- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Determining Oxidative Damage by Lipid Peroxidation Assay in Rat Serum
Published: Vol 9, Iss 12, Jun 20, 2019 DOI: 10.21769/BioProtoc.3263 Views: 9506
Reviewed by: Davide BottaHua WangAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

OrganoPlate Micro-fluidic Microvessel Culture and Analysis
Abidemi Junaid and Thomas Hankemeier
Jul 5, 2021 4559 Views
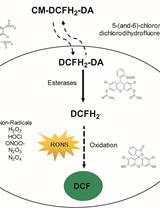
Measurement of Reactive Oxygen and Nitrogen Species in Living Cells Using the Probe 2',7'-Dichlorodihydrofluorescein
Waleska Dornas and Detlef Schuppan
Dec 20, 2021 3898 Views
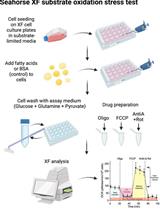
Mass Spectrometry-based Lipidomics, Lipid Bioenergetics, and Web Tool for Lipid Profiling and Quantification in Human Cells
Liang Cui [...] Kuan Rong Chan
Aug 20, 2023 3039 Views
Abstract
It has been well-established that malondialdehyde (MDA), which is generated during the process of lipid peroxidation, is a commonly known biomarker for oxidative stress. Therefore, the serum levels of MDA are detected by using the lipid peroxidation assay with commercially available kit to determine the induction of oxidative stress in rat models.
Keywords: Oxidative stressBackground
As lipid peroxidation is the degradation of lipids that occurs as a result of oxidative damage and contributes to the pathology of many diseases, some end-products of the chain reaction of lipid peroxidation such as malondialdehyde (MDA), 4-Hydroxynonenal (4-HNE) and 8-iso-Prostaglandin F2alpha (8-isoprostane) in serum samples have been detected and quantified for the identification of oxidative damage (Marrocco et al., 2017). In the current lipid peroxidation assay protocol, the serum levels of MDA could be specifically and reliably quantified based on the condensation reaction between MDA and thiobarbituric acid (TBA) by modifying a commercially available kit (MAK085D; Sigma, St Louis, MO) without n-butanol precipitation step (Tang et al., 2019).
Materials and Reagents
- Pipette tips
- 96-well flat bottom microplate (Sigma-Aldrich, catalog number: CLS3610)
- Separate tube (Corning, catalog number: CLS430791)
- Double-distilled water (ddH2O)
- Ice
- 60 μl freshly collected rat serum
- TBA (Sigma-Aldrich, catalog number: MAK085D)
- Glacial acetic acid (Sigma-Aldrich, catalog number: A6283)
- 4.17 M MDA standard (Sigma-Aldrich, catalog number: MAK085E)
- 42 mM Sulfuric acid solution (Sigma-Aldrich, catalog number: 84736)
- Phosphotungstic acid solution (PTA) (Sigma-Aldrich, catalog number: MAK085B)
- Butylated hydroxytoluene (BHT) (Sigma-Aldrich, catalog number: MAK085C)
- MDA standard (see Recipes)
Equipment
- Pipettes (Bio-rad, P10, P200 and P1000)
- Vortexer (Ratek, catalog number: VM1)
- Centrifuge (Thermoline, model: K1015 Pro)
- Incubator (Robbins Scientific, catalog number: 2000)
- Microplate reader (Perkin Elmer, catalog number: 1420 Victor3)
Software
- Microsoft Excel 2010
Procedure
The flow chart in Figure 1 shows all the steps described in this protocol.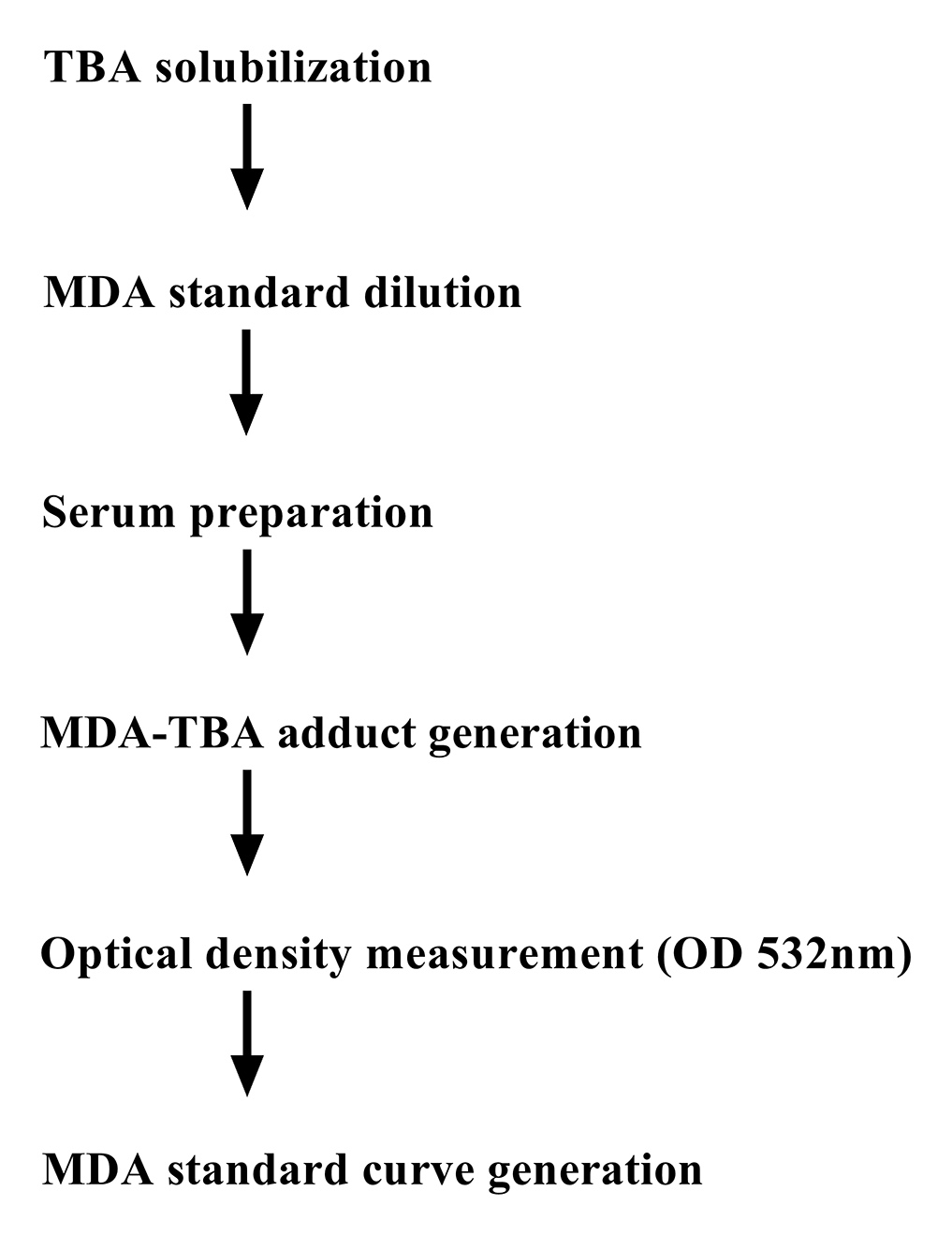
Figure 1. Schematic diagram of the assay procedure
- Reconstitute a bottle of TBA with 7.5 ml glacial acetic acid followed by adjusting with ddH2O to make 25 ml TBA solution.
- Dilute 10 μl of 4.17 M MDA standard in 407 μl of ddH2O to make 0.1 M MDA standard.
- Dilute 10 μl of 0.1 M MDA standard in 490 μl of ddH2O to make 2 mM MDA standard.
- Use 2 mM MDA standard to generate MDA standard curve dilutions (Recipe 1).
- Mix 20 μl of serum with 500 μl of 42 mM sulfuric acid.
- Add 125 μl of PTA followed by vortexing.
- Centrifuge at 13,000 x g at room temperature for 5 min.
- Add 200 μl of BHT to 10 ml of ddH2O in a separate tube followed by vortexing.
- Collect and resuspend serum pellet with 102 μl of BHT/ddH2O on ice.
- Adjust the final volume to 200 μl with ddH2O followed by vortexing.
- Incubate at 37 °C for 2 h.
- Mix 600 μl of TBA solution with 200 μl MDA standard/serum sample to generate 800 μl of MDA-TBA adduct.
- Incubate at 95 °C for 1 h and then place on ice bath for 15 min.
- Pipette 200 μl of MDA-TBA adduct into a 96-well microplate in duplicate.
- Measure the absorbance at a wavelength of 532 nm on the microplate reader.
- Establish the standard curve using the serial dilutions of MDA standard.
- Calculate MDA concentrations in serum samples.
Data analysis
- Average the duplicate values for each reading.
- Set the mean value of the blank (Standard #1) as the background.
- Correct for the background by subtracting the blank value from all readings.
- Use the corrected values of MDA standards to plot a standard curve by using Microsoft Excel 2010 (Figure 2).
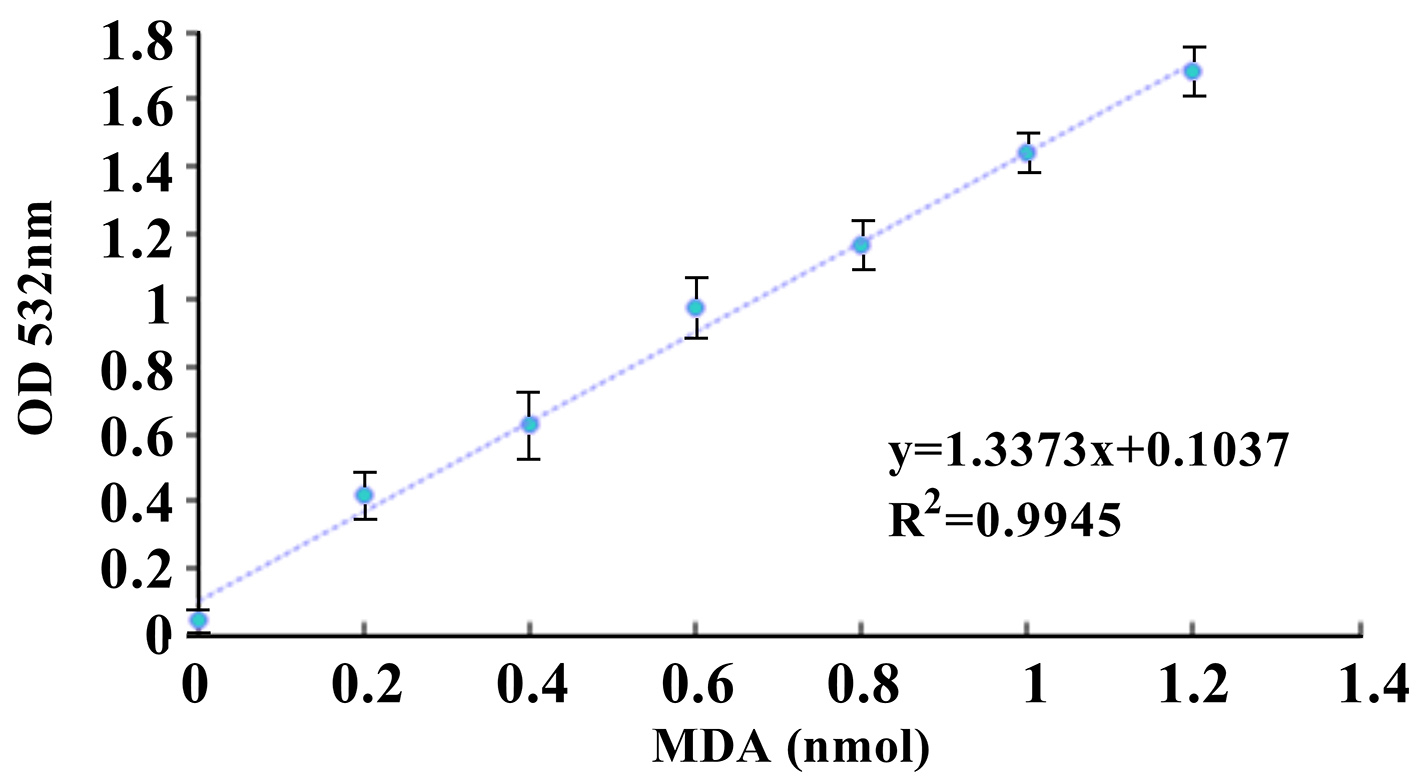
Figure 2. MDA standard curve. The standard curve was generated by using linear regression in Microsoft Excel 2010. - Obtain MDA amount in the sample wells based on the standard curve by using linear regression.
- Concentration of MDA in the test serum samples is calculated as:
MDA concertation = (A /0.02 ml) x 4 x D
where,
A = Amount of MDA in sample calculated from the standard curve (nmol).
0.02 ml = Original serum volume used (200 μl).
4 = Correction for using 200 μl of reaction mix from 800 μl of MDA-TBA adduct.
D = Sample dilution factor (if sample is diluted to fit within the standard curve range).
Notes
- Equilibrate all materials and prepared reagents to room temperature just prior to use and gently agitate.
- Under the current experimental settings, the whole blood sample was centrifuged at 2,000 x g at 4 °C for 10 min to collect serum. The serum was then immediately apportioned into 0.5 ml aliquots and stored at -20 °C.
- A new standard curve must be set up each time the assay is run.
- N-butanol precipitation step could be performed to enhance assay sensitivity where MDA-TBA adduct concentration is low in plasma samples.
Recipes
- MDA standard
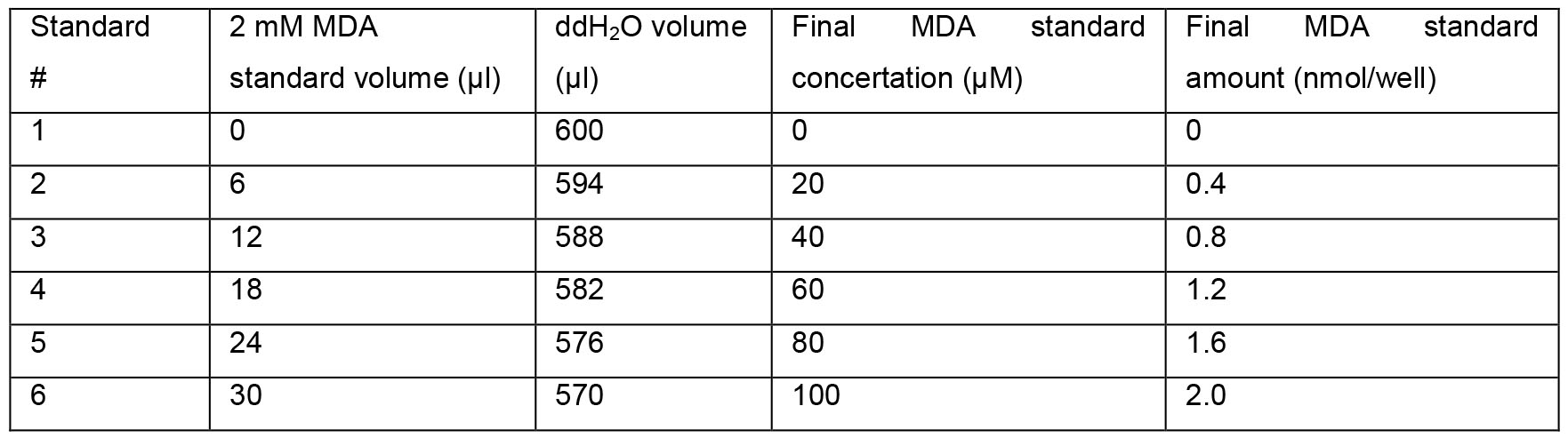
Acknowledgments
CJX is supported by National Health and Medical Research Council of Australia (No. 1127396) and Natural Science Foundation of China (No. 81671928).
Competing interests
The authors declared that they have no conflict of interests to this work.
References
- Marrocco, I., Altieri, F. and Peluso, I. (2017). Measurement and clinical significance of biomarkers of oxidative stress in humans. Oxid Med Cell Longev 2017: 6501046.
- Tang, Q., Su, Y. W., Fan, C. M., Chung, R., Hassanshahi, M., Peymanfar, Y. and Xian, C. J. (2019). Release of CXCL12 from apoptotic skeletal cells contributes to bone growth defects following dexamethasone therapy in rats. J Bone Miner Res 34(2): 310-326.
Article Information
Copyright
© 2019 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Tang, Q., Su, Y. and Xian, C. J. (2019). Determining Oxidative Damage by Lipid Peroxidation Assay in Rat Serum. Bio-protocol 9(12): e3263. DOI: 10.21769/BioProtoc.3263.
Category
Developmental Biology > Cell signaling > Stress response
Cell Biology > Cell metabolism > Lipid
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









