- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Assessment of Humoral Alloimmunity in Mixed Lymphocyte Reaction
Published: Vol 9, Iss 2, Jan 20, 2019 DOI: 10.21769/BioProtoc.3139 Views: 11596
Reviewed by: Vemika ChandraEmilie BattivelliAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Identification and Sorting of Adipose Inflammatory and Metabolically Activated Macrophages in Diet-Induced Obesity
Dan Wu [...] Weidong Wang
Oct 20, 2025 2232 Views
Selective Enrichment and Identification of Cerebrospinal Fluid-Contacting Neurons In Vitro via PKD2L1 Promoter-Driven Lentiviral System
Wei Tan [...] Qing Li
Nov 20, 2025 1335 Views
Revisiting Primary Microglia Isolation Protocol: An Improved Method for Microglia Extraction
Jianwei Li [...] Guohui Lu
Dec 5, 2025 1460 Views
Abstract
Humoral alloimmunity remains a significant and unresolved problem that constrains allograft survival. Thus, there is a need for the development of an easy, preferably non-radioactive, and inexpensive protocol for assessing the effect of various drug treatments on humoral alloimmunity. In order to satisfy this demand, we developed such a protocol in which de novo alloantibodies production is induced in one-way mixed lymphocyte reaction (MLR). The amount and capacity of the generated alloantibodies in the supernatant of each one-way MLR is assessed using an antibody-mediated cell dependent cytotoxicity (CDC) assay. The principle of the assay relies on the assessment of cellular survival of resting PBMCs isolated from the same donors, supplemented with the alloantibodies from the one-way MLR supernatant. The lesser is the cellular survival, the higher the production of alloantibodies in one-way MLR and consequently the more potent the humoral alloimmunity.
Keywords: Humoral alloimmunityBackground
Kidney transplantation is the treatment of choice for patients with end-stage kidney disease. Modern immunosuppression strategies considerably increased the one-year graft survival by decreasing the frequency and the potency of acute cellular-mediated rejection episodes, though long-term graft survival remains problematic mainly due to chronic antibody-mediated rejection, a case without effective treatment (Abramowicz et al., 2018). Thus, there is a need for the development of an easy, preferably non-radioactive, and inexpensive protocol for assessing humoral alloimmunity during various drug interventions, as well as for preclinical evaluation of new medications.
We developed such a protocol in order to test the effect of certain compounds on humoral alloimmunity (Eleftheriadis et al., 2017a, 2017b and 2017c). Although this protocol has been applied in human peripheral blood mononuclear cells (PMBCs), it could also be easily modified for using cells from other species, for instance from different mouse strains.
Our protocol was based on previous studies, supporting that one week is enough for the development of IgM and IgG alloantibodies in one-way mixed lymphocyte reaction (MLR) with a stimulator to responder cell ratio equal to 1:1 (Rumke et al., 1982), along with the well-established antibody-mediated complement-dependent cytotoxicity (CDC) assay (Konishi et al., 2008). Mitomycin C was used to treat the stimulator cell population. Mitomycin C is an anti-tumor antibiotic agent, which acts via the inhibition of DNA and RNA synthesis, rendering the lymphocytes unable to proliferate or differentiate in effector cells. Prolonged treatment with mitomycin-C may also induce apoptosis. The responder PBMCs produced alloantibodies against the mitomycin-C treated stimulator cells.
To determine humoral alloimmunity, de novo alloantibodies production is induced in one-way MLR. Amount and capacity of generated alloantibodies in the supernatant of each one-way MLR are assessed using an antibody-mediated cell dependent cytotoxicity (CDC) assay. The principle of the assay relies on the assessment of cellular survival of resting PBMCs isolated from the same donors, supplemented with the alloantibodies from the one-way MLR supernatant (Figure 1). Cell survival was measured by performing a well-known cell proliferation assay [Tetrazolium salt 2,3-bis-(2-methoxy-4-nitro-5-sulfophenyl)-2H-tetrazolium-5-carboxanilide, XTT cell proliferation assay kit].
We avoided direct cytotoxicity assessment through the non-radioactive lactate dehydrogenase release assay (LDH) mainly because cellular cytotoxicity inevitably occurs in the context of MLRs (Sato et al., 1999) and lactate dehydrogenase is released in the supernatant by dead cells only. Thus, different drug treatments may exert different effects on the MLR mixed cell populations resulting in different amounts of LDH released in the supernatant, a fact that would provide erroneous data in the subsequent antibody-mediated CDC assay. 
Figure 1. Assessment of humoral alloimmunity in mixed-lymphocyte reaction. The flow-chart depicts the protocol to assess humoral alloimmunity using MLR. In one-way MLR, responder PBMCs produce alloantibodies against the mitomycin-C treated stimulator cells. Blood A and B correspond to blood donors A and B, respectively. Blood B sample will be used to generate the resting cells of that donor that will constitute the responder cells against the mitomycin-C treated stimulator cells of Blood A. Resting cells, or responder cells of Blood A will be used against the mitomycin-C treated stimulator cells of another blood donor’s sample (e.g., Blood B). Quantitative measurement of alloantibodies in the supernatant of the MLR is assessed by adding this supernatant to resting PBMCs (from the same individual who provided the stimulator PMBCs used for the MLR) using an antibody-mediated CDC assay and measuring cell viability with XTT assay.
Materials and Reagents
- Filtered P1000 tips DF1000ST (Gilson, catalog number: F171705)
- Filtered P200 tips DF200ST (Gilson, catalog number: F171503)
- Filtered P20 tips DF30ST (Gilson, catalog number: F171303)
- Cover glasses 22 x 22 mm (VWR, catalog number: 631-0125)
- Serological pipette 10 ml sterile (Greiner Bio-One, catalog number: 607180/10ml)
- Cell culture 6-well plate, flat bottom, tissue culture treated (Eppendorf, catalog number: 0030720113)
- Cell culture 24-well plate, flat bottom, tissue culture treated (Eppendorf, catalog number: 0030722116)
- Cell culture 96-well plate, flat bottom, tissue culture treated (Eppendorf, catalog number: 0030730119)
- K3-EDTA vacuum blood collection tube (C.D.RICH, catalog number: CDRP007)
- Conical tubes 15 ml (Corning, catalog number: CLS430790)
- Conical tubes 50 ml (Corning, catalog number: CLS430828)
- Pasteur Plastic Pipettes, 3 ml, sterile (Fisher Scientific GmbH, catalog number: 3124975)
- Microcentrifuge Tube, 1.7 ml, Snap Cap, Clear, sterile (Corning B.V., catalog number: 3621)
- Sterile blood syringes, 20 ml (Nipro, catalog number: 200812)
- Low-Tox-H rabbit complement (Cedarlane, catalog number: CL3331)
- Ficoll Histopaque-1077 (Sigma-Aldrich, Merck Millipore, catalog number: 10771-100ML)
- Trypan blue solution (Sigma-Aldrich, Merck Millipore, catalog number: T8154-100ML)
- RPMI 1640 cell culture medium (Sigma-Aldrich, Merck Millipore, catalog number: R7388-100ML)
- RPMI 1640 cell culture medium without phenol red (Gibco, catalog number: 11835-063 500ML)
- Fetal bovine serum (Sigma-Aldrich, Merck Millipore, catalog number: F2442-500ML)
Note: The fetal bovine serum has to be heat inactivated at 55 °C for 1 h. - Antibiotic-antimycotic solution 100x (Sigma-Aldrich, Merck Millipore, catalog number: A5955)
- Dulbecco’s phosphate buffer saline (Sigma-Aldrich, Merck Millipore, catalog number: D8537-500ML)
- XTT cell proliferation assay kit (Trevigen, catalog number: 4891-025-K)
- Mitomycin C (Sigma-Aldrich, Merck Millipore, catalog number: M0503)
Equipment
- Pipetman Starter kit (Gilson, catalog number: F167300)
- CappTempo motorized pipette controller (Capp, catalog number: T100-B)
- Cell Counting Chamber Neubauer (Marienfeld, catalog number: 0640010)
- Safety Cabinet Safeflow 1.2 (BioAir, catalog number: LD60000)
- CO2 Incubator (Heraeus, model: BB6220)
- Inverted Microscope (Zeiss, model: Axiovert 40C)
- Varispeed Refrigerated Centrifuge (Centurion Scientific, model: K280R)
- Heated Bath Circulator (Lauda, model: M20)
- Refrigerator Cell Plate Centrifuge (Eppendorf, model: 5810R)
- Multimode Plate Reader (Perkin Elmer, model: Enspire)
- Microcentrifuge (Eppendorf, model: 5415 R)
- Any Styrofoam box that will serve as an ice bucket
Software
- The SPSS software (version 13; SPSS Inc, Chicago, IL, USA) (for statistical analysis of the data)
Procedure
- Blood collection
Notes:- To exclude pre-sensitization events, male subjects without blood transfusion history or previous organ transplantation are preferred. Subjects must be healthy and not related.
- A written informed consent must be obtained from each individual enrolled in the study.
- The ethics committee of your respected institute/country must formally approve your study, providing also a study protocol number.
- The whole procedure requires aseptic conditions, the use of sterile reagents and plastic ware is highly recommended.
- With a sterile blood syringe, draw about 20 ml of blood from each individual and immediately dispense it into a pre-labeled K3-EDTA vacuum blood collection tube (one tube/donor).
- Invert the tube several times for the blood to mix properly with K3-EDTA and, keep the collection tubes in a cold environment (4 °C, incubation time can be no longer than 2 h) until PBMCs isolation commences.
- PBMCs isolation
Notes:- The whole procedure must be performed inside a sterile cabinet (hood).
- Whether you use a swinging bucket centrifuge rotor, or a rotor with a fixed angle will not significantly affect the total numbers of isolated PMBCs.
- During Step B1, layering of blood on the Ficoll Histopaque should be done really gently in order not to disturb the Ficoll layer (Ficoll allows for elimination of dead cells).
- Following these instructions, a proper isolation should yield 1.8-2.3 x 106 PBMCs/1 ml of blood of a healthy individual.
- The ratio of blood to Ficoll Histopaque volume should be 1:1.
- Dependent on the volume of blood collected, add the same total volume of Ficoll into a 50 ml conical tube per donor. Slowly add the blood of each individual with a plastic Pasteur pipette at the surface of the Ficoll without disturbing it.
- Immediately centrifuge the tubes containing Ficoll and blood at 400 x g for 25 min at room temperature (RT). Make sure that the deceleration speed of the centrifuge machine is 0.
- Using a sterile plastic Pasteur pipette carefully breach the plasma layer and, without disturbing the Ficoll Histopaque layer, aspirate the white PBMCs layer (“buffy” coat) formed in the interphase between blood plasma and Ficoll Histopaque (Figure 2). Pour the PBMCs into a new 15 ml conical tube.

Figure 2. Isolation of PBMCs. Layers of Ficoll Histopaque and whole blood prior to (A) and after (B) centrifugation showing the separation of layers in the conical tube. - Add RPMI 1640 cell culture medium without phenol red up to 15 ml for each PBMCs collected.
- Pellet PBMCs by centrifuging samples at 315 x g for 10 min at RT. Revert deceleration speed to normal (6 on Centurion Scientific centrifuge).
- Remove the supernatant, wash cell pellet with 10 ml of RPMI 1640 cell culture medium without phenol red and centrifuge samples at 315 x g for 10 min at RT.
- Remove the supernatant and resuspend each cellular pellet with 2 ml of complete RPMI medium (10% FBS, 1% antibiotic-antimycotic solution) and count them using Neubauer chamber. Mix 90 μl of trypan blue solution and 10 μl of cells into a clean Eppendorf tube, load the Neubauer chamber and begin counting immediately under a microscope at the lowest magnification. Prolonged overstaining with trypan blue solution stains all cells blue. Always count the viable cells that appear as spherical white cells. Use of trypan blue allows for distinction of dead cells from live cells, since dead cells are stained in blue. A detailed protocol for cell counting using a Neubauer chamber was discussed in (Phelan and Lawler, 2001).
- Assessment of humoral alloimmunity
Notes:- As already mentioned, a 7-day incubation period in Step C1f is necessary for the development of IgM and IgG alloantibodies. Any drug treatment should last for the entirety of the period.
- During Step C3a, seed the appropriate number of cells that you will need to generate enough wells to assess humoral immunity in non-treated cells, treated cells and control cells. We recommend to perform nine to twelve identical experiments, each in triplicate, and the results should refer to the mean of the three measurements.
- Low-Tox-H rabbit complement is provided as a lyophilized compound. Shortly before the use of Low-Tox-H rabbit complement, reconstitute it with 1 ml of ice-cold distilled water (reconstitute during the 30 min incubation on ice during Step C3f). According to the manufacturer's instructions, the complement must be stored at 4 °C for a maximum of one hour after reconstitution.
- Have access to an ELISA plate reader capable of reading wavelengths at 490 nm and 630 nm, respectively.
- Make sure to remove any bubble formed in the XTT 96-well plate before reading the plate with an ELISA plate reader machine. Any foam or bubble will negatively impact OD readings. The tip of a syringe works best for bubble or foam removal.
- Preparation of one-way MLR
- Resuspend 0.5 x 106 PBMCs in 500 μl with complete RPMI from each sample and transfer them into a sterile 1.7 ml microcentrifuge tube. Treat them with 50 μg/ml mitomycin C for 30 min at 37 °C. These are the stimulator cells.
- Centrifuge cells at 300 x g for 5 min at RT, remove supernatant and wash cells with Dulbecco’s phosphate buffer saline.
- Repeat Step C1b two more times.
- Seed 0.5 x 106 stimulator cells/well resuspended in 250 μl complete RPMI into a 24-well plate.
- To each well already containing the 0.5 x 106 stimulator cells, add 0.5 x 106 resting PBMCs (resuspended in 250 μl complete RPMI) from a different donor; these are the responder cells (e.g., stimulator cells from donor A with responder cells from donor B). Together stimulator and responder cells constitute the MLR.
- Incubate MLR for 7 days at 37 °C in a CO2 incubator with or without the treatment that is under your investigation. The treatment of choice should be added right at the start of the 7-day incubation period. Culture media must not be changed during the 7-day incubation.
- Preparation of resting PBMCs
- In parallel to the preparation of one-way MLR (Steps C1a-C1f), resting PBMCs samples should be prepared. Resuspend 1 x 106 PBMCs in 2 ml of complete RPMI and transfer the cells into one well of 6-well plate. Prepare as many wells as needed. These cells are the resting PBMCs.
- Incubate resting PBMCs for 7 days at 37 °C in a CO2 incubator, without changing the culture media during the 7-day incubation period.
- Antibody-mediated complement-dependent cytotoxicity
- Once the 7-day period is over, count the viable resting PBMCs from each well. Counting of cells from each resting PBMCs sample is laborious. It has to be done for each sample, as it will ensure accurate counting of resting cells, excluding proliferation or apoptotic rates during the 7-day period that may greatly vary between PBMCs populations from different blood donor individuals.
- Following counting, resuspend 0.5 x 105 resting PBMCs into 50 μl of complete RPMI and seed 50 μl of cells/well into a flat bottom 96-well plate. The above resting PBMCs must be derived from the same individual that also provided the stimulator PBMCs of the one-way MLR (as seen in Figure 1, e.g., PBMC A individual). Humoral alloimmunity will be evaluated in the person who provided the responder PBMCs (as seen in Figure 1, e.g., responder PBMC B individual).
- Once the 7-day period is over, centrifuge each 24-well plate containing the MLR, from Step C1f, on a Cell Plate Centrifuge at 300 x g for 10 min at RT.
- Collect the supernatant from each MLR. The volume of the supernatant should be around 450-500 μl. Please be gentle during handing and pipetting, in order not to accidentally pick any centrifuged cells.
- Add 50 μl from each MLR supernatant or 50 μl of complete RPMI (control) into the wells of that 96-well plate containing the resting target PBMCs. In addition, in an empty well add 100 μl of complete RPMI that will serve as the blank well for the XTT assay. It is recommended to generate nine to twelve control wells for statistical analysis.
- Incubate the 96-well plate on ice for 30 min inside the hood.
- After the 30 min incubation, add 11 μl of rabbit complement Low-Tox-H rabbit complement (10% [v/v], working concentration) in each well (samples, controls, blanks) at a final concentration of 10%. Now the total volume in each well should be 111 μl.
- Incubate the 96-well plate at 37 °C for 2 h in a CO2 incubator.
- Immediately before the end of the incubation period, prepare the XTT working solution from the XTT assay kit that will be used to measure cell survival via colorimetry. Briefly and according to the manufacturer’s protocol, add 100 μl of the XTT activator to 5 ml of XTT reagent in order to prepare the XTT working solution.
- Once the 2 h incubation is over (Step C3i), add 50 μl of the XTT working solution to each well of the 96-well plate and mix gently.
- Return the plate at 37 °C for another hour in the CO2 incubator.
- Read the absorbance of your samples using an ELISA Plate Reader set at 490 nm with a reference correction at 630 nm.
- After subtracting background OD values derived from the blank wells, calculate cell survival in each sample by the equation: Cell survival (%) = (XTT assay OD of the evaluated condition/XTT assay OD of the control) x 100. The lesser the cell survival, the higher the production of alloantibodies in one-way MLR and, consequently the more potent the humoral alloimmunity.
Data analysis
We recommend to perform at least nine to twelve one-way MLRs for each different condition, for instance with or without various tested substances, run in triplicates. Once all cell survival ratios are assessed (Step C3m), you can implement the data into SPSS software. If two conditions are evaluated, paired t-test should be used for comparison of means, since each MLR (consisted of PBMCs from two different donors) is unique in experiments with human subjects. For more than two evaluated conditions one-way repeated-measures ANOVA followed by Bonferroni’s correction test should be used. If you apply this protocol to two different mouse strains, and you consider that subjects of each strain are genetically matching, unpaired t-test and one-way ANOVA followed by the Bonferroni’s correction test respectively are the appropriate statistical tests.
An example of an antibody-mediated CDC is depicted in Figure 3, showing inhibition of humoral immunity by the specific proteasome inhibitor CEP18770 at a concentration of 10 nM. Proteasome plays a key role in cellular protein turnover and protein homeostasis. Moreover, it is well known that plasma cells are susceptible to proteasome inhibition since they produce large amounts of antibodies and consequently exhibit high protein turnover. Direct inhibition of the proteasome in plasma cells leads to impaired protein turnover, eliciting the unfolded protein response, which in its turn will induce apoptosis (Walsh et al., 2012). The presented published results render proteasome as an excellent target for the development of new immunosuppressive medications in the field of transplantation (Eleftheriadis et al., 2017a).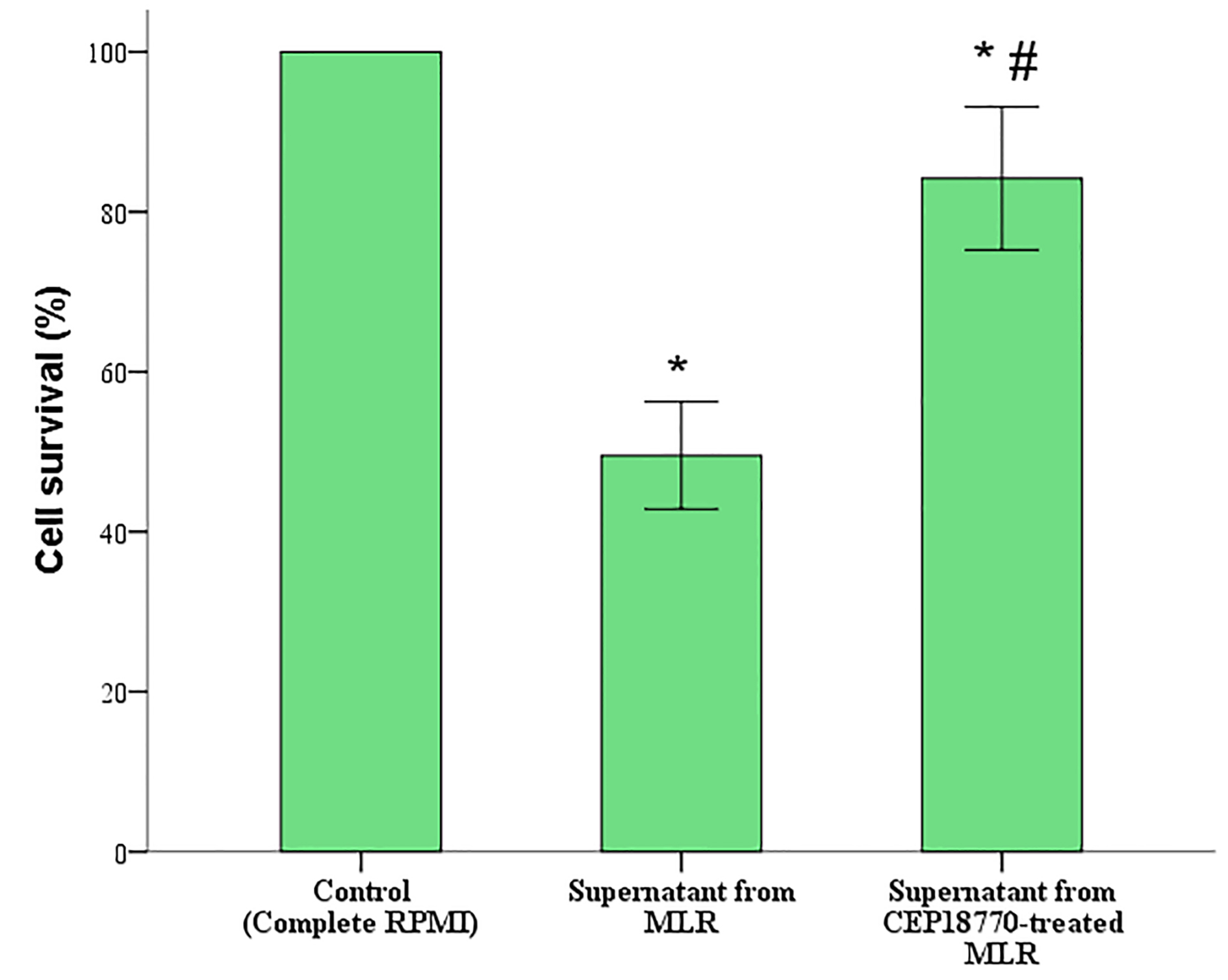
Figure 3. Proteasome inhibitor CEP-18770 efficiently inhibits humoral alloimmunity. One-way MLRs were performed in the presence or not of 10 nM CEP-18770 for a 7-day period. Supernatants were collected and used in an antibody-mediated CDC assay against resting target PBMCs similar to those used as stimulator cells in the respective MLRs. Supernatants derived from CEP-18770-treated MLRs exhibited smaller antibody-mediated CDC than those derived from untreated MLRs. Marks denoted as * and #, indicate a P < 0.05 compared between the conditions. Error bars correspond to SD. Twelve such experiments were performed, each in triplicates and the results refer to the mean of the three measurements.
Notes
Inconsistent variability between ODs during XTT assay should not emerge, and triplicate ensures reproducibility of data. We obtained consistent results by performing this protocol many times. Three studies based on this protocol have already been published (Eleftheriadis et al., 2017a, 2017b and 2017c).
Acknowledgments
The sources of the Nephrology Department, Faculty of Medicine, University of Thessaly, which is funded by the Ministry of Health and the Ministry of Education, Research and Religious Affairs, have funded the development of this protocol.
Competing interests
Declaration of interest: None declared.
Ethics
An informed consent was obtained from each individual enrolled in the study and the Ethics Committee of the University of Thessaly, Faculty of Medicine, Larissa, Greece approved the study protocol (Number of approval: 558/10-2-2017).
References
- Abramowicz, D., Oberbauer, R., Heemann, U., Viklicky, O., Peruzzi, L., Mariat, C., Crespo, M., Budde, K. and Oniscu, G. C. (2018). Recent advances in kidney transplantation: a viewpoint from the Descartes advisory board. Nephrol Dial Transplant 33(10): 1699-1707.
- Eleftheriadis, T., Pissas, G., Antoniadi, G., Liakopoulos, V. and Stefanidis, I. (2017a). A comparative analysis between proteasome and immunoproteasome inhibition in cellular and humoral alloimmunity. Int Immunopharmacol 50: 48-54.
- Eleftheriadis, T., Pissas, G., Mavropoulos, A., Liakopoulos, V. and Stefanidis, I. (2017b). Comparison of the effect of the aerobic glycolysis inhibitor dichloroacetate and of the Krebs cycle inhibitor LW6 on cellular and humoral alloimmunity. Biomed Rep 7(5): 439-444.
- Eleftheriadis, T., Pissas, G., Sounidaki, M., Antoniadi, G., Antoniadis, N., Liakopoulos, V. and Stefanidis, I. (2017c). In human cell cultures, everolimus is inferior to tacrolimus in inhibiting cellular alloimmunity, but equally effective as regards humoral alloimmunity. Int Urol Nephrol 49(9): 1691-1697.
- Konishi, E., Kitai, Y. and Kondo, T. (2008). Utilization of complement-dependent cytotoxicity to measure low levels of antibodies: application to nonstructural protein 1 in a model of Japanese encephalitis virus. Clin Vaccine Immunol 15(1): 88-94.
- Phelan, M. C. and Lawler, G. (2001). Cell counting. Curr Protoc Cytom Appendix 3: Appendix 3A.
- Rumke, H. C., Terpstra, F. G., Huis, B., Out, T. A. and Zeijlemaker, W. P. (1982). Immunoglobulin production in human mixed lymphocyte cultures: implications for co-cultures of cells from patients and healthy donors. J Immunol 128(2): 696-701.
- Sato, T., Deiwick, A., Raddatz, G., Koyama, K. and Schlitt, H. J. (1999). Interactions of allogeneic human mononuclear cells in the two-way mixed leucocyte culture (MLC): influence of cell numbers, subpopulations and cyclosporin. Clin Exp Immunol 115(2): 301-308.
- Walsh, R. C., Alloway, R. R., Girnita, A. L. and Woodle, E. S. (2012). Proteasome inhibitor-based therapy for antibody-mediated rejection. Kidney Int 81(11): 1067-1074.
Article Information
Copyright
© 2019 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Pissas, G. and Eleftheriadis, T. (2019). Assessment of Humoral Alloimmunity in Mixed Lymphocyte Reaction. Bio-protocol 9(2): e3139. DOI: 10.21769/BioProtoc.3139.
Category
Immunology > Immune cell function > Lymphocyte
Cell Biology > Cell isolation and culture > Cell isolation
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










