- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
One-step Derivation of Functional Mesenchymal Stem Cells from Human Pluripotent Stem Cells
Published: Vol 8, Iss 22, Nov 20, 2018 DOI: 10.21769/BioProtoc.3080 Views: 9927
Reviewed by: Antoine de MorreeShweta GargAnthony Flamier

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Spheroid Sheets: A Scalable Platform for Producing Tissue Membrane Constructs
Quang Bach Le [...] Deepak Choudhury
Nov 20, 2025 1536 Views
A Protocol to Induce Brown and Beige Adipocyte Differentiation From Murine and Human Adipose-Derived SVF
Rohit Raj Yadav [...] Narendra Verma
Dec 5, 2025 1643 Views
Optimization of Adipogenic Differentiation Protocol for Murine and Human Cell Culture Models
Junwan Fan [...] Wenyan He
Jan 20, 2026 214 Views
Abstract
Mesenchymal stem cells (MSCs) are invaluable cell sources for understanding stem cell biology and potential application in tissue engineering and regenerative medicine. The current issues of MSCs that demand to be further addressed are limited donors, tissue sources and limited capacity of ex vivo expansion. Here, we describe a simple and easy protocol for generating functional mesenchymal stem cells from human pluripotent stem cells (hPSCs) via one-step low glucose medium switch strategy in feeder-free culture system. In this protocol, human induced pluripotent stem cells (hiPSCs) and H9 human embryonic stem cells (hESCs) were successfully differentiated into MSCs, named hiPSC-MSCs and hESC-MSCs, respectively. The derived hiPSC-MSCs and hESC-MSCs exhibited common MSC characteristics as MSCs derived from human bone marrow (hBM-MSCs), including expressing MSC surface markers and possessing capability of tri-lineage differentiation in vitro (adipogenesis, osteogenesis and chondrogenesis). As compared with other available protocols, our protocol can be applied to generate a large number of MSCs from hPSCs with high efficiency, low-cost manner, moreover, not involving embryoid body, mouse feeder-cell, flow sorting, and pathway inhibitors (such as SB203580 and SB431542). We believe that this protocol could provide a robust platform to reach the future demand for producing the industrial scale of MSC from hPSCs for autologous cell-based therapy.
Keywords: Mesenchymal stem cells (MSCs)Background
Mesenchymal stem cells (MSCs) are adult stem cell populations derived from postnatal tissues. They retain fast proliferation rate, self-renewal capabilities, and lineage-specific differentiation potential. Moreover, MSCs are capable of producing growth factors, having anti-apoptosis properties, lacking carcinogenicity (Trounson and McDonald, 2015), and no adverse effects noted thus far in clinical trials (Lalu et al., 2012). Therefore, MSCs represent promising cell candidates for stem cell-based therapy and regenerative medicine (Tuan et al., 2003; Chen et al., 2008). Functional MSCs have been well established from various tissues, including bone marrow (BM) (Noth et al., 2002), adipose tissue (Zuk et al., 2001), peripheral blood (da Silva Meirelles et al., 2006), umbilical cord blood (Lee et al., 2004; Schuh et al., 2009), and other human organs (Campagnoli et al., 2001; Crisan et al., 2008). Among them, BM is regarded as the most accessible and reliable source for MSCs in adults, considering the wide distribution and well-characterized isolation methodology (Mezey et al., 2000). MSCs derived from human bone marrow (hBM-MSCs) have been clinically applied for a variety of injuries/disorders. Allogeneic hBM-MSCs transplantation-based tissue regeneration provides the possibility to re-establish bone-associated elements (Horwitz et al., 1999; Ghasroldasht et al., 2014). And autologous hBM-MSC transplantation could increase cartilage formation and repair the cartilage defects (Wakitani et al., 2007; Wu et al., 2013). However, the potential therapeutic applications of hBM-MSCs have been hindered by limited number of cell acquisitions, the limited cell long-term expandability in vitro, high cell heterogeneity and invasive procedure of cell isolation from human donors (Kretlow et al., 2008; Uccelli et al., 2008). Therefore, obtaining large quantities of functional MSCs from limited human donors is a challenge.
The large population of functional MSCs derived from human pluripotent stem cells (hPSCs) by in vitro differentiation, including human embryonic stem cells (hESCs) and human induced pluripotent stem cells (hiPSCs) could be an alternative to hBM-MSCs for therapeutic applications. MSCs derived from hESCs and hiPSCs (hESC-MSCs and hiPSC-MSCs) can overcome potential barriers for clinical therapeutic applications of hBM-MSCs, without capacity and function decline (Brown et al., 2014; Hynes et al., 2014). First, hPSCs represent infinitive cells ex vivo and in vivo, thus providing renewable cell sources for MSC generation with a high yield from various persons, without invasive procedure. Secondly, due to the reduced immune rejection and no tumorigenesis, MSCs derived from hPSCs can be used for autologous transplantation and personalized cell therapy (Peng et al., 2016; Gao et al., 2017). Thirdly, there has a great advantage to use isogenic hMSCs for autologous MSCs transplantation, in which the hPSCs can be genetically modified by endonuclease on patient PSCs with genetic mutations, and then differentiated into isogenic hMSCs. So far, hESCs are already used to generate clinically compliant MSCs, which displayed similar gene expression profiles and function as hBM-MSCs (Lian et al., 2007). Meanwhile, hiPSC-MSCs can significantly relieve hind-limb ischemia, in which hiPSC-MSCs were even more beneficial than hBM-MSCs due to superior survival and engraftment capability (Lian et al., 2010).
How to rapidly produce a large quantity of genetically identical MSCs from hPSC as a substitute for hBM-MSCs? Simple methods have been reported to effectively induce hPSCs into functional MSCs. Identical batches of hESC-MSCs can be reproducibly generated from hESCs in differentiation medium supplemented with 5 ng/ml FGF2 and 5 ng/ml PDGF-AB (Lian et al., 2007). Camilla and the colleagues successfully established mesenchymal progenitors from hESCs by single cell passage within high glucose medium and 10 ng/ml bFGF (Karlsson et al., 2009). MSCs can also be differentiated from hiPSCs in knockout medium containing 10 ng/ml bFGF, 10 ng/ml PDGF-AB, and 10 ng/ml epidermal growth factor (Lian et al., 2010). Furthermore, MSCs can be generated from hiPSCs via embryoid body formation with p38-MAPK inhibitor SB203580 treatment, in which hiPSC-MSCs could have less tumorigenicity (Wei et al., 2012). Our team previously developed a simple and efficient method to derive hiPSC-MSCs by culturing hiPSCs in low glucose medium for 2 weeks, and a serial passaging without embryoid body formation and flow sorting (Zou et al., 2013; Kang et al., 2014). However, a monolayer of feeder cells involved in the induction process has been a risk factor for therapeutic application (Kang et al., 2015). Another single step method to transform hESCs/hiPSCs into MSCs is to culture hESCs/hiPSCs in serum-free medium supplemented with TGF-β pathway inhibitor SB431542 for 10 days. This differentiation process was in an embryoid body independent and feeder cell-free manner. However, these hESC/hiPSC-MSCs showed limited adipogenic differentiation capacity, and this induction procedure was SB431542 dependent (Chen et al., 2012). In 2015, Fei Liu’s group also described a modified protocol to improve the differentiation of hiPSCs toward MSCs in mTESR1 medium supplemented with TGFβ inhibitor SB431542 in 7.5% CO2 incubation, followed by repeatedly passaging cells without flow cytometric sorting. 88% of derived hiPSC-MSCs were positive for MSC surface markers, but the 7.5% CO2 condition is not universal for cell culture, and SB431542 is needed for this induction process (Zhao et al., 2015).
Different methods for deriving functional MSCs from hPSCs have their own advantages, such as a simple protocol, or high efficiency. However, some current methods still have technical limitations regarding the MSCs production, including being time-consuming (more than 1.5 months), having less adipogenic differentiation capability of cells, and requiring additional chemicals or uncommon culture condition. Therefore, here, we updated a simple, efficient method for differentiating hESCs/iPSCs into MSCs. This protocol is free of embryoid body formation and feeder cell, approximately takes 28-30 days to generate high-quality MSCs. This system would provide a promising platform for generating a large scale of hiPSC-MSCs/hESC-MSCs from single donor for regenerative medicine.
Materials and Reagents
- Materials
- 15 ml and 50 ml Falcon tubes (Falcon, catalog numbers: 352097, 352098)
- 1.5 ml and 2 ml centrifuge tubes (Eppendorf, catalog numbers: 0030120086, 0030120094)
- 96-mutiwell, 24-well, and 6-well tissue culture plates (Corning, catalog numbers: 3599, 3524, 3516)
- T25, T75 and T175 tissue culture flasks (Thermo scientific, catalog numbers: 156367, 156499, 159910)
- 1 ml, 5 ml, 10 ml and 25 ml disposable plastic serological pipettes (Corning, catalog numbers: 4012, 4051, 4101, 4251)
- Pipette tips (10, 200 and 1,000 μl) (BBI, catalog numbers: F602215-0001, F604224-0001, F600222-0001)
- 0.22 μm and 0.45 μm disposable sterile filters (Millipore, catalog numbers: SLGV033RB, SLHV033RS)
- 1.5 ml cryogenic vials (Nalgene, catalog number: 5000-1020) and cryovial storage racks
- Cells
- Human induced pluripotent stem cells (hiPSCs, generated in our laboratory)
- Human embryonic stem cells (H9, hESCs) (WiCel Research Institute, catalog number: WA09; a gift from Associate Professor Yonglun Luo, Aarhus University)
- Human BM-MSC (hBM-MSCs, LONZA, catalog number: PT-2501)
- Antibodies
Note: The antibodies are used to validate MSC surface antigens.- APC Mouse Anti-Human CD34 (BD, catalog number: 560940)
- V450 Mouse Anti-Human CD45 (BD, catalog number: 560368)
- PE Mouse Anti-Human CD73 (BD, catalog number: 561014)
- FITC Mouse Anti-Human CD90 (BD, catalog number: 555595)
- PerCP-Cy5.5 Mouse Anti-Human CD105 (BD, catalog number: 560819)
- Reagents
- TeSR-E8 medium (Stem Cell Technologies, catalog number: 05990)
- Vitronectin XF (Stem Cell Technologies, catalog number: 07180)
- Cell Adhere Dilution Buffer (Stem Cell Technologies, catalog number: 07183)
- DPBS without calcium and magnesium (1x, DPBS w/o Ca2+ and Mg2+) (HyClone, catalog number: SH30028.02)
- ReLeSR (Stem Cell Technologies, catalog number: 05872)
- Rock inhibitor Y-27632 (Stem Cell Technologies, catalog number: 72302)
- Dimethyl Sulfoxide (DMSO) (Sigma-Aldrich, catalog number: D5879)
- DMEM-low glucose (1 g/L) (Gibco, catalog number: 31885023)
- Highly Qualified Fetal Bovine Serum (HQ-FBS) (Gibco, catalog number: 26140079)
- GultaMAX (100x) (Gibco, catalog number: 35050061)
- Non-Essential Amino Acids Solution (NEAA, 100x) (Gibco, catalog number: 11140050)
- Penicillin-Streptomycin (100x) (Gibco, catalog number: 15140122)
- 0.05% Trypsin-EDTA (1x) (Gibco, catalog number: 25300062)
- Gelatin (Sigma-Aldrich, catalog number: G1890)
- Autoclaved H2O (made in our laboratory)
- 75% Ethanol (vol/vol) (made in our laboratory)
- Isopropanol (Fisher Scientific, catalog number: A416-1)
- 37% Formaldehyde (vol/vol) (Polysciences, catalog number: 00625-1)
- StemPro Osteogenesis Differentiation Kit (Gibco, catalog number: A1007201)
- StemPro Chondrogenesis Differentiation Kit (Gibco, catalog number: A1007101)
- StemPro Adipogenesis Differentiation Kit (Gibco, catalog number: A1007001)
- Alizarin Red S (Sigma-Aldrich, catalog number: A5533)
- Toluidine blue (Sigma-Aldrich, catalog number: 89640)
- Oil Red O solution (Sigma-Aldrich, catalog number: O1391)
- Paraffin (Leica Biosystems, catalog number: 39601095)
- Complete TeSR-E8 Medium for hiPSCs and hESCs (see Recipes)
- Vitronectin XF coated culture vessels (see Recipes)
- Rock inhibitor (100x Stock solution, see Recipes)
- TeSR-E8 freezing medium (see Recipes)
- MSC low glucose medium (see Recipes)
- 1% (wt/vol) gelatin stock solution (see Recipes)
- 0.1% gelatin-coated vessels (see Recipes)
- 2x MSC freezing medium (see Recipes)
- 4% formaldehyde solution (see Recipes)
Equipment
- Pipette aid (Eppendorf, catalog number: 4430000018)
- Mechanical Palm Click Counter (Jinnan, model: JN28-HP)
- Inverted phase-contrast microscope (x4, x10, x20, x40 objectives) (Olympus, model: CKX31)
- Tissue culture centrifuge with multiple rotors (Eppendorf, model: 5401000060)
- Humidified CO2 incubators (Heal Force, model: HF90)
- -80 °C freezers (Thermo Scientific, model: Forma 900)
- Liquid nitrogen tank (Chart Biomedical, model: YDS-50B-125)
- Isopropanol cell freezing container (Nalgene, model: 5100-0001)
- Water bath (Shanghai Yiheng, model: HWS-26)
- Hemocytometer (Marienfeld, model: AP-0650030)
- Vacuum apparatus (Sciencetool, model: BV240)
- Biosafety cabinet (Suzhou Antai Air Technology, model: BSC-1304IIA2)
- Flow cytometer (BD Biosciences, model: LSR Fortessa)
- -20 °C freezer (Homa)
Software
- Flow Jo (Version 10)
- Image J (Version 1.5.1)
Procedure
- Thawing and recovering human iPSCs or human ESCs (hiPSCs/hESCs) (6-well plate format)
- Pre-warm a required volume of TeSR-E8 medium to room temperature in a 50 ml Falcon tube (see Note 1).
- Prepare an appropriate quantity of Vitronectin XF coated 6-well plates at least 1 h before cell thawing (see Recipes).
- Remove the cryogenic vial of frozen hiPSCs/hESCs from liquid nitrogen quickly and immerse the vial into a 37 °C water bath.
- Hold the cap to constantly swirl cryogenic vial without submerging the cap.
- When most cells are thawed/only a few ice crystals remain, take the cryogenic vial out of the water bath, spray the cryogenic vial with 70% ethanol and move the vial into biosafety culture cabinet.
- Open the cap, and then gently transfer the cell suspension into a 15 ml Falcon tube containing TeSR-E8 medium prepared before (see Note 2).
- Rinse the cryogenic vial with 1 ml TeSR-E8 medium and add the solution into the same Falcon tube, gently mix cells by pipetting.
- Centrifuge at 300 x g for 5 min at room temperature, and then discard the supernatant.
- Resuspend the hiPSC/hESC pellets with 2 ml TeSR-E8 medium by gently pipetting up and down several times.
- Add 20 μl ROCK inhibitor (100x, stock solution) to the cell suspension for increasing survival of hPSCs (see Note 3).
- Discard the excess Vitronectin XF from the pre-coated 6-well plate, and slowly transfer hiPSC/hESCs suspension into the plate. One cryogenic vial of hiPSC/hESC can be plated into 1-2 wells of the 6-well plate.
- Gently move the culture plate back and forth, left and right motions several times to disperse cells across the whole well (see Note 4).
- Place the plate in a 37 °C, 5% CO2 incubator and culture overnight.
- The next day, replace the medium with fresh TeSR-E8 medium without Rock inhibitor Y-27632.
- After that, feed cells daily with 2 ml TeSR-E8 medium per well in the 6-well plate, the hiPSCs/hESCs are allowed to grow for 4-5 days. When cells reach 70%-80% confluence, they can be passaged (see Note 5).
- Passage hiPSCs and hESCs using ReLeSR (6-well plate format)
Generally, hiPSCs and hESCs can grow as colonies when plated on Vitronectin XF, with typical undifferentiated, dense and compact morphology and a high nucleus-to-cytoplasm ratio (Figure 1B, Day -2). Ideally, hiPSCs and hESCs can be passaged every 4-5 days (70%-80% confluence) as either single cell or cell aggregates. We routinely use ReLeSR for maintenance of hiPSCs and hESCs as cell aggregates (see Note 6).
The following steps are for passaging hiPSCs/hESCs from wells of the 6-well plate in TeSR-E8 medium and Vitronectin XF systems. If using other culture vessels, adjust volumes of reagents according to surface area (see Table 1).
Table 1. Recommended volume of related reagents for different culture vessels
- Prior to starting, coat new 6-well plates with Vitronectin XF at least 1 h and pre-warm the required volume of TeSR-E8 medium to room temperature.
- Remove the spent medium from the wells and wash cells once with 2 ml DPBS w/o Ca2+ and Mg2+. No need to remove differentiated regions of hiPSCs/hESCs.
- Remove DPBS w/o Ca2+ and Mg2+, and then add 1 ml ReLeSR into each well of 6-well plate. Swirl the plate to distribute ReLeSR over the entire cell surface, and then aspirate ReLeSR within 1 min.
- Incubate hiPSCs/hESCs for 5 min at room temperature (see Note 7), and check cell dissociation under the inverted phase-contrast microscope.
- When hiPSCs/hESCs start to separate and round up at the edge, they are ready to be removed from the wells, and then add 1 ml TeSR-E8 medium to each well of the 6-well plate.
- Disperse the big colonies into small clumps by gently pipetting the mixtures up and down 5-6 times. Collect all cell aggregates into a 15 ml tube (see Note 8).
- Rinse the well with another 1 ml TeSR-E8 medium and transfer into the same 15 ml tube.
- Remove the residual Vitronectin XF solution from the pre-coated plate. And evenly split the cell aggregates at the desired density into the plate. We usually split the hiPSCs/hESCs at the ratio of 1:6 (see Note 9).
- Supplement an appropriate volume of TeSR-E8 medium to each well, so that each well can contain 2 ml TeSR-E8 medium finally.
- Add 20 μl of Rock inhibitor Y-27632 (100x, working concentration 10 μM) into the culture system to increase survival of hiPSCs/hESCs.
- Gently move the plate in several times quick, back and forth, side to side motions to uniformly distribute the cell aggregates across the surface of the plate, and then return the plate into the 37 °C incubator (see Note 10).
- Refresh the TeSR-E8 medium daily and visualize cultures to monitor growth condition until the next passaging or banking (see Note 11).
- Derive MSCs from hiPSCs and hESCs with a single step medium switch
We use a simple and efficient protocol to differentiate hiPSCs and hESCs into MSCs in feeder-free culture system. The timetable diagram of this differentiation procedure is shown in Figure 1A.- Day -2: hiPSCs and hESCs preparation in 6-well plate
Before hiPSCs/hESCs differentiation into MSCs, hiPSCs/hESCs were plated as clumps into Vitronectin XF coated 6-well plates by ReLeSR treatment in TeSR-E8 medium (Figure 1B, Day -2).- Pre-warm the TeSR-E8 medium. Meanwhile, prepare Vitronectin XF-coated 6-well plate as before.
- Seed cell clumps of hiPSCs/hESCs into Vitronectin XF-coated 6-well plate at the desired density as Steps B2-B11 in the Procedure B.
- Culture the cells overnight at 37 °C to maximally make cells to attach wells.
- Day -1: Fresh TeSR-E8 medium renewal
- Check cell growth condition under microscope to ensure that the majority of passaged hiPSC/hESC aggregate attach onto new wells and start to proliferate.
- Remove spent medium and feed cells with 2 ml fresh TeSR-E8 medium in 6-well plates, and continue to incubate cells at 37 °C, 5% CO2.
- Monitor the cell growth until hiPSCs/hESCs become at least 40%-50% confluent for MSC differentiation.
- Day 0: MSC low-glucose medium switch
The hiPSCs/hESCs should normally reach at least 40%-50% confluence before initiating differentiation (see Note 12). The hiPSC/hESC colonies display typical compact morphology with clear border, composed of tightly packed cells (Figure 1B, Day 0).- Pre-warm MSC low glucose medium in a 37 °C water bath.
- When the passaged hiPSCs/hESCs reach the desired confluence in the 6-well plate, gently remove the spent TeSR-E8 medium, and then wash the hiPSC/hESC colonies twice each with 2 ml MSC low glucose medium.
- Aspirate the spent MSC low glucose medium thoroughly, add 2 ml fresh MSC low glucose medium into hiPSCs/hESCs for each well of the 6-well plate.
- Thereafter, refresh MSC low glucose medium every other day.
- Days 2-13: Fresh MSC low glucose medium renewal
In first 2 days after MSC low glucose medium treatment, many cells die due to the culture pressure from differentiation in MSC low glucose medium. Feed cells with fresh MSC low glucose medium every other day.- Gently aspirate the spent MSC low glucose medium and dead cells.
- Wash the differentiated cells twice with DPBS w/o Ca2+ and Mg2+, add 2 ml fresh MSC low glucose medium into culture wells and continue to culture cells at 37 °C, 5% CO2 (see Note 13).
- Observe cells under microscope to check cell morphology changes and record the photos (see Note 14) (Figure 1B, Day 7).
- Day 14: Plating heterogeneous cell derivations onto 0.1% gelatin-coated T25 flask
At this time point, the cell derivations are confluent, however with heterogeneous cell populations (Figure 1B, Day 14). The cell derivations can be passaged onto 0.1% gelatin-coated flasks.- Pre-warm 0.05% trypsin-EDTA and MSC low glucose medium to room temperature, and pre-coat the T25 flask with 0.1% gelatin in a 37 °C incubator.
- Aspirate the spent MSC low glucose medium from the 6-well plates.
- Wash the cells once with 1 ml DPBS w/o Ca2+ and Mg2+, and then discard.
- Add 1 ml 0.05% trypsin-EDTA to wells of the 6-well plate. Swirl plate to ensure that the trypsin solution completely covers the entire cell surface.
- Incubate the cell derivations at 37 °C for 5 min (see Note 15). Gently tap plate to dislodge cells if necessary.
- Add 2 ml MSC low glucose medium into the well for neutralizing trypsinization. And then pipette cells up and down several times (normally 8-10 times) to disassociate cell derivations into single cells.
- Collect the cell suspension into a 15 ml Falcon tube, wash the culture wells with an additional 2 ml MSC low glucose medium, and pool all the cell suspension into the same 15 ml Falcon tube.
- Centrifuge the cell suspension at 300 x g for 5 min, and then discard supernatant.
- Break the cell pellets by finger tapping, and then resuspend the cell pellets gently with 3 ml MSC low glucose medium to make a single cell suspension.
- Plate all the cell suspension into the 0.1% gelatin-coated T25 flask previously prepared.
- Supplement additional medium into the T25 flask, ensure the T25 flask can contain at least 5 ml medium.
- Culture the cells in the 37 °C, 5% CO2 incubator and refresh MSC low glucose medium every other day until the cells become 80% confluent.
- Days 15-20: Fresh MSC low glucose medium renewal
At the first 2 days after plating cell derivations into the 0.1% gelatin-coated flask, not all cells can attach flasks. The attached cells are the expected cells we require. In 5-7 days, the attached cells can become confluent in the T25 flask. The MSC low glucose medium is renewed every other day. - Days 21-29: Enrich hiPSC-MSCs and hESC-MSCs by serial trypsinization (8 days)
When the differentiated cells in the T25 flask reach 80%-90% confluence, they can be split into new T25 flasks. The most of cells present fibroblast-like morphology (spindle-shape) (Figure 1B, Day 21).- Prepare 0.1% gelatin-coated T25 flasks, warm MSC low glucose medium and 0.05% trypsin-EDTA as before.
- Harvest cell pellets as in the Steps C5b-C5h.
- Finger tapping to break the cell pellets, and resuspend the cells in 4 ml MSC low glucose medium to make a single cell suspension.
- Count the total cell number using a hemocytometer and adjust the density of cells to 2.5 x 105 cells/ml using the MSC low glucose medium.
- Seed the cells into new 0.1% gelatin-coated T25 flasks at a cell density of 1 x 104 cell/cm2 (i.e., 2.5 x 105 cells in T25 flask) (see Note 16). Meanwhile, we designate this generation of cells as passage 0 (P0) at this point, and name the cells as hiPSC-MSCs/hESC-MSCs.
- Supplement additional MSC low glucose medium into the T25 flasks, ensure the T25 flasks contain 5 ml MSC low glucose medium.
- Thereafter, replace the MSC low glucose medium every other day.
- Consecutively passage the hiPSC-MSCs/hESC-MSCs by trypsinization when they are 80%-90% confluent (normally within 4-5 days). The derived MSCs can gradually become mature population with 2 more times passages. Designate the corresponding generations as P1, P2. (Figure 1B, MSC, P2). Large expand these hiPSC-MSCs/hESC-MSCs for banking, characterizations and later experiments.
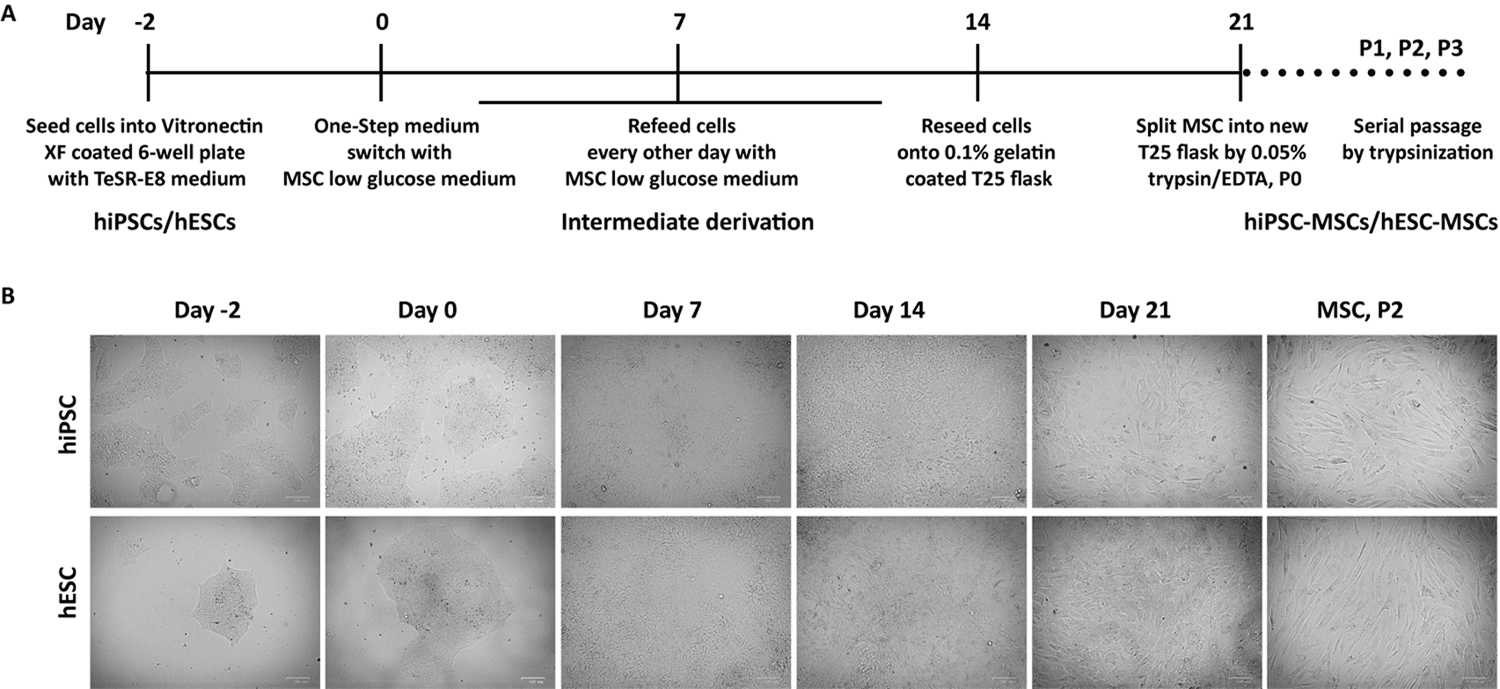
Figure 1. Differentiation of hiPSCs and hESCs into MSCs by one-step derivation in feeder-free system. A. Schematic diagram of efficient differentiating hiPSCs and hESCs into MSCs by MSC low glucose medium treatment and serial passages with trypsinization in feeder-free culture system. B. Representative morphology of cells at key phages during MSC derivation from hiPSCs and hESCs at Days -2, 0, 7, 14 and 21, as well as P2 of mature hiPSC-MSCs and hESC-MSCs. P2, Passage 2. Scale bars: 100 μm. - Day -2: hiPSCs and hESCs preparation in 6-well plate
- Cryopreserving hiPSC-MSCs and hESC-MSCs
After serial trypsinization, the hiPSC-MSCs/hESC-MSCs can become more mature. Therefore they can be cryopreserved after passage 2.- Prepare fresh 2x MSC freezing medium, and put it on ice until use.
- Make MSC pellets by following the Steps C5b-C5h.
- Dislodge the cell pellets by finger tapping, and resuspend the cells in MSC low glucose medium to make a single cell suspension.
- Count the total cell number using a hemocytometer and adjust the cell density to 2 x 106 cell/ml.
- Add the same volume of cold 2x MSC Freezing Medium into the MSC cell suspension, and mix them thoroughly.
- Aliquot 1 ml cell mixture into each cryogenic vial. Each cryogenic vial contains 1 x 106 cells (see Note 17).
- Quickly put cryogenic vials into the freezing container, and keep it at -80 °C overnight.
- The next day, transfer cryogenic vials with cells into -180 °C freezer or liquid nitrogen tank for long-term storage.
- MSC Surface antigens analysis by flow cytometry
The hiPSC-MSCs/hESC-MSCs express similar surface antigens as hBM-MSCs. Antibodies against human antigens CD34, CD45, CD73, CD90, and CD105 can be used in this assay (see Antibodies). In this assay, hBM-MSCs can be used as positive control cells. Negative markers: CD34–, CD45–; Positive markers: CD73+, CD90+, and CD105+.
The results of MSC surface antigens analysis in hiPSC-MSCs, hESC-MSCs, and hBM-MSCs were shown in Figure 2.- The MSC pellets are collected as the Steps C5b-C5h.
- Break the cell pellets by finger tapping, and resuspend cells with cold 1% FBS in DPBS w/o Ca2+ and Mg2+.
- Count the total cell number using a hemocytometer, and adjust the cell density to 2 x 106 cells/ml.
- Transfer 2 x 105 cells (100 μl cell suspension) into 1.5 ml centrifuge tubes (see Note 18).
- Add corresponding amount MSC antibodies (see Antibodies) to each tube, and gently mix them by pipetting.
- Incubate for 30 min at room temperature in the dark.
- Wash cell once with 1% FBS in DPBS w/o Ca2+ and Mg2+ and finally resuspend cell samples in 250 µl 4% formaldehyde-DPBS w/o Ca2+ and Mg2+.
- Analyze samples with a flow cytometer (LSR Fortessa) (see Note 19).
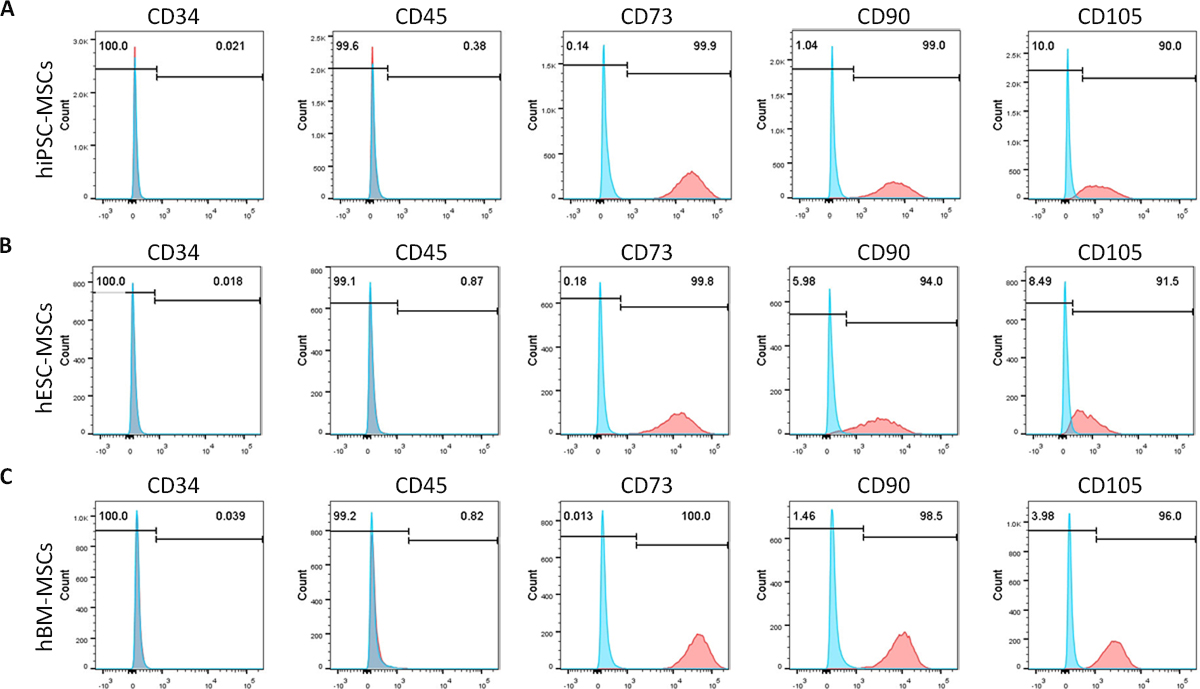
Figure 2. Flow cytometric analysis of MSC surface antigens in hiPSC-ESCs, hESC-MSCs and hBM-MSCs. Expression percentages of MSC surface antigens were analyzed in hiPSC-MSCs (A), hESC-MSCs (B) and hBM-MSCs (C). Unstained cells were used as the control. Percentage of cells positive for each marker is calculated by normalizing to the unstained cells. Negative markers: CD34–, CD45–; Positive markers: CD73+, CD90+, and CD105+. - Characterization of hiPSC-MSCs and hESC-MSCs by in vitro multi-potent differentiation
The hiPSC-MSCs and hESC-MSCs possess multipotent differentiation potential (Osteogenesis, Chondrogenesis, and Adipogenesis). We recommend typically performing the in vitro differentiation assay using the early passage of cells (passage 5) to characterize the multipotency of hiPSC-MSCs and hESC-MSCs (see Note 20) (Figure 3).- Osteogenic differentiation of hiPSC-MSCs and hESC-MSCs
Note: Osteogenic differentiation can be performed using the StemPro Osteogenesis Differentiation Kit.- When the hiPSC-MSCs and hESC-MSCs reach 80% confluence in a T75 flask, wash cells with DPBS w/o Ca2+ and Mg2+.
- Harvest cells with 0.05% Trypsin-EDTA.
- Count the cells and seed the cell suspension into 0.1% gelatin-coated 24-well plates at a cell density of 1 x 104 cells/cm2 with 0.5 ml MSC low glucose medium (i.e., 2 x 104 cells/well in 24-well plate).
- Incubate the cells in a 37 °C, 5% CO2 incubator for 3 days in MSC low glucose medium.
- Wash cells once with DPBS w/o Ca2+ and Mg2+, and then add 0.5 ml Osteogenesis differentiation medium.
- Feed cultures every 3 days up to 3 weeks with the osteogenic differentiation medium.
- After 21 days differentiation cultivation, fix the cells with 4% formaldehyde for 30 min.
- Stain cells with 2% Alizarin Red S for calcium deposit identification (Figure 3A), and then acquires images using standard light microscopy.
- Chondrogenic differentiation of hiPSC-MSCs and hESC-MSCs
Note: Chondrogenic differentiation can be performed using the StemPro Chondrogenesis Differentiation Kit.- At the day of differentiation, the hiPSC-MSCs and hESC-MSCs should be 80% confluent.
- Aspirate MSC low glucose medium, rinse the cells with DPBS w/o Ca2+ and Mg2+, and collect the cells with 0.05% Trypsin-EDTA.
- Count the cells and adjust hiPSC-MSCs/hESC-MSCs to a cell density of 1.6 x 107 cell/ml with the MSC low glucose medium.
- Seed only 5 μl droplets of hiPSC-MSCs/hESC-MSCs solution (8 x 104 cell/well) in the center of 96-multiwell plates.
- Place the plates back into the 37 °C, 5% CO2 incubator for 2 h micromass cultivation.
- After micromass cultivating, supplement 100 μl the chondrogenesis medium to culture wells.
- Incubate the cells in the 37 °C, 5% CO2 incubator for 3 weeks and feed cultures every 3 days with the chondrogenic differentiation medium.
- Paraffin embed chondrogenic pellets, section, and dye cells by toluidine blue staining to evaluate extracellular chondrocyte matrix (Figure 3B), and then acquire images using standard light microscopy.
- Adipogenic differentiation of hiPSC-MSCs and hESC-MSCs
Note: Adipogenic differentiation can be performed using the StemPro Adipogenesis Differentiation Kit.- Observe cells under a microscope to ensure that the hiPSC-MSCs and hESC-MSCs are at 80% confluence before differentiation.
- Dissociate the cells into a single cell suspension by treatment with 0.05% Trypsin-EDTA.
- Count the cells and seed the cell suspension into 0.1% gelatin-coated 24-well plates at a density of 1 x 104 cells/cm2 with 0.5 ml pre-warmed MSC low glucose medium (i.e., 2 x 104 cells/well in 24-well plate).
- Incubate the cells in MSC low glucose medium in a 37 °C, 5% CO2 incubator for 3 days.
- Wash cells once with DPBS w/o Ca2+ and Mg2+, and then add 500 μl adipogenic differentiation medium.
- Renew medium every 3 days, and continuously culture cells with adipogenic differentiation medium for 2 weeks.
- After 14 days induction cultivation, fix the cells with 4% formaldehyde for 30 min.
- Apply Oil Red O staining to validate the lipid droplets (Figure 3C), and then acquire images using standard light microscopy.
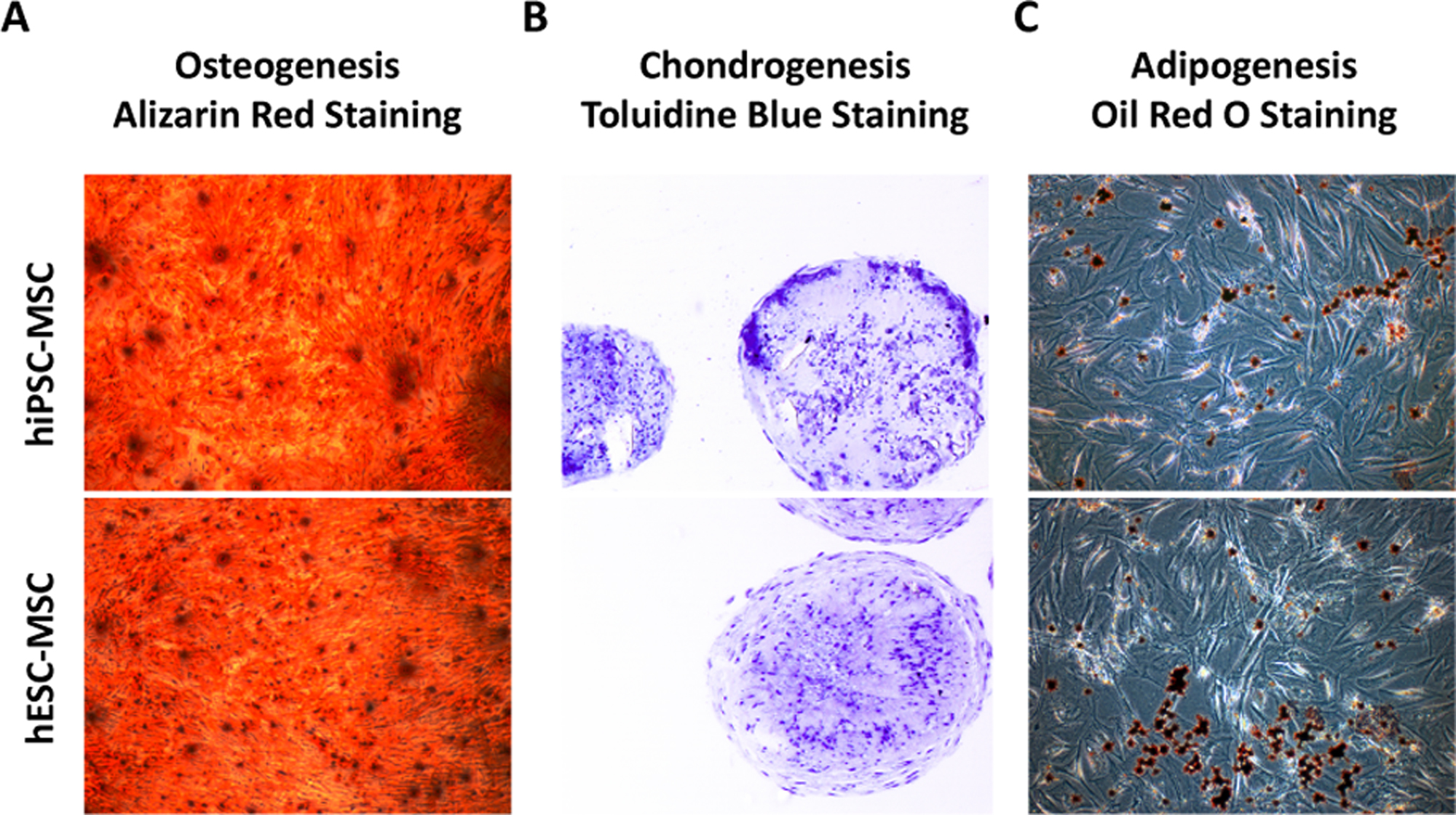
Figure 3. Tri-lineage differentiation of hiPSC-MSCs and hESC-MSCs. A. Alizarin Red staining of calcium deposition of hiPSC-MSCs and hESC-MSCs in osteogenic differentiation medium for 3 weeks. Magnification: 50x. B. Toluidine blue staining of cartilaginous extracellular matrix of hiPSC-MSCs and hESC-MSCs after 3 weeks in chondrogenic differentiation medium. Magnification: 100x. C. Oil Red O staining of lipid droplets of hiPSC-MSCs and hESC-MSCs in adipogenic differentiation medium for 2 weeks. Magnification: 50x. - Osteogenic differentiation of hiPSC-MSCs and hESC-MSCs
Data analysis
- At least 20,000 events are recorded on BD Fortessa flow cytometer when analyzing MSC surface antigens characterization using antibodies against CD34, CD45, CD73, CD90 and CD105. Flow Jo software is used to analyze the flow cytometry data.
- The staining photos of osteocytes, chondrocytes and adipocytes can be analyzed using ImageJ software.
Notes
- Do not warm the TeSR-E8 medium in a 37 °C water bath.
- Transfer hiPSCs/hESCs into the Falcon tube with drop-wise manner, meanwhile gently move the Falcon tube back and forth for mixing the hPSCs, which can reduce osmotic shock to the hPSCs and ensure cell viability.
- The addition of Rock inhibitor Y-27632 (100x stock solution) into the hPSC medium 24 h greatly improves the plating efficiency and the viability of hPSCs after thawing without affecting their pluripotency.
- Gently move culture plates, do not swirl them, as it will bring all cells to the center of the plates.
- The hiPSC/hESC colonies should become visible within 2 days after thawing, and in 4-5 days the hiPSCs/hESCs should be ready for passaging.
- Enzymes such as collagenase IV or dispase does not work well with hPSCs cultured in TeSR-E8 medium system. ReLeSR is an enzyme-free reagent suitable for dissociating hiPSCs and hESCs into cell clumps for routine passaging without manual removal of differentiated regions.
- The treatment time of ReLeSR may vary depending on the hPSC lines and quality of colonies. The optimal dissociation time with ReLeSR for some certain hPSC lines may take longer than 5 min.
- Gently pipetting the cell suspension several times can break up hPSC colonies into small cell clumps after ReLeSR treatment. Meanwhile, avoid creating single cell suspension. Sometimes there are some cells which do not detach after ReLeSR treatment. Collecting them is unnecessary.
- The split ratio can vary for different hPSC lines. The established hPSC cultures can be split with the ratio of 1:6 to 1:12 (i.e., cell clumps from one well can be plated into six to twelve wells).
- Uneven distribution of cell slumps may increase the differentiation of hiPSC/hESCs. Avoid disturbing the plate for overnight to attach cells maximally.
- It is normal to see some cell debris the day after passaging. When hiPSCs/hESCs reach 80% confluence, they can be passaged or frozen down with TeSR-E8 Freezing Medium. Do not allow hiPSC/hESCs to be over-confluent to avoid spontaneous differentiation.
- The hiPSCs/hESCs should be at least 40% confluent 2 days after seeding. If the cells are not enough to initialize differentiation, the cells can be incubated for 1 or 2 more days before MSC differentiation. However, the hiPSCs/hESCs should not be over confluent before MSC differentiation. Otherwise, they will go through random differentiation.
- In the first several days of hiPSC/hESC differentiation into MSCs, the differentiated cells should be washed with DPBS w/o Ca2+ and Mg2+ when renewing medium due to many dead cells. Afterward, if the spent medium is clean and there are no many dead cells, the step of washing cells is unnecessary.
- The cell morphology changes can be gradually visualized as early as 7 days post-differentiation from tight and flat colonies into fibroblast-like cells. Most of cells derived from hiPSCs and hESCs in this differentiation procedure would present typically spindle-shaped morphologies at day 14.
- It is important to observe cell monolayer under a microscope. When most of cells detach at edge from the surface of the culture vessels, they are ready to be removed. The incubation time for some cell lines may be less than 5 min.
- The cell density of 1 x 104 cell/cm2 is a good density for the cell growth of MSCs derived from hiPSCs/hESCs. At this time point, we can also directly split cells into the new T25 flasks at 1:3-1:4 ratio.
- Generally, the hiPSC-MSCs/hESC-MSCs in the T75 flask can be frozen into 4-6 cryogenic vials, when they are 80% confluent.
- Remember to prepare groups for unstained control.
- We recommend analyzing samples on the same day for best results. If samples cannot be analyzed at the same day, the samples can be kept in the dark at 4 °C for several days until analysis.
- Typically, hiPSC-MSCs and hESC-MSCs will gradually lose their multipotency at late passage number (after passage 15).
Recipes
- Complete TeSR-E8 Medium for hiPSCs and hESCs
- Thaw TeSR-E8 Supplement (25x) at room temperature (15-25 °C) or 4 °C overnight. Do not thaw at 37 °C. Mix thoroughly. Once thawed, use the supplement immediately, do not refreeze
- To prepare 500 ml Complete TeSR-E8 Medium, aseptically mix thoroughly 480 ml TeSR-E8 Basal Medium and 20 ml TeSR-E8 Supplement (25x)
- Complete TeSR-E8 Medium can be stored at 4 °C for up to 2 weeks. Alternatively, aliquot 50 ml Complete TeSR-E8 Medium and store at -20 °C for up to 6 months
- Before use, warm complete TeSR-E8 medium at room temperature. Do not warm the medium in a 37 °C water bath
- Vitronectin XF coated culture vessels (6-well plate format as an example)
- Thaw the vial of Vitronectin XF at room temperature. If not used immediately, aliquot Vitronectin XF and immediately freeze at -80 °C. Avoid repeated freeze-thaw cycles. The Vitronectin XF can be stored at -80 °C for up to several months
- To prepare the working solution, dilute Vitronectin XF in a ratio of 1:25 in Cell Adhere Dilution Buffer to reach a final working concentration of 10 μg/ml (i.e., use 40 μl Vitronectin XF per 1 ml Cell Adhere Dilution Buffer)
- Add 240 μl thawed Vitronectin XF into a 15 ml Falcon tube containing 6 ml Cell Adhere Dilution Buffer. Gently mix the solution up and down by pipetting. Do not vortex
- Immediately use 1 ml diluted Vitronectin XF solution to coat each well of a 6-well plate (refer to Table 1 for recommended coating volumes for different culture vessels)
- Gently swirl the culture plate to spread the Vitronectin XF solution so as to evenly cover the entire surface of culture plate
- Incubate the Vitronectin XF coated plate at room temperature for at least 1 h. If not used immediately, the culture plate can be sealed with parafilm and stored at 4 °C for a week. Prior to use, warm the stored culture plate to room temperature for at least 30 min
- Remove the excess Vitronectin XF solution from the culture plate before plating cells.
- Immediately seed hiPSCs or hESCs onto the Vitronectin XF coated culture plate with an appropriate volume of Complete TeSR-E8 Medium (2 ml medium for one 6-well plate, Refer to Table 1)
- Rock inhibitor (100x Stock solution)
ROCK inhibitor Y-27632 is a cell-permeable, highly selective inhibitor of ROCK pathway. And Y-27632 is light sensitive. Therefore, it should be protected from light.- Dissolve 1 mg Y-27632 into 3.12 ml cooled distilled water to prepare 100x stock solution (1 mM)
- Aliquot the stock solution into 100 μl aliquots and store at -20 °C for up to 6 months. Y-27632 is effective at a final working concentration of 10 μM
- TeSR-E8 freezing medium
- To make 10 ml fresh TeSR-E8 freezing medium, aseptically mix 9 ml Complete TeSR-E8 Medium and 1 ml DMSO in a 15 ml Falcon tube (Medium: DMSO = 9:1).
- Store the TeSR-E8 freezing medium on ice until use. Discard any remaining freezing medium after use.
- MSC low glucose medium
MSC low glucose medium contains DMEM-low glucose (1 g/L) supplemented with 10% HQ-FBS, 1% Penicillin-Streptomycin, 1% NEAA and 1% GlutaMAX.- To prepare 500 ml MSC low glucose medium, mix 50 ml HQ-FBS, 5 ml Penicillin-Streptomycin, 5 ml NEAA and 5 ml GlutaMAX, and then fill up to 500 ml with DMEM-low glucose medium
- Store the MSC low glucose medium at 4 °C for up to 2 months
- 1% (wt/vol) gelatin stock solution
The gelatin stock solution contains 1% (wt/vol) gelatin in distilled H2O.- Weigh 10 g gelatin powder and add it into 1,000 ml distilled H2O
- Heat and stir the gelatin solution on a hot plate until the gelatin powder completely dissolves
- Autoclave the gelatin stock solution at 115 °C for 15 min to make homogenous solution (1% stock solution)
- Dispense the gelatin stock solution into a 500 ml bottle, and store the 1% gelatin solution at room temperature up to several months
- 0.1% gelatin-coated vessels (T25 flask format as an example)
- Dilute the 1% gelatin stock solution into 0.1% gelatin working solution with distilled water (10x dilution)
- Add at least 5 ml 0.1% gelatin working solution to cover the entire bottom of a T25 flask
- Incubate the T25 flask for 1 h at 37 °C. The coated vessels can be stored at 4 °C for several weeks, if not used immediately
- Remove excess gelatin solution, and then seed cell suspension directly onto the 0.1% gelatin-coated T25 flask
- 2x MSC freezing medium
We use 2x MSC Freezing medium to cryopreserve MSCs. The 2x MSC freezing medium contains 80% HQ-FBS and 20% DMSO.- To make 10 ml 2x MSC freezing medium, aseptically mix 8 ml HQ-FBS and 2 ml DMSO in a 15 ml Falcon tube
- Chill the Falcon tube with 2x MSC Freezing Medium on ice before use
- Mix the 2x MSC Freezing medium with cell suspension at 1:1 ratio when using it. Always make fresh freezing medium and discard any remaining freezing medium after use
- The final medium approximately contains 50% MSC low glucose medium, 40% HQ-FBS, and 10% DMSO
- 4% formaldehyde solution
This formaldehyde solution contains 37% formaldehyde and DPBS at 4:34 ratio. Formaldehyde is a hazardous reagent, so wear gloves and lab coat while handing it.- To make 37 ml 4% formaldehyde solution, add 4 ml 37% formaldehyde into 33 ml DPBS w/o Ca2+ and Mg2+
- Thoroughly mix the solution and keep at 4 °C before use. Always prepare fresh 4% formaldehyde solution before use
Acknowledgments
This protocol is developed and modified from our previous studies published in Scientific Reports (Zou et al., 2013) and Stem Cell Research and Therapy (Kang et al., 2015). This work was supported by Shenzhen Science and Technology Innovation Committee (CKCY2016082917391652, CKCY2016082917372416, and JCYJ20160226192924528). We also acknowledge FACS core facility in Aarhus University.
Competing interests
The authors declare that no competing interests exist.
References
- Brown, P. T., Squire, M. W. and Li, W. J. (2014). Characterization and evaluation of mesenchymal stem cells derived from human embryonic stem cells and bone marrow. Cell Tissue Res 358(1): 149-164.
- Campagnoli, C., Roberts, I. A., Kumar, S., Bennett, P. R., Bellantuono, I. and Fisk, N. M. (2001). Identification of mesenchymal stem/progenitor cells in human first-trimester fetal blood, liver, and bone marrow. Blood 98(8): 2396-2402.
- Chen, Y. S., Pelekanos, R. A., Ellis, R. L., Horne, R., Wolvetang, E. J. and Fisk, N. M. (2012). Small molecule mesengenic induction of human induced pluripotent stem cells to generate mesenchymal stem/stromal cells. Stem Cells Transl Med 1(2): 83-95.
- Chen, Y., Shao, J. Z., Xiang, L. X., Dong, X. J. and Zhang, G. R. (2008). Mesenchymal stem cells: a promising candidate in regenerative medicine. Int J Biochem Cell Biol 40(5): 815-820.
- Crisan, M., Yap, S., Casteilla, L., Chen, C. W., Corselli, M., Park, T. S., Andriolo, G., Sun, B., Zheng, B., Zhang, L., Norotte, C., Teng, P. N., Traas, J., Schugar, R., Deasy, B. M., Badylak, S., Buhring, H. J., Giacobino, J. P., Lazzari, L., Huard, J. and Peault, B. (2008). A perivascular origin for mesenchymal stem cells in multiple human organs. Cell Stem Cell 3(3): 301-313.
- da Silva Meirelles, L., Chagastelles, P. C. and Nardi, N. B. (2006). Mesenchymal stem cells reside in virtually all post-natal organs and tissues. J Cell Sci 119(Pt 11): 2204-2213.
- Gao, W. X., Sun, Y. Q., Shi, J., Li, C. L., Fang, S. B., Wang, D., Deng, X. Q., Wen, W. and Fu, Q. L. (2017). Effects of mesenchymal stem cells from human induced pluripotent stem cells on differentiation, maturation, and function of dendritic cells. Stem Cell Res Ther 8(1): 48.
- Ghasroldasht, M. M., Irfan-Maqsood, M., Matin, M. M., Bidkhori, H. R., Naderi-Meshkin, H., Moradi, A. and Bahrami, A. R. (2014). Mesenchymal stem cell based therapy for osteo-diseases. Cell Biol Int 38(10): 1081-1085.
- Horwitz, E. M., Prockop, D. J., Fitzpatrick, L. A., Koo, W. W., Gordon, P. L., Neel, M., Sussman, M., Orchard, P., Marx, J. C., Pyeritz, R. E. and Brenner, M. K. (1999). Transplantability and therapeutic effects of bone marrow-derived mesenchymal cells in children with osteogenesis imperfecta. Nat Med 5(3): 309-313.
- Hynes, K., Menicanin, D., Mrozik, K., Gronthos, S. and Bartold, P. M. (2014). Generation of functional mesenchymal stem cells from different induced pluripotent stem cell lines. Stem Cells Dev 23(10): 1084-1096.
- Kang, R., Luo, Y., Zou, L., Xie, L., Lysdahl, H., Jiang, X., Chen, C., Bolund, L., Chen, M., Besenbacher, F. and Bünger, C. (2014). Osteogenesis of human induced pluripotent stem cells derived mesenchymal stem cells on hydroxyapatite contained nanofibers. RSC Adv 4(11): 5734-5739.
- Kang, R., Zhou, Y., Tan, S., Zhou, G., Aagaard, L., Xie, L., Bunger, C., Bolund, L. and Luo, Y. (2015). Mesenchymal stem cells derived from human induced pluripotent stem cells retain adequate osteogenicity and chondrogenicity but less adipogenicity. Stem Cell Res Ther 6: 144.
- Karlsson, C., Emanuelsson, K., Wessberg, F., Kajic, K., Axell, M. Z., Eriksson, P. S., Lindahl, A., Hyllner, J. and Strehl, R. (2009). Human embryonic stem cell-derived mesenchymal progenitors--potential in regenerative medicine. Stem Cell Res 3(1): 39-50.
- Kretlow, J. D., Jin, Y. Q., Liu, W., Zhang, W. J., Hong, T. H., Zhou, G., Baggett, L. S., Mikos, A. G. and Cao, Y. (2008). Donor age and cell passage affects differentiation potential of murine bone marrow-derived stem cells. BMC Cell Biol 9: 60.
- Lalu, M. M., McIntyre, L., Pugliese, C., Fergusson, D., Winston, B. W., Marshall, J. C., Granton, J., Stewart, D. J. and Canadian Critical Care Trials, G. (2012). Safety of cell therapy with mesenchymal stromal cells (SafeCell): a systematic review and meta-analysis of clinical trials. PLoS One 7(10): e47559.
- Lee, O. K., Kuo, T. K., Chen, W. M., Lee, K. D., Hsieh, S. L. and Chen, T. H. (2004). Isolation of multipotent mesenchymal stem cells from umbilical cord blood. Blood 103(5): 1669-1675.
- Lian, Q., Lye, E., Suan Yeo, K., Khia Way Tan, E., Salto-Tellez, M., Liu, T. M., Palanisamy, N., El Oakley, R. M., Lee, E. H., Lim, B. and Lim, S. K. (2007). Derivation of clinically compliant MSCs from CD105+, CD24- differentiated human ESCs. Stem Cells 25(2): 425-436.
- Lian, Q., Zhang, Y., Zhang, J., Zhang, H. K., Wu, X., Zhang, Y., Lam, F. F., Kang, S., Xia, J. C., Lai, W. H., Au, K. W., Chow, Y. Y., Siu, C. W., Lee, C. N. and Tse, H. F. (2010). Functional mesenchymal stem cells derived from human induced pluripotent stem cells attenuate limb ischemia in mice. Circulation 121(9): 1113-1123.
- Mezey, E., Chandross, K. J., Harta, G., Maki, R. A. and McKercher, S. R. (2000). Turning blood into brain: cells bearing neuronal antigens generated in vivo from bone marrow. Science 290(5497): 1779-1782.
- Noth, U., Osyczka, A. M., Tuli, R., Hickok, N. J., Danielson, K. G. and Tuan, R. S. (2002). Multilineage mesenchymal differentiation potential of human trabecular bone-derived cells. J Orthop Res 20(5): 1060-1069.
- Peng, K. Y., Lee, Y. W., Hsu, P. J., Wang, H. H., Wang, Y., Liou, J. Y., Hsu, S. H., Wu, K. K. and Yen, B. L. (2016). Human pluripotent stem cell (PSC)-derived mesenchymal stem cells (MSCs) show potent neurogenic capacity which is enhanced with cytoskeletal rearrangement. Oncotarget 7(28): 43949-43959.
- Schuh, E. M., Friedman, M. S., Carrade, D. D., Li, J., Heeke, D., Oyserman, S. M., Galuppo, L. D., Lara, D. J., Walker, N. J., Ferraro, G. L., Owens, S. D. and Borjesson, D. L. (2009). Identification of variables that optimize isolation and culture of multipotent mesenchymal stem cells from equine umbilical-cord blood. Am J Vet Res 70(12): 1526-1535.
- Trounson, A. and McDonald, C. (2015). Stem cell therapies in clinical trials: progress and challenges. Cell Stem Cell 17(1): 11-22.
- Tuan, R. S., Boland, G. and Tuli, R. (2003). Adult mesenchymal stem cells and cell-based tissue engineering. Arthritis Res Ther 5(1): 32-45.
- Uccelli, A., Moretta, L. and Pistoia, V. (2008). Mesenchymal stem cells in health and disease. Nat Rev Immunol 8(9): 726-736.
- Wakitani, S., Nawata, M., Tensho, K., Okabe, T., Machida, H. and Ohgushi, H. (2007). Repair of articular cartilage defects in the patello-femoral joint with autologous bone marrow mesenchymal cell transplantation: three case reports involving nine defects in five knees. J Tissue Eng Regen Med 1(1): 74-79.
- Wei, H., Tan, G., Manasi, Qiu, S., Kong, G., Yong, P., Koh, C., Ooi, T. H., Lim, S. Y., Wong, P., Gan, S. U. and Shim, W. (2012). One-step derivation of cardiomyocytes and mesenchymal stem cells from human pluripotent stem cells. Stem Cell Res 9(2): 87-100.
- Wu, L., Cai, X., Zhang, S., Karperien, M. and Lin, Y. (2013). Regeneration of articular cartilage by adipose tissue derived mesenchymal stem cells: perspectives from stem cell biology and molecular medicine. J Cell Physiol 228(5): 938-944.
- Zhao, Q., Gregory, C. A., Lee, R. H., Reger, R. L., Qin, L., Hai, B., Park, M. S., Yoon, N., Clough, B., McNeill, E., Prockop, D. J. and Liu, F. (2015). MSCs derived from iPSCs with a modified protocol are tumor-tropic but have much less potential to promote tumors than bone marrow MSCs. Proc Natl Acad Sci U S A 112(2): 530-535.
- Zou, L., Luo, Y., Chen, M., Wang, G., Ding, M., Petersen, C. C., Kang, R., Dagnaes-Hansen, F., Zeng, Y., Lv, N., Ma, Q., Le, D. Q., Besenbacher, F., Bolund, L., Jensen, T. G., Kjems, J., Pu, W. T. and Bunger, C. (2013). A simple method for deriving functional MSCs and applied for osteogenesis in 3D scaffolds. Sci Rep 3: 2243.
- Zuk, P. A., Zhu, M., Mizuno, H., Huang, J., Futrell, J. W., Katz, A. J., Benhaim, P., Lorenz, H. P. and Hedrick, M. H. (2001). Multilineage cells from human adipose tissue: implications for cell-based therapies. Tissue Eng 7(2): 211-228.
Article Information
Copyright
© 2018 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Zhou, Y., Liao, J., Fang, C., Mo, C., Zhou, G. and Luo, Y. (2018). One-step Derivation of Functional Mesenchymal Stem Cells from Human Pluripotent Stem Cells. Bio-protocol 8(22): e3080. DOI: 10.21769/BioProtoc.3080.
Category
Stem Cell > Adult stem cell > Mesenchymal stem cell
Stem Cell > Pluripotent stem cell > Cell differentiation
Cell Biology > Cell isolation and culture > Cell differentiation
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link













