- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Quantitative Transformation Efficiency Assay for Bacillus subtilis
Published: Vol 8, Iss 23, Dec 5, 2018 DOI: 10.21769/BioProtoc.3109 Views: 9030
Reviewed by: Modesto Redrejo-RodriguezMelanie BerkmanAnna McLoon

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
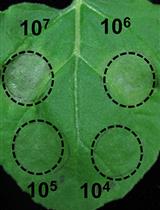

A Fast and Easy Method to Study Ralstonia solanacearum Virulence upon Transient Gene Expression or Gene Silencing in Nicotiana benthamiana Leaves
Wenjia Yu and Alberto P. Macho
Aug 5, 2021 4699 Views
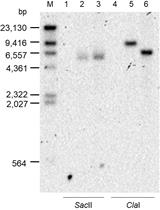
A New Tool for the Flexible Genetic Manipulation of Geobacillus kaustophilus
Ryotaro Amatsu [...] Ken-ichi Yoshida
Sep 5, 2022 1905 Views
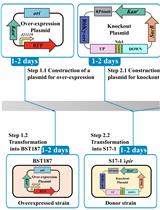
Efficient Genetic Transformation and Suicide Plasmid-mediated Genome Editing System for Non-model Microorganism Erwinia persicina
Tingfeng Cheng [...] Lei Zhao
Mar 20, 2024 2430 Views
Abstract
Bacillus subtilis (B. subtilis) is a model Gram-positive organism used to study a variety of physiological states and stress responses, one of which is the development of competence. Competence refers to the physiological state of a cell which allows it to be transformed naturally. Through induction of competence, the efficiency of natural transformation can be quantified by plating colony forming units (CFU) and transforming units (TFU). Here we describe a protocol for quantifying relative transformability using B. subtilis.
Keywords: Natural transformationBackground
Natural transformation is a type of horizontal gene transfer that involves a bacterium importing exogenous DNA from the environment and recombining that DNA into its genome. This process differs from induced transformation in that naturally transformable bacteria produce proteins that facilitate the import of DNA. Modification of the cell membrane, such as through electroporation or heat shock, is typically essential for induced transformation. Natural transformation can lead to the acquisition of new genetic elements, such as antibiotic resistance or catabolic genes, and may allow bacteria to adapt rapidly to their environment. The various proteins involved with the development of competence, or the physiological state of a cell which allows it to be transformed naturally, and the mechanism of natural transformation are not yet fully understood. Quantitative analysis of transformation efficiency differences between a mutant and wild type provides insight into the contribution and potential functions of the gene in question. Beginning with the description of requirements for transformation of laboratory strains of Bacillus subtilis by Anagnostopoulos and Spizizen (1961), numerous variations on the original media and conditions for transformation, both qualitative and quantitative methods, have been described including but not limited to those by Dubnau and Davidoff-Abelson (1971) and Cutting and Vander Horn (1990). The experimental procedure described here is similar to those prior, focusing on currently available and preferred materials (such as casein hydrolysate, which has been found to produce varying efficiencies based on lot and manufacturer), and includes description of statistical analysis of the results.
Materials and Reagents
- Sterile pipette tips
- Sterile 18 mm glass tubes (Fisher Scientific, catalog number: S63300)
- Sterile 13 mm glass tubes (Fisher Scientific, catalog number: 14-961-27)
- Sterile stainless steel closures (18 mm neck size) (VWR, catalog number: 71000-330)
- Sterile stainless steel closures (13 mm neck size) (VWR, catalog number: 71000-322)
- Sterile 1.7 ml microfuge tubes (Denville Scientific, catalog number: C2170)
- Sterile 60 ml Luer Lock syringes (Fisher Scientific)
- Sterile Acrodisc® 13 mm syringe filters (0.2 µm pore size) (VWR, catalog number: 28142-340)
- Sterile syringe filter disc (0.2 µm pore size) (VWR, catalog number: 28145-477)
- 1.5 ml Cuvettes (Dot Scientific, catalog number: 2410)
- Aluminum foil
- 100 x 15 mm Petri dishes (Crystalgen, catalog number: S-3006)
- Sterile glass beads (diameter 0.2-0.3 mm) (Sigma-Aldrich, catalog number: G1277)
- Sterile streaking sticks (flat toothpicks) (VWR, catalog number: 500029-808)
- Bacillus subtilis (B. subtilis)
Note: This includes the control strains and tested strains. For control strains, B. subtilis subsp. subtilis str. 168 or B. subtilis str. PY79 are typically used. - LB broth (Sigma-Aldrich, catalog number: L3022-1KG)
- LB agar (VWR, DifcoTM, catalog number: 90003-346)
- NaCl (Dot Scientific Inc., catalog number: 7647-14-5)
- KCl (RPI, catalog number: P41025-500.0)
- Na2HPO4 (J.T. Baker, catalog number: 7558-79-4)
- KH2PO4 (VWR, catalog number: 7778-77-0)
- NaOH (Sigma-Aldrich, catalog number: 221465-500G)
- K2HPO4 (Dot Scientific Inc., catalog number: DSP41300-1000)
- Glucose (Amresco Inc., catalog number: 50-99-7)
- Sodium citrate dihydrate (Na3C6O7•2H2O) (VWR, SX00445-1 EMD)
- Casein hydrolysate (EMD, catalog number: 1.02245.0500)
- Potassium glutamate monohydrate (Sigma-Aldrich, catalog number: G1501-100G)
- 100 ng/µl genomic DNA with selectable marker
Note: The marker on the donor DNA should not be present in any of the strains being tested. - MgSO4 (Sigma-Aldrich, catalog number: M7506)
- Ammonium iron citrate (J.T. Baker, catalog number: 1185-57-5)
- Antibiotics (for selection)
- 10x phosphate buffered saline (PBS) (see Recipes)
- 10x competence medium (MC) (see Recipes)
- 1 M MgSO4 (see Recipes)
- 1,000x Ferric Ammonium Citrate (see Recipes)
Equipment
- Pipettes
- Roller drum with disc for 18 mm tubes (New Brunswick, TC-7 style motor with disc for 18mm tubes)
Note: A disc for 16 mm tubes may be used if a disc for 18 mm tubes is unavailable. A 16 mm tube can be used in replacement of the 13 mm tube in an 18 mm tube setup (Figure 1). - Vortexer (VWR, catalog number: 58816-121)
- 37 °C incubator with shelves
- Microcentrifuge capable of 6,200 x g
- UV/vis Spectrometer (Thermo Fisher Scientific, model: 4001/4)
- Autoclave
Procedure
- Preparation of preparatory cultures
- Streak desired B. subtilis cells onto LB agar, with selective antibiotic, when appropriate.
- Incubate plates at 37 °C overnight or RT for 2-3 days.
- In a 13 mm glass tube placed within an 18 mm glass tube, inoculate 1 ml of LB culture with a single colony of B. subtilis approximately 12 h before starting preculture and grow the cells overnight at RT (Figure 1).
Note: This setup helps with transferring liquid cultures, as some pipettes may not reach the bottom of the 18 mm glass tube.
Figure 1. Two possible ways to set up a 13 mm tube within an 18 mm tube. In setup 1, the end of a 1 ml pipette tip is cut off and inserted into the 18 mm tube, with the 13 mm tube set on top of the tip. In setup 2, two 13 mm glass tube caps are placed on top of each other and placed into the bottom of the 18 mm tube.
- Transformation assay setup
- Grow cells rolling at 37 °C for one hour.
Note: Roller drum should be set to 9 RPM. - Transfer 1 ml of each culture to a 1.7 ml microfuge tube.
- Harvest cells by centrifugation (6,200 x g; 2 min, room temperature). There should be a visible pellet at the bottom of the microfuge tube.
- Pipette to remove supernatant and discard.
- Resuspend cells in 1 ml of 1x PBS.
- Centrifuge again (6,200 x g; 2 min, room temperature).
- Pipette to remove supernatant and discard.
- Resuspend cells in 1 ml of 1x PBS.
- Centrifuge again (6,200 x g; 2 min, room temperature).
- Discard supernatant and resuspend cells in 500 µl of freshly made 1x MC (see Recipes).
- Check OD600, using 150 µl of resuspended cells and 450 µl of 1x PBS (Blank with 150 µl of 1x MC and 450 µl of 1x PBS).
Note: OD600 should be 0.2-1.2. - Inoculate washed cells into a new 13 mm tube in an 18 mm tube and fill to 1 ml of 1x MC, so the culture has a starting OD600 of 0.01 to 0.05.
Note: 1 µl of the selective antibiotic pertaining to the original strain resistance (not the donor DNA antibiotic marker) can be added to prevent any accidental contamination. - Grow cells while rolling at 37 °C for 4.5 to 5 h.
- After 4.5 to 5 h, transfer 300 µl of each culture to a new 13 mm glass tube.
- Add 3 µl of genomic DNA (at a concentration of 100 ng/µl) and 3 µl of 1 M MgSO4 to each culture tube.
Note: Include one culture without DNA added as a control for each experiment. - Continue rolling cells at 37 °C for 2 h.
- Grow cells rolling at 37 °C for one hour.
- Serial dilutions and plating
- Retrieve cultures from 37 °C.
- Create 1:10 serial dilutions of the cultures in 1x PBS to a 10-6 dilution.
- Plate 100 µl of desired dilutions onto LB (for CFU) and selective plates (for TFU) using glass beads (Figure 2).
Note: A 10-6 dilution should yield 30-300 colonies for the wild-type TFU. For mutants, dilution plating is dependent on the type of defect present, so initial experiments are necessary to determine the range of dilutions required for plating.
Figure 2. Representation of how the plating may look when comparing wild-type cells to mutants. In Row 1, the wild-type and ∆lacA strains are plated at a 10-2 dilution. The ∆comK and ∆ecsB strains are plated with undiluted culture. All plates in Row 1 contain the selectable antibiotic relevant to the marker present on the donor DNA. Row 2 is plated on LB plates with 10-6 dilutions. Each column is a different genotype. Column 1 is wild type, column 2 is ∆comK (a gene necessary for transformation), column 3 is ∆lacA (a gene not related to natural transformation), and column 4 is ∆ecsB (putatively involved with natural transformation). No colonies formed on the undiluted ∆comK selective plate, a few colonies are visible on the ∆ecsB selective plate, and colony formation was at the level of wild type for ∆lacA. CFU can also can be compared to wild type at this step. - Place plates at 37 °C overnight, or leave at RT for 2-3 days.
Note: TFU plates may need an extra day for colonies to grow due to selection. - Count colonies on each plate. The colonies on each plate must be multiplied by their dilution factor to obtain CFUs and TFUs.
Note: It may be helpful to count using a marker to mark colonies while using a tally counter to keep track of the number of colonies.
Data analysis
- Perform the quantitative transformation efficiency assay a minimum of three times per strain, and each time plate duplicates of each dilution. A positive control (wild-type strain) and negative control (wild type with no donor DNA added) should also be implemented during each replicate.
Note: Each replicate is performed using different colonies from the same plate. - For data analysis, transformation efficiency is calculated using the formula ηS = Cr/(CTpD), where Cr is the average number of TFU (transformants), CT is the average total number of CFU, and pD is the DNA concentration in micrograms per milliliter, from biological replicates. Each replicate is normalized to WT and run at the same time. The limit of detection of the assay is 0.5 transformants/CFU/µg of genomic DNA (as in Chilton et al., 2017; for PY79 with transforming DNA selected with 100 µg/ml spectinomycin).
- For statistical analysis, relative efficiency values are converted to arcsin values with the following equation to allow for normally distributed input for Dunnett’s multiple-comparison test (for comparison of all mutants together with the control): DS = [180 sin-1 (ηS/6)]/π. Dunnett's multiple-comparison test is performed on converted data with the multcomp package in the R statistical analysis software. Relative transformation efficiencies of WT and mutants are log transformed to generate normally distributed data, and t-tests comparing each mutant pairwise with corresponding controls are used to test for significant differences (as in Chilton et al., 2017).
Note: When testing a single strain against WT, an ANOVA or t-test can be used.
Recipes
- 10x phosphate buffered saline (PBS) (0.5 L, pH 7.4)
40 g NaCl
1 g KCl
7.26 g Na2HPO4 (anhydrous)
1.2 g KH2PO4
ddH2O to 0.4 L
Dissolve solids
Adjust pH to pH 7.4 with 50% NaOH
Bring total volume to 0.5 L with ddH2O
Sterilize by autoclaving
Store at room temperature
Dilute 1:10 in ddH2O to obtain 1x PBS
Note: ddH2O provides consistency from batch to batch compared to dH2O. - 10x competence medium (MC) (100 ml) (as in Konkol et al., 2013)
10.7 g K2HPO4
5.2 g KH2PO4
20 g Dextrose anhydrous (glucose)
0.88 g Na3C6O7•2H2O (sodium citrate dihydrate)
1 ml 1,000x Ferric Ammonium Citrate (2.2% stock) (see below)
1 g Casein Hydrolysate EDM
2.2 g Potassium Glutamate monohydrate
ddH2O to 100 ml
Mix thoroughly; filter sterilize using 0.2 µm filter and syringe
Store at -20 °C
Note: To make 1 ml of fresh 1x MC, dilute 100 µl 10x MC into 900 µl ddH2O. Once the final concentration of 1 ml is obtained, add 3 µl of 1 M MgSO4. - 1 M MgSO4 (0.5 L)
60 g MgSO4 (Magnesium Sulfate Anhydrous)
ddH2O to 0.5 L
Sterilize by autoclaving
Store at room temperature - 1,000x Ferric Ammonium Citrate (100 ml)
2.2 g Ammonium iron citrate
ddH2O to 100 ml
Filter sterilize using screw cap filter
Wrap in aluminum foil
Store at room temperature
Note: This solution is light sensitive.
Acknowledgments
This protocol was adapted from Chilton et al. (2017). This work was funded in part by a grant from the Rita Allen Foundation.
Competing interests
The authors have no conflicts of interest or competing interests with respect to this work.
References
- Anagnostopoulos, C. and Spizizen, J. (1961). Requirements for transformation in Bacillus subtilis. J Bacteriol 81(5): 741-746.
- Chilton, S. S., Falbel, T. G., Hromada, S. and Burton, B. M. (2017). A conserved metal binding motif in the Bacillus subtilis competence protein ComFA enhances transformation. J Bacteriol 199(15): e00272-17.
- Cutting, S. M. and Vander Horn, P. B. (1990). Genetic Analysis. In: Harwood, C.R. and Cutting, S. M. (Eds.). Molecular Biological Methods for Bacillus. John Wiley & Sons, Ltd., West Sussex, UK. pp 27-74.
- Dubnau, D. and Davidoff-Abelson, R. (1971). Fate of transforming DNA following uptake by competent Bacillus subtilis. I. Formation and properties of the donor-recipient complex. J Mol Biol 56(2): 209-221.
- Konkol, M. A., Blair, K. M. and Kearns, D. B. (2013). Plasmid-encoded ComI inhibits competence in the ancestral 3610 strain of Bacillus subtilis. J Bacteriol 195(18): 4085-4093.
Article Information
Copyright
© 2018 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Loyo, C. L. and Burton, B. M. (2018). Quantitative Transformation Efficiency Assay for Bacillus subtilis. Bio-protocol 8(23): e3109. DOI: 10.21769/BioProtoc.3109.
Category
Microbiology > Microbial physiology > Competence
Microbiology > Microbial genetics > Transformation
Molecular Biology > DNA > Transformation
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link








