycosylated. The use of E. coli as an expression system can avoid these problems in addition to providing other benefits such as ease of culture and purification. Therefore, we have established a protocol for optimized expression, purification, and refolding of active recHPL from E. coli. This method requires only two days of culturing, is easy to scale up at low cost, and is convenient because it circumvents the need to control the glycosylation pattern of the recombinant protein. Although the recHPL is non-glycosylated and native porcine pancreatic lipase (PPL) is glycosylated, both lipases show lipolytic activity equal to native PPL against tributyrin and triolein as substrates (Kawaguchi et al., 2018). It may be useful as an alternative to the PPL that has been used in the development of pharmaceutical products. For example, pancreatic enzyme replacement therapy for pancreatic exocrine insufficiency (Lowe and Whitcomb, 2015), and screening of inhibitor compounds to provide an important avenue for anti-obesity therapy (Seyedan et al., 2015). Other promising applications of recHPL are industrial biocatalysts for biotechnological processes (Sharma et al., 2001).
Materials and Reagents
- 50 ml tubes (Labcon, catalog number: 3181-345-008)
- 15 ml tubes (Labcon, catalog number: 3131-345-008)
- 1.5 ml tubes (BIO-BIK, catalog number: ST-0150F)
- 0.2 ml PCR tubes with Dome Cap (Axygen, Funakoshi, catalog number: PCR-02D-C)
- Pipette tips 10 μl (BIO-BIK, catalog number: 1050N), 200 μl (BIO-BIK, catalog number: PT-200), 1,000 μl (BIO-BIK, catalog number: 804C)
- 96-well microplates (Greiner Bio-One, catalog number: 655801)
- Disposable 2 ml polystyrene columns (Pierce, catalog number: 29920)
- Ultrafiltration discs, 10 kDa NMW (Millipore, catalog number: PBGC02510)
- Cellulose acetate membrane filter, 0.45 μm (Advantec, catalog number: C045A090C)
- GL chromatodisc 0.45 μm (GL Science, 25N)
- PVDF membrane (Millipore, catalog number: IPVH00010)
- Dialysis membrane (As One, catalog number: 2-316-02)
- HiPrep 16/60 Sephacryl S-200 HR (GE Healthcare, catalog number: 17-1166-01)
- Sterile Petri dishes (As One, catalog number: 1-7484-01)
- E. coli DH5α competent cells (Takara Bio, catalog number: 9057)
- Stirred Ultrafiltration cells (Millipore, catalog number: 1532701)
- Primers (Invitrogen)
Primer 1 (forward), 5′-AAAGAAGTTTGCTACGAAAGACTC-3′
Primer 2 (reverse), 5′-GATTGTGCCACACTCCCACTCG-3′
Primer 3 (forward), 5′-ACCAAAAGATGTCAACACCCGCTTCC-3′
Primer 4 (reverse), 5′-GACCAAGGCAATATATGGAGGGGTC-3′
Primer 5 (forward), 5′-ATGGTAGGTC TCAAATGAAAGAAGTTTGCTACGAAAGACTCG-3′
Primer 6 (reverse), 5′-ATGGTAGGTCTCAGCGCTACACGGTGTGAGGGTGAGCAG-3′
Primer 7 (forward), 5′-CCCAGCAGAACCTTGCTTTCAGG-3′
Primer 8 (reverse), 5′-CTTCCCAGATTCCGTCTATGTCC-3′ - pASK-IBA3plus plasmid (IBA, catalog number: 2-1402-000)
- Strep-Tactin-Sepharose (IBA, catalog number: 2-1201-010)
- Human MTC Panel I (Clontech Laboratories, catalog number: 636742)
- PfuTurbo DNA Polymerase (Agilent Technologies, catalog number: 600250)
- KOD-Plus-Mutagenesis Kit (Toyobo Co., Ltd., catalog number: SMK-101)
- MgSO4·7H2O (Wako, catalog number: 138-00415)
- Anti-lipase antibodies (Abcam, catalog number: ab96100)
- HRP-conjugated goat anti-rabbit antibodies (Kirkegaard and Perry Laboratories, catalog number: 074-15-061)
- BSA, Albumin from bovine serum, Low salt (Wako, 017-15124)
- BsaI (BioLabs, catalog number: R0535)
- PCR purification kit (Wizard SV® Gel and PCR Clean-Up system) (Promega, catalog number: A9281)
- Ligase enzyme T4 DNA Ligase (Toyobo Co. Ltd, Osaka)
- BigDye TerminatorTM Purification Kit (Thermo Fisher Scientific, catalog number: 4376484)
- FastDigest® Eco31I (Thermo Fisher Scientific, catalog number: FDO293)
- 10x FastDigest®Green Buffer (Thermo Fisher Scientific, catalog number: B72)
- Ligation high ver.2 (TOYOBO, catalog number: LGK-201)
- Ampicillin sodium (Wako, catalog number: 016-10373)
- Bacto Agar (Becton, Dickinson and Co., catalog number: 214010)
- Luria Bertani (LB) Broth Base (Invitrogen, catalog number: 12780-052)
- Anhydrotetracycline (IBA, catalog number: 2-0401-002)
- Urea (Wako, catalog number: 217-00171)
- Dithiothreitol (DTT) (Wako, catalog number: 049-08972)
- Desthiobiotin (Sigma-Aldrich, catalog number: D1411)
- Ethylenediaminetetraacetic acid (EDTA) (Wako, catalog number: 345-01865)
- Sodium chloride (NaCl) (Kanto Chemical Co., Inc., catalog number: 37144-86)
- Sodium hydroxide (NaOH) (Wako, catalog number: 198-13765)
- Phenylmethylsulfonyl fluoride (PMSF) (Nacalai Tesque, catalog number: 27327-94)
- Tris (hydroxylmethyl) aminomethane (Wako, catalog number: 204-07881)
- Triton X-100 (Sigma-Aldrich, catalog number: T9284)
- L-Arginine hydrochloride (Wako, catalog number: 018-04625)
- CBB R-250 (Wako, catalog number: 031-17922)
- Methanol (Wako, catalog number: 25183-70)
- Acetic acid (Wako, catalog number: 017-00251)
- L-Cystine dihydrochloride (Wako, catalog number: 03405322)
- L-Cysteine (Wako, catalog number: 039-20652)
- Glycerol (Wako, catalog number: 075-00611)
- CaCl2 (Wako, catalog number: 031-00435)
- Tributyrin (Wako, catalog number: 207-02392)
- Colipase, Pig (Bio-Rad, catalog number: 2148-1004)
- Sodium taurodeoxycholate
- LB agar plates (see Recipes)
- LB medium (see Recipes)
- Buffer A (see Recipes)
- Lysis buffer (see Recipes)
- Buffer A* (see Recipes)
- Equivalent buffer (see Recipes)
- Elution buffer (see Recipes)
- Refolding buffer (see Recipes)
- Running buffer (see Recipes)
- Reaction buffer (see Recipes)
Equipment
- 300 ml shaking flask (Iwaki, catalog number: 4070FK300)
- 500 ml glass beaker (e.g., As One, catalog number: 2-5091-06)
- Pipettes (Eppendorf, Reference 10-100 μl, 0.5-10 μl, catalog numbers: 4920000059, 4920000024)
- Magnetic stirring bar (NALGENE, catalog number: 93-5521-5)
- -80 °C freezer (Nihon Freezer MY BIO, model: VT-208)
- Zymoreactor II thermal cycler (ATTO)
- Prism 3100 Avant sequencer (Applied Biosystems)
- Shaking incubator (e.g., Thomas, model: AT-12R)
- Refrigerated bench-top centrifuge (e.g., Eppendorf, model: 5417R)
- High speed centrifuge (Hitachi, model: CR20GIII)
- Sonicator (Branson, model: Advanced Sonifier 250)
- Spectrophotometer (DeNovix, model: DS-11)
- Microplate spectrophotometer (DS Pharma Biomedical, model: Viento XS)
- Fraction collector (e.g., Advantec, model: SF-2100)
- AmiconTM, Bioseparations Stirred Cell (Merck, catalog number: 8010)
- AKTA Purifier chromatography system (GE Healthcare)
- pH-stat AT-510 (Kyoto Electronics Manufacturing Co, Ltd.)
Software
- UNICORN (GE Healthcare), control software for the AKTA purifier chromatography system
- Gen 5 (Bio Tek), data collection and analysis software for the microplate reader
- AT-Win (Kyoto Electronics Manufacturing Co, Ltd.), the control software for the pH-stat, automatic potentiometric titrator, AT-510
Procedure
This procedure to prepare recHPL includes five steps: (A) construction of expression plasmid, (B) expression in E. coli, (C) isolation, (D) refolding, and (E) purification.
- Construction of HPL expression plasmid
ST II-fused recHPL was cloned into the pASK-IBA3plus plasmid to allow expression in E. coli. The construct contains a C-terminal Strep-tag II to facilitate separation of recHPL (Figure 1).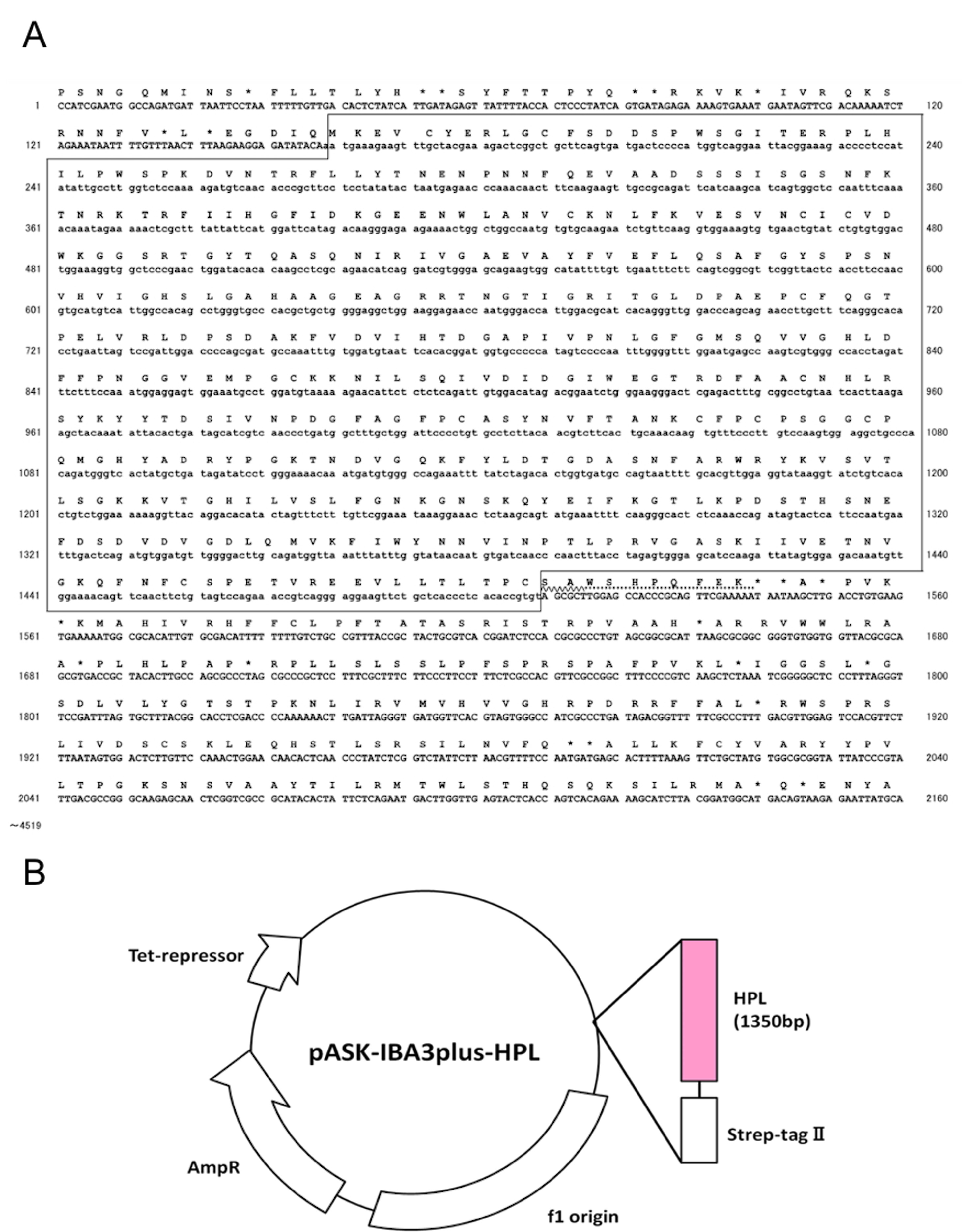
Figure 1. Sequence of recHPL. A. Part of the nucleotide sequence of an HPL-pASK-IBA3plus plasmid and the deduced amino acid sequence. The entire sequence of HPL was confirmed using an autosequencer. Boxed region: the coding sequence of HPL, wavy underlines: the sequence of the linker, dotted underlines: the sequence of the ST II-tag. B. Structure of pASK-IBA 3Plus-HPL plasmid.- Following the PCR protocol detailed below, amplify the coding sequence of HPL using the human pancreas cDNA library, Human MTC Panel I as the template for PCR using a Zymoreactor II (ATTO) thermal cycler with following conditions:
[Primers]
Primer 1 (forward), 5′-AAAGAAGTTTGCTACGAAAGACTC-3′
Primer 2 (reverse), 5′-GATTGTGCCACACTCCCACTCG-3′
[Reaction mixture]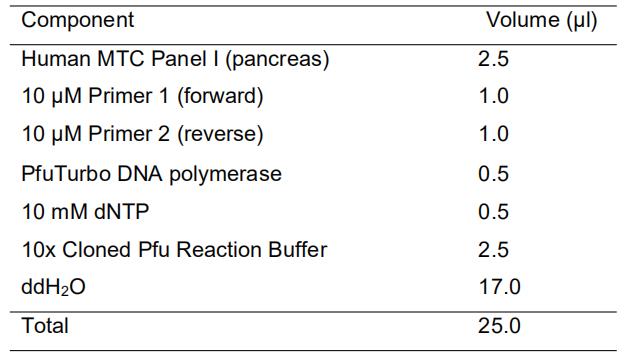
[PCR conditions]30 cycles of 95 °C 60 s (Denaturation) 55 °C 60 s (Annealing) 72 °C 60 s (Extension) - Separate the amplicon by agarose gel electrophoresis using 1% agarose gel.
- Cut out the DNA band (expected size 1,350 bp) on the gel and purify with Wizard® SV Gel and PCR Clean-Up System to obtain insert DNA.
- Perform a second PCR to replace the Eco311 restriction endonuclease recognition sequence GGTCTC contained in the insert DNA with GGTCAC using a Zymoreactor II (ATTO) thermal cycler with following conditions.
[Primers]
Primer 3 (forward), 5′-ACCAAAAGATGTCAACACCCGCTTCC-3′
Primer 4 (reverse), 5′-GACCAAGGCAATATATGGAGGGGTC-3′
[Reaction mixture]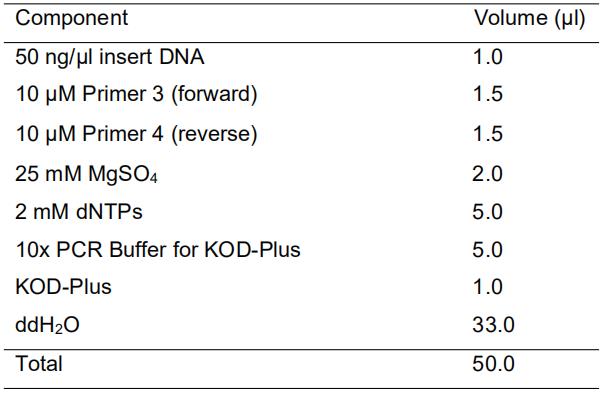
[PCR conditions]10 cycles of 94 °C 120 s 98 °C 10 s 68 °C 360 s - Separate the amplicon by agarose gel electrophoresis using 1% agarose gel.
- Cut out the DNA band (expected size 1,350 bp) on the gel and purify it with Wizard® SV Gel and PCR Clean-Up System to obtain insert DNA.
- Perform a third PCR to obtain insert DNA with Eco31 I restriction endonuclease recognition site added at both ends using Zymoreactor II (ATTO) thermal cycler under the following condition:
[Primers]
Primer 5 (forward), 5′-ATGGTAGGTC TCAAATGAAAGAAGTTTGCTACGAAAGACTCG-3′
Primer 6 (reverse), 5′-ATGGTAGGTCTCAGCGCTACACGGTGTGAGGGTGAGCAG-3′
[Reaction mixture]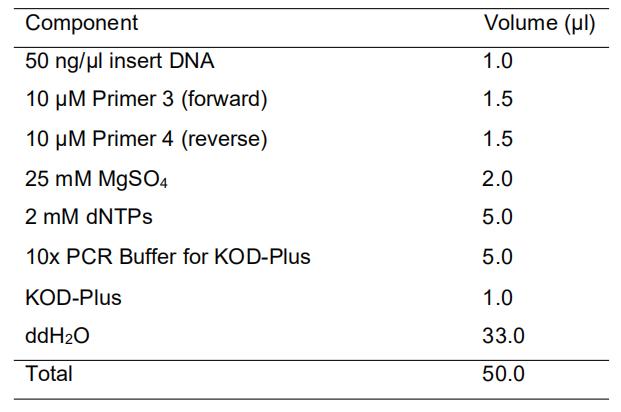
[PCR conditions]Initial denaturation 94 °C 120 s 25 cycles of 94 °C 15 s 65 °C 30 s 68 °C 60 s - Purify the sequencing reactions using a BigDye TerminatorTM Purification Kit.
- Digest the amplicon 2 μl by adding FastDigest® Eco31I 1 μl, 10x FastDigest Green Buffer 2 μl, and ddH2O 15 μl at 37 °C for 5 min.
- Digest pASK-IBA3Plus 2 μl by adding FastDigest® Eco31I 1 μl, 10x FastDigest Green Buffer 2 μl, and ddH2O 15 μl at 37 °C for 5 min.
- Separate the digested samples by agarose gel electrophoresis using 1% agarose gel.
- Cut out the DNA bands (expected sizes of insert DNA and pASK-IBA3Plus are 1,350 bp and 3,247 bp, respectively) on the gel and purify with Wizard® SV Gel and PCR Clean-Up.
- Ligate the digested insert DNA with the digested pASK-IBA3plus to obtain an HPL-pASK-IBA3plus plasmid by DNA Ligation Kit, Ligation high Ver.2.
- Sequence all constructed plasmids using the Prism 3100 Avant sequencer to confirm the sequences of the designed recombinant HPLs. Use the following primers:
Primer 7 (forward), 5′-CCCAGCAGAACCTTGCTTTCAGG-3′
Primer 8 (reverse), 5′-CTTCCCAGATTCCGTCTATGTCC-3′
- Following the PCR protocol detailed below, amplify the coding sequence of HPL using the human pancreas cDNA library, Human MTC Panel I as the template for PCR using a Zymoreactor II (ATTO) thermal cycler with following conditions:
- Expression of recHPL in E. coli
- Transform 5 μl (10-fold dilution) of HPL-pASK-IBA3plus plasmid (ca. 2.5 ng) into 50 μl of E. coli DH5α competent cells in a 1.5 ml tube.
- Incubate for 30 min on ice.
- Incubate at 42 °C for 45 s.
- Add 900 μl of LB medium and culture at 37 °C for 1 h.
- Plate 100 μl of transformants on LB agar plates including 100 μg/ml ampicillin at 37 °C for 18 h.
- Pick up a single colony and grow the transformants in 5 ml LB medium including 100 μg/ml ampicillin in a 50 ml tube at 37 °C for 18 h.
- Transfer the bacterial suspension (3 ml) into 300 ml fresh LB medium including 100 μg/ml ampicillin in a shaking flask and culture for 3 h at 37 °C at 150 rpm.
- Add 30 μl of 2 mg/ml anhydrotetracycline at a final concentration of 200 μg/L when the absorbance of the culture measured at 550 nm by a spectrophotometer (DeNovix) has reached 0.5 to induce expression of recombinant protein.
- Incubate for 5 h at 30 °C with shaking at 120 rpm.
- Centrifuge at 6,100 x g for 15 min at 4 °C using a high speed centrifuge (Hitachi) to harvest cells.
- Discard the supernatant, resuspend the bacterial pellet in 15 ml of buffer A, and freeze at -80 °C in a 50 ml tube until use.
- Isolation of recHPL
- Thaw the frozen bacterial pellet.
- Lyse the bacterial cell pellet by sonication for 2 min (30 s x 4 times, output: 4, duty cycle: 70%) on ice.
- Centrifuge at 4,400 x g for 15 min at 4 °C using a high speed centrifuge (Hitachi).
- Discard the supernatant and resuspend the pellet in 15 ml of lysis buffer.
- Centrifuge at 28,000 x g for 10 min at 4 °C.
- Discard the supernatant and wash the pellet twice with 30 ml of buffer A.
- Centrifuge at 28,000 x g for 10 min at 4 °C.
- Discard the supernatant and solubilize the washed inclusion bodies with 10 ml of buffer A* with agitation for 30 min at 4 °C.
- Add 50 μl of 2 M DTT at a final concentration of 10 mM to the supernatant.
- Incubate for 1 h at 37 °C with agitation at 100 rpm.
- Dilute the solubilized proteins with 10 ml of buffer A*.
- Dialyze against 150 ml of the equivalent buffer using the dialysis membrane (pore diameter = 50 Å) at 4 °C for 2 h to remove urea in a glass beaker with gentle stirring using a magnetic stirrer.
- Centrifuge the dialyzed solution at 28,000 x g for 10 min at 4 °C.
- Aspirate the supernatant and transfer it to a 50 ml tube.
- Add 20 ml of ice-cold equivalent buffer to the 2 ml of Strep-Tactin-Sepharose in a polystyrene column using a Pasteur pipette.
- Remove the equilibrated Strep-Tactin-Sepharose and add to the dialyzed protein solution in the 50 ml tube.
- Incubate at 4 °C for 30 min with gentle shaking at around 30 rpm.
- Pack the suspension mixture of Strep-Tactin-Sepharose into a polystyrene column.
- Wash with 20 ml of equivalent buffer at a flow rate of 3 drops/s, collecting 70-drop fractions using a fraction collector at 4 °C (Figure 2).
- Elute ST II-fused recHPL with elution buffer at a flow rate of 10 drops/s, collecting 22-drop fractions using a fraction collector at 4 °C.
- Measure the absorbance of 200 μl of fractions using 96-well microplates, a microplate spectrophotometer (DS Pharma Biomedical), and software Gen 5 at 280 nm at room temperature.
- Combine fractions with high A280 readings corresponding to the eluted protein and store the eluted peak fractions in a 15 ml tube at -80 °C.
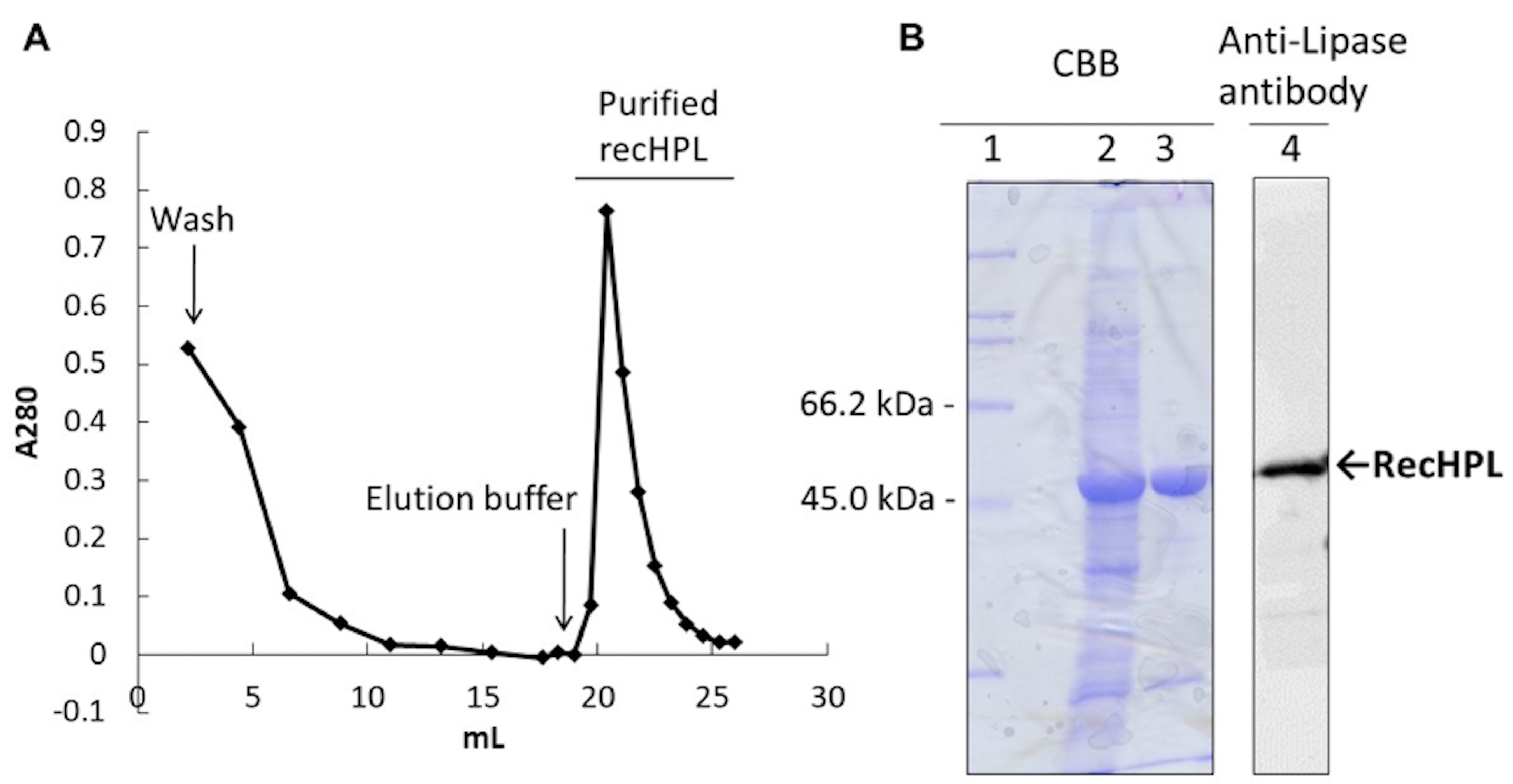
Figure 2. Separation of recHPL by Strep-Tactin-Sepharose column. A. Affinity chromatography profile on a Strep-Tactin-Sepharose column (Schmidt et al., 1996). Elution was started from the fraction indicated by the arrow, and the peak was obtained at elution volume 19-26 ml. Column volume: 1.9 ml (i.e., 7 mm x 50 mm), flow rate: 10 s/drop, temperature: 4 °C. B. SDS-PAGE and Western blot analysis of the recHPL-ST II. From left lane, “CBB” shows a CBB-stained 12% polyacrylamide gel. “Anti-HPL” shows an electroblotted PVDF membrane blocked with 3% BSA and stained using the primary antibody, anti-lipase antibodies (1:3,000) at room temperature for 1 h and the secondary antibody, HRP-conjugated goat anti-rabbit antibodies (1:5,000) at room temperature for 1 h followed by developed using ECL reagent (GE Healthcare). 1, Marker proteins; 2, solubilized protein from inclusion body; 3 and 4, recHPL-ST II purified on the Strep-Tactin-Sepharose column.
- Refolding of RecHPL (Figure 3)
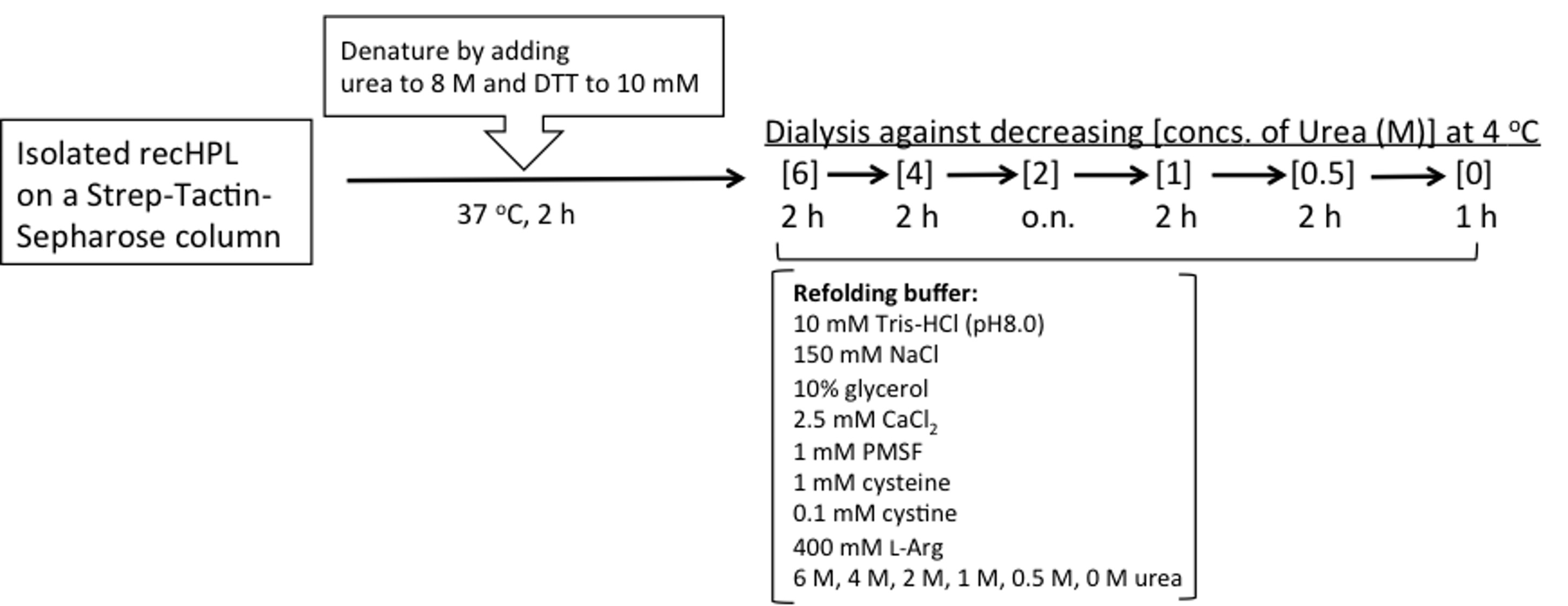
Figure 3. Procedure for refolding of isolated recHPL- Thaw the frozen eluted peak fractions and measure the absorbance at 280 nm at room temperature.
- Add urea to a final concentration of 8 M to 7-12 ml of the purified 0.1-0.2 mg/ml recHPL in peak fractions.
- Add 2 M DTT to a final concentration of 10 mM.
- Incubate for 2 h at 37 °C with agitation at approximately 100 rpm.
- Dialyze against 300 ml of refolding buffer containing a step-wise decrease in urea concentration (6 M and 4 M) at 4 °C for 2 h.
- Dialyze against 300 ml of refolding buffer containing 2 M urea at 4 °C overnight.
- Dialyze against 300 ml of refolding buffer containing step-wise decrease in urea concentration (1 M and 0.5 M) at 4 °C for 2 h.
- Dialyze against 300 ml of refolding buffer containing 0 M urea at 4 °C for 1 h.
- Centrifuge at 21,000 x g for 5 min and collect the supernatant to remove protein aggregates at 4 °C.
- Purification of recHPL (Figure 4)
Note: Store all reagents and carry out all procedures at 4 °C.
- Add the supernatant of the refolded recHPL to an AmiconTM Bioseparations Stirred Cell with an ultrafiltration disc (10 kDa NMWL).
- Concentrate recHPL by 2-10 times using the stirred cell.
- Filter the concentrated recHPL through the GL chromatodisc 0.45 μm (GL Science, 25N)
- Filter the running buffer for FPLC through a cellulose acetate membrane filter, 0.45 μm.
- Inject 1 ml of refolded recHPL into a HiPrep 16/60 Sephacryl S-200 HR column (column volume: 120 ml), pre-equilibrated with running buffer on an AKTA Purifier FPLC system at a flow rate of 0.5 ml/min.
- Elute the column with 180 ml of running buffer at a flow rate of 0.5 ml/min, collecting 2 ml fractions.
- Measure the absorbance of fractions at 280 nm and store approximately 5 fractions (each 2 ml) of the second peak as active recHPL.
- Determine the lipolytic activity of recHPL using tributyrin as substrate by automated titration of fatty acids released with 10 mM NaOH using the pH-stat AT-510 (Kawaguchi et al., 2018).
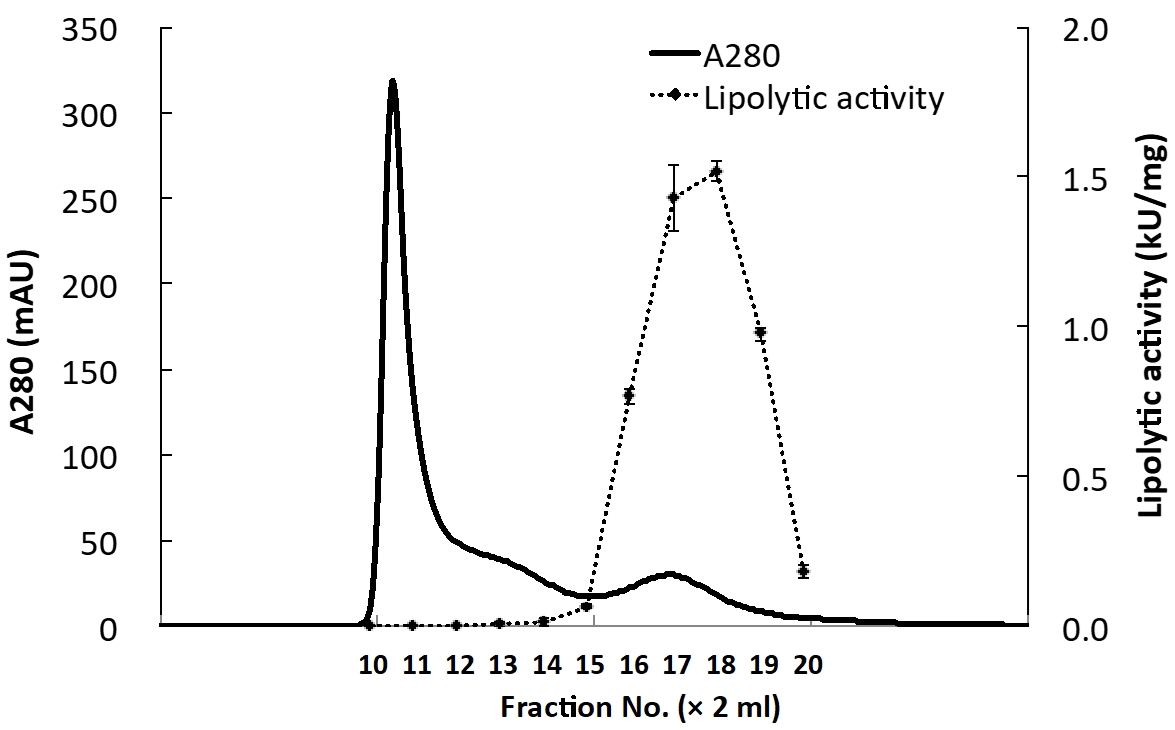
Figure 4. Purification of active recHPL by gel-filtration FPLC. Elution profile of recHPL on size exclusion. RecHPL after refolding was loaded onto a HiPrep 16/60 Sephacryl S-200 column (column volume: 120 ml) and eluted using 10 mM TBS containing 400 mM L-Arg at a flow rate of 0.5 ml/min at 4 °C. Protein peaks were detected by measuring absorbance at 280 nm using a UPC-900 monitor and AKTA Purifier. Active recHPL-ST II (1.8-6 mg) was obtained from 1 L of culture, and it showed a specific activity of 1.2-1.9 kU/mg. Specific activity of the fractions was measured using 0.11 M tributyrin emulsified in 1 ml reaction buffer as the substrate. The measurements were performed containing colipase in excess, 0.5 μg/50 μl. After addition of each fraction eluted by gel filtration FPLC in Figure 4 (0.5 μg/50 μl), the NaOH consumption was recorded at a reaction temperature of 37 °C using the automatic potentiometric titrator, pH-stat AT-510 (Kyoto Electronics Manufacturing Co. Ltd.). One unit was defined as the amount of enzyme that released 1.0 μmol of free fatty acid per min.
Data analysis
Protein expression is verified by Coomassie staining and Western blotting using anti-lipase antibodies (Abcam, ab96100). Chromatograms were recorded using UNICORN software. Gen 5 software was used for data collection and analysis for the microplate reader. The pH-stat AT-510 was controlled by AT-Win software.
Notes
- Prepare 0.2 M PMSF stock solution in ethanol and dilute to buffer just before use.
- In a comparison of TBS supplemented with 8 M urea, 1% Triton X-100, or 1% SDS, TBS containing 8 M urea is the most efficient method of solubilizing recHPL-ST II from E. coli lysates.
- A Strep-Tactin-Sepharose column is an open column using gravity to move the solvent. The flow rate is controlled by the hydrostatic pressure on top of the column and the stopcock at the bottom of the column.
- All the processes for isolation, dialysis for refolding, and purification of recHPL should be performed at 4 °C except for the denaturation processes with 8 M urea and 10 mM DTT, which must be done at 37 °C.
- The addition of L-Arg showed a remarkable effect on the refolding of recHPL-ST II, resulting in a high yield, which must be attributable to the inhibition of aggregation of recHPL-ST II by L-Arg.
Recipes
- LB agar plates
20 g/L LB Broth Base
15 g/L bacto agar
Autoclave at 121 °C, 2.6 bar for 20 min to sterilize
Add ampicillin (100 μg/ml) and pour the medium into sterile Petri dishes - LB medium
20 g/L LB Broth Base
Autoclave at 121 °C, 2.6 bar for 20 min to sterilize - Buffer A
10 mM Tris-HCl (pH 8.0)
150 mM NaCl
1 mM EDTA - Lysis buffer
10 mM Tris-HCl (pH 8.0)
150 mM NaCl
1 mM EDTA
0.5% Triton X-100 - Buffer A*
10 mM Tris-HCl (pH 8.0)
150 mM NaCl
1 mM EDTA
8 M urea
1 mM PMSF - Equivalent buffer
10 mM Tris-HCl (pH 8.0)
150 mM NaCl
400 mM L-Arg
1 mM EDTA
1 mM PMSF - Elution buffer
10 mM Tris-HCl (pH 8.0)
150 mM NaCl
400 mM L-Arg
1 mM EDTA
1 mM PMSF
2.5 mM desthiobiotin - Refolding buffer
10 mM Tris-HCl (pH 8.0)
150 mM NaCl
10% glycerol
2.5 mM CaCl2
1 mM PMSF
1 mM cysteine
0.1 mM cystine
400 mM L-Arg
Varing urea concentration: 6 M, 4 M, 2 M, 1 M, 0.5 M, and 0 M urea - Running buffer
10 mM Tris-HCl (pH 8.0)
150 mM NaCl
400 mM L-Arg - Reaction buffer
1 mM Tris-HCl (pH 7.5)
4 mM sodium taurodeoxycholate
100 mM NaCl
5 mM CaCl2
Acknowledgments
We sincerely thank Daiwa Securities Health Foundation (HO) and the Japan Foundation of Applied Enzymology (HO) for support with research funds. We sincerely thank Prof. Yusuke Suzuki, Ms. Chihiro Tomita, Ms. Rina Naradate, Ms. Tomoko Higami, Dr. Kosuke Nakamura, and Ms. Hiromi Sakagami for assisting in this work. We thank Mrs. K. Ono for English editing.
Competing interests
The authors declare there are no conflicts of interest.
References
- Canalias, F., Visvikis, A., Thioudellet, C. and Siest, G. (1994). Stable expression of enzymatically active human pancreatic lipase in V79 cells: purification and characterization of the recombinant enzyme. Clin Chem 40(7 Pt 1): 1251-1257.
- Kawaguchi, N., Date, K., Suzuki, Y., Tomita, C., Naradate, R., Higami, T., Nakamura, K., Aikawa, K. and Ogawa, H. (2018). A novel protocol for the preparation of active recombinant human pancreatic lipase from Escherichia coli. J Biochem 164(6): 407-414.
- Lowe, M. E. and Whitcomb, D. C. (2015). Next generation of pancreatic enzyme replacement therapy: Recombinant microbial enzymes and finding the perfect lipase. Gastroenterology 149(7): 1678-1681.
- Schmidt, G., Koepke, J., Frank, R. and Skerra, A. (1996). Molecular interaction between the Strep-tag affinity peptide and its cognate target, streptavidin. J Mol Biol 255: 753-766.
- Seyedan, A., Alshawsh, M. A., Alshagga, M. A., Koosha, S. and Mohamed, Z. (2015). Medicinal plants and their inhibitory activities against pancreatic lipase: A review. Evid Based Complement Alternat Med 2015: 973143.
- Sharma, R., Chisti, Y. and Banerjee, C. (2001). Production, purification, characterization, and applications of lipases. Biotechnol Adv 19: 627-662.
- Thirstrup, K., Carriere, F., Hjorth, S., Rasmussen, P. B., Woldike, H., Nielsen, P. F. and Thim, L. (1993). One-step purification and characterization of human pancreatic lipase expressed in insect cells. FEBS Lett 327(1): 79-84.
- Yang, Y. and Lowe, M. E. (1998). Human pancreatic triglyceride lipase expressed in yeast cells: purification and characterization. Protein Expr Purif 13(1): 36-40.
Article Information
Copyright
© 2019 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Kawaguchi, N., Ogawa, H. and Date, K. (2019). Preparation and Purification of Active Recombinant Human Pancreatic Lipase in Escherichia coli. Bio-protocol 9(13): e3286. DOI: 10.21769/BioProtoc.3286.
Category
Microbiology > Heterologous expression system > Escherichia coli
Biochemistry > Protein > Expression
Molecular Biology > Protein > Activity
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link