- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Detection of Cell Death in Planarians
Published: Vol 8, Iss 19, Oct 5, 2018 DOI: 10.21769/BioProtoc.3039 Views: 7788
Reviewed by: Ivan ZanoniYang FuAchille Broggi

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Quantification of Autophagosomes in Human Fibroblasts Using Cyto-ID® Staining and Cytation Imaging
Barbara Hochecker [...] Jörg Bergemann
Jul 5, 2024 1674 Views
Microfluidic Cultures of Basal Forebrain Cholinergic Neurons for Assessing Retrograde Cell Death by Live Imaging
Srestha Dasgupta [...] Wilma J. Friedman
Jan 5, 2025 1888 Views
Real-time IncuCyte® Assay for the Dynamic Assessment of Live and Dead Cells in 2D Cultures
Arlene K. Gidda [...] Sharon M. Gorski
Feb 5, 2025 2943 Views
Abstract
Planarians are freshwater flatworms, well known for their ability to regenerate a complete organism from any piece of their body. Furthermore, planarians are constantly growing and degrowing throughout their lives, maintaining a functional and proportioned body. These properties rely on the presence of a population of adult stem cells and on the tight control of their cell renewal, which is based on the balance between the proliferation of new cells and their differentiation, and the death of unnecessary cells. Due to the importance of these two processes in planarian biology, over the years, researchers have optimized molecular techniques to detect both cell proliferation and cell death in planarians. Here, we present the two main protocols currently used for cell death detection and quantification in the planarian field: Caspase-3 activity quantification and TUNEL assay.
Keywords: Caspase-3 activityBackground
Cell renewal in adult organisms is a complex mechanism based on three processes: (a) the elimination of selected cells by cell death; (b) the replacement of eliminated cells through cell division, typically involving adult stem cells and their descendants; and (c) the differentiation of newly generated cells and their integration with preexisting tissue (Pellettieri and Sanchez Alvarado, 2007; González-Estévez and Saló, 2010). In planarians, cell renewal must be continuously coordinated, since they grow and degrow depending on food availability and temperature (Baguñá and Romero, 1981). It is known that the changes in size result mainly from changes in cell number, rather than in cell size, so the ratio of dying/proliferating cells is controlled by environmental conditions (González-Estévez and Saló, 2010). Planarians are able to tolerate long starvation periods, and during this time, they degrow up to minimum sizes. Under these stressful conditions, food reserves from gastrodermis and mesenchyma are the firsts to be used, and at more extreme points, the sexual strains digest the sexual organs, and become asexual (González-Estévez and Saló, 2010; Miller and Newmark, 2012). When food is available, planarians are able to grow back, and in the sexual strains, the reproductive organs reappear. These cycles of grow and degrow occur throughout planarian lives without damage to the animal.
During planarian starvation, cell death increases to re-organize the organs and structures, and planarian adult stem cells (neoblasts) self-renewal is maintained at basal levels, resulting in a decrease of planarian body size (Figures 1A and 1B) (González-Estévez et al., 2012). The tissue remodeling is critical during planarian starvation because it maintains a proportioned planarian body. It was shown that JNK signaling, and Gtdap-1 are controlling the planarian body re-scaling during degrowing through the modulation of apoptotic cell death (González-Estévez et al., 2007; Almuedo-Castillo et al., 2014).
Because cell death is a relevant process in planarian, in the last few years molecular techniques to detect and measure cell death have been developed and optimized. Here, we will explain step-by-step the two main protocols used to detect cell death in planarians: measurement of Caspase-3 activity and TUNEL assay.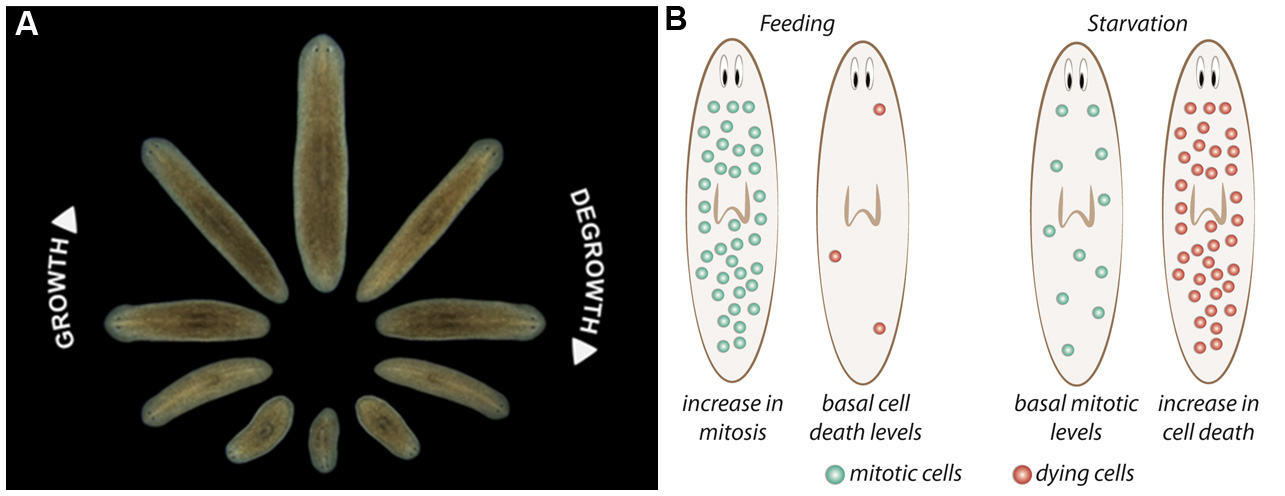
Figure 1. Planarian homeostasis. A. Planarians are able to grow and degrow during their lives, maintaining their body proportions and functionality. Image provided by Gustavo Rodriguez-Esteban. B. After a stimulus, proliferation and/or cell death can change in planarians. After feeding, neoblast proliferation increases throughout planarian body, and cell death is reduced to minimum levels, resulting in the increase of animal’s size. Conversely, when planarians are in starvation, neoblast proliferation is maintained at basal levels and cell death increases, which not only results in a decrease in body size but allows the reorganization of the tissues. Image from Nídia de Sousa Ph.D. thesis (de Sousa, 2017).
Part I: Caspase-3 activity assay
The Caspase-3 activity assay is a fluorescent assay that detects the activity of Caspase-3 in cell lysates using the fluorogenic substrate acetyl Asp-Glu-Val-Asp 7-amido-4-methylcoumarin (Ac-DEVD-AMC). It is based on the hydrolysis of Ac-DEVD-AMC by Caspase-3, resulting in the release of the fluorescent 7-amino-4-methylcoumarin (AMC). AMC that can be detected using a luminescence spectrophotometer with excitation at 380 nm and emission between 420 nm and 460 nm. Cleavage of the substrate only occurs in lysates in which Caspase-3 is present, which is a gene required for apoptosis; therefore, the amount of AMC produced is proportional to the number of apoptotic cells in the sample.
Materials and Reagents
- Petri dish (VWR, catalog number: 391-0439 )
- Slides (VWR, catalog number: 631-1551 )
- Razor blade (MARTOR, catalog number: NO. 743 )
- Eppendorf tubes (VWR, Eppendorf, catalog number: 700-5239 )
- 15 ml Falcons (LF Equipamentos, catalog number: 166 )
- Spectrophotometry Cuvettes (VWR, catalog number: 634-0677BTU )
- 96-well plate (VWR, Corning, catalog number: 734-1664 )
- MilliQ water
- Ice
- Micro BCA Protein Assay Kit (Thermo Fisher Scientific, PierceTM, catalog number: 23235 )
- Tris-HCl, pH 8 (Sigma-Aldrich, catalog number: 93362 )
- EDTA, pH 8 (Sigma-Aldrich, catalog number: 1233508 )
- Triton X-100 (Sigma-Aldrich, catalog number: X100 )
- HEPES pH 7.5 (Sigma-Aldrich, catalog number: H3375 )
- Glycerol 10% (Sigma-Aldrich, catalog number: G5516 )
- DTT (Sigma-Aldrich, catalog number: 646563 )
- Caspase-3 inhibitor Z-DEVD-FMK (Merck, catalog number: 264155 )
- Caspase-3 substrate Ac-DEVD-AMC (BD Biosciences, PharmingenTM, catalog number: 556449 )
- Lysis buffer (see Recipes)
- Assay buffer (see Recipes)
Equipment
- Oven
- Luminescence spectrophotometer
- Pipettes
- Vortex
- Centrifuge
- Platform shaker
- 4 °C refrigerator
- -20 °C freezer
- 37 °C incubator
Procedure
- Protein extraction (Figure 2A)
- Prepare lysis buffer (see Recipe 1)
- In a Petri dish with ice, place a slide and transfer planarians to the slide. Remove the excess of water and chop planarians into small pieces with a razor blade (removing the mucus from the razor blade with a piece of paper when necessary).
Note: A minimum of 5 planarians with 3-5 mm per condition. - Add 100 μl of lysis buffer per planarian to the slide and use it to aspirate planarian fragments into an Eppendorf tube. Keep it on ice.
- Homogenize by pipetting with a P200 and vortexing for 5 sec, maintaining, whenever possible, the Eppendorf tubes on ice.
- Centrifuge at 13,000 x g for 10 min at 4 °C.
- Transfer the supernatant fraction to a new tube at 4 °C. For long-term storage keep at -20 °C.
- Protein concentration determination using Micro BCA Protein Assay Kit (Figure 2B)
- Perform the necessary BSA protein dilutions to obtain the standard curve following the manufacturer’s recommendations.
- Prepare the working reagent following the manufacturer’s recommendations.
- Dilute your samples 250 times (e.g., 2 μl of sample + 498 μl MilliQ water).
- To each spectrophotometry cuvette, add 500 μl of Working Reagent + 500 μl of diluted sample. This procedure should also be done to the samples used to obtain the standard curve.
- Incubate the spectrophotometry cuvettes in an oven for 1 h at 60 °C.
- Remove the samples from the oven and measure the absorbance (λ = 562 nm) in a spectrophotometer.
Note: All the samples should be read in no more than 10 min. - Construct a standard curve using the values of the protein dilutions performed in Step B1. Subtract the value of the blank sample (with no protein, obtained in Step B1) to avoid background measures. Calculate the protein concentration of your samples. Multiply the value obtained for the dilution factor.
- Caspase-3 activity measurement (Figure 2C)
- Prepare the assay buffer (see Recipe 2)
- Load the 96-well plate.
- Start with the blank wells. The blank solution is composed of 125 μl assay buffer + 25 μl of lysis buffer. Blank wells must be done in triplicate.
Note: All the samples loaded in 96-well plate must be done in triplicate. This means that for each condition (blank, negative control and samples) you must have at least three wells. - Load the experimental negative control. For the negative control, protein from a control sample should be used. Load in each well 123 μl of assay buffer + 20 μg of protein (x μl of protein extract + y μl of lysis buffer until 25 μl of volume) + 1.5 μl of Caspase-3 inhibitor Z-DEVD-FMK. Negative controls must be done in triplicate.
Note: Experimental control is used to confirm that you are measuring Caspase-3 activity. Using the Caspase-3 inhibitor is supposed to obtain low levels of Caspase-3 activity. It is essential to have one control but as many as desired can be included. - In the remaining wells load 123 μl of assay buffer + 20 μg of protein (x μl of protein extract + y μl of lysis buffer to 25 μl of final volume). All the samples must be done in triplicate.
- Incubate the plate for 15 min in the dark at 37 °C, in agitation.
- Load 2 μl of Caspase-3 substrate Ac-DEVD-AMC to all plate wells and incubate for 2 h in the dark at 37 °C.
- Measure enzyme activity through luminescence, using a luminescence spectrophotometer (excitation 380 nm, emission 440 nm).
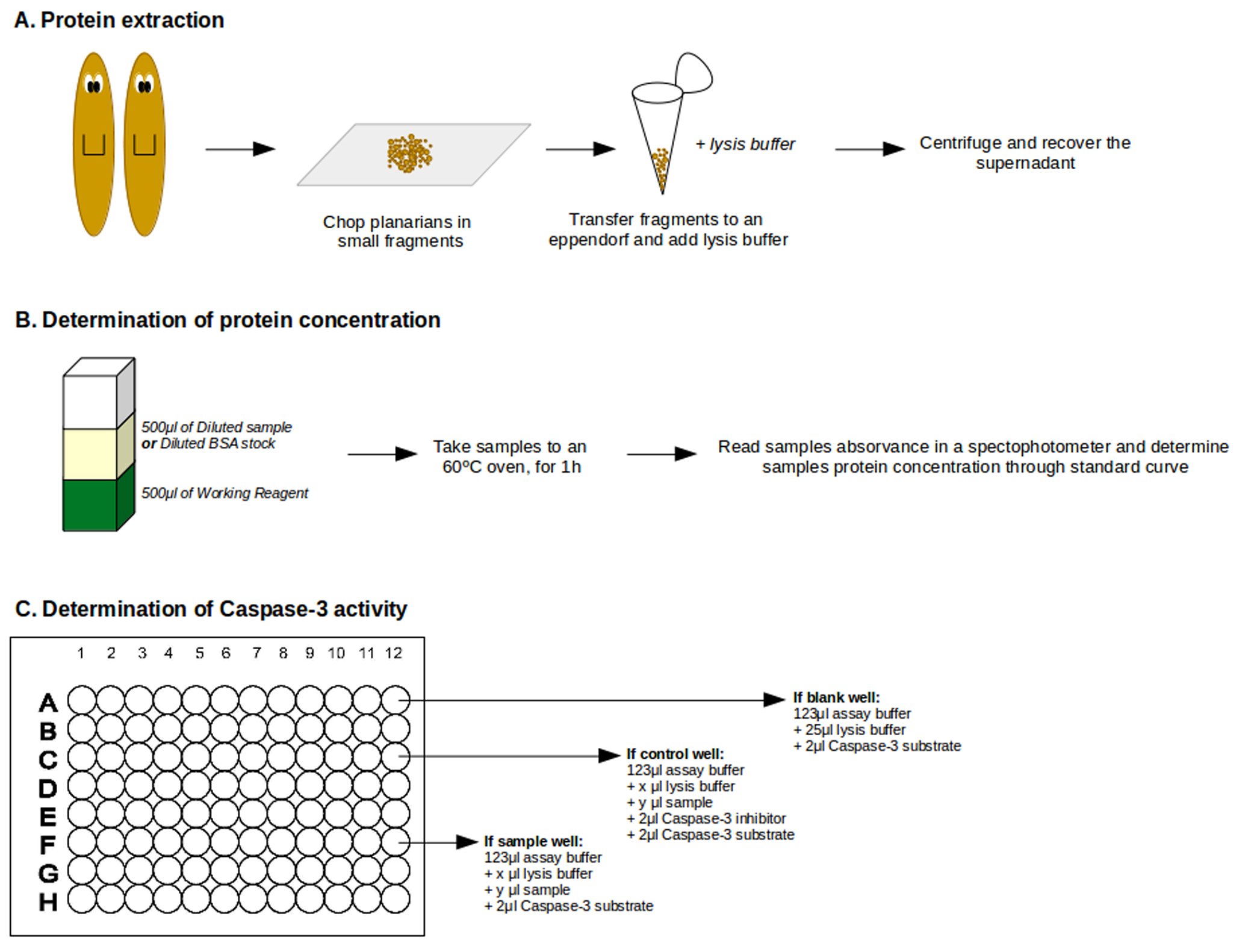
Figure 2. General view of Caspase-3 assay protocol. A. Cartoon exemplifying protein extraction. B. Step-by-Step of protein concentration determination. C. Example of how load plate wells.
Data analysis
- Average the three technical replicates to obtain a single read to each sample.
- Subtract the value of the blank sample to avoid background reads.
- Plot the negative controls, experimental controls, and target samples.
Part II: TUNEL assay
Terminal deoxynucleotidyl transferase (TdT) dUTP Nick-End Labeling (TUNEL) assay has been designed to detect apoptotic cells that present extensive DNA degradation. The method is based on the ability of TdT to label blunt ends of double-stranded DNA breaks independent of a template.
Materials and Reagents
- Glass vials (Sigma-Aldrich, catalog number: 27151 )
- 96-well plate (VWR, catalog number: 734-2781 )
- Glass slide (VWR, catalog number: 631-1551 )
- Coverslip (Merck, catalog number: S7117 )
- ApopTag Red In Situ Apoptosis Detection Kit (Merck, catalog number: S7165 )
- 10% n-acetyl cysteine (diluted in PBS 1x) (Sigma-Aldrich, catalog number: A9165 )
- PBS 1x
- 37% formaldehyde (Merck, catalog number: 104003 )
- 1% SDS (diluted in PBS 1x) (Sigma-Aldrich, catalog number: L3771 )
- 30% H2O2 (Merck, catalog number: 107209 )
- Bleaching solution
- TdT enzyme (from ApopTag Kit)
- Anti-digoxigenin-rhodamine (from ApopTag Kit)
- 70% glycerol (diluted in PBS 1x)
- PBSTx (see Recipes)
- PBSTB (see Recipes)
Equipment
- Platform shaker
- Pipettes
- 37 °C incubator
- 4 °C refrigerator
- Confocal microscope
Software
- ImageJ
Procedure
- Place animals in glass vials and remove the planarian water using a pipette.
Note: Animals should be between 3 mm and 5 mm large. - Incubate the animals with 5 ml of 10% N-acetyl cysteine (diluted in PBS 1x) for 5 min at RT, rocking.
Notes:- Although glass vials are rocking, ensure that planarians are properly separated and in contact with the solution.
- This step kills the animals and removes their thick coating of mucus.
- Remove N-acetyl cysteine and fix the animals with 5 ml of 4% formaldehyde (diluted in PBSTx) for 20 min at RT, rocking.
Note: Every liquid exchange should be performed carefully through pipetting but not touching planarians with the tips, to avoid tissue damage. - Remove the fixative and incubate the animals with 5 ml of 1% SDS (diluted in PBS 1x) for 20 min at RT, rocking.
- Remove the previous solution and wash animals twice with 5 ml of PBSTx for 5 min each at RT, rocking.
- Remove the washing solution and bleach animals with 5 ml of 6% H2O2 (diluted in PBSTx), overnight under white light, without rocking.
Note: An overnight is around 16 h. Please be sure to perform this bleaching time. - Remove the bleaching solution and wash the animals twice with 5 ml of PBSTx for 5 min each at RT, rocking.
Note: On this step, animals can be stored at 4 °C for a few days (no longer than 3 days). - Remove the washing solution and rinse animals in 5 ml of PBS 1x.
- Carefully, transfer the animals to the 96-well plate.
Note: It is important to be sure that all animals in the well are in contact with the solution. If you have several animals, it is better to distribute them for two or three wells. - Using a pipette remove PBS 1x and incubate the animals with 100 μl of TdT enzyme diluted in Reaction buffer (both from the ApopTag Kit) for 4 h at 37 °C, rocking.
- Remove the TdT enzyme and rinse animals with 100 μl of stop/wash buffer (from the ApopTag Kit).
- Remove the previous solution and wash animals twice with 100 μl of PBSTB (PBSTx with 0.25% BSA) for 5 min at RT, rocking.
- Remove the washing solution and stain animals with 100 μl of anti-digoxigenin-rhodamine diluted in Blocking solution (both from the ApopTag Kit) for 4 h at RT, on dark, rocking.
Note: From this step until the end of the protocol, animals should be maintained in the dark. - Remove the previous solution and wash stained animals 4 times with 100 μl of PBSTB each for 10 min at RT, rocking. Keep washing overnight in PBSTB at 4 °C, rocking.
- Wash animals until obtain a clear signal (Figure 3) with PBSTB at RT, rocking.
Note: Change the solution every 20/30 min. - Remove the previous solution and add 70% glycerol to the animals.
- Transfer the animals to a glass slide and mount with a coverslip. Add glycerol if necessary.
Note: To avoid glycerol evaporation and sample disruption, apply transparent nail varnish around the coverslip. Be aware to not apply varnish over the animals.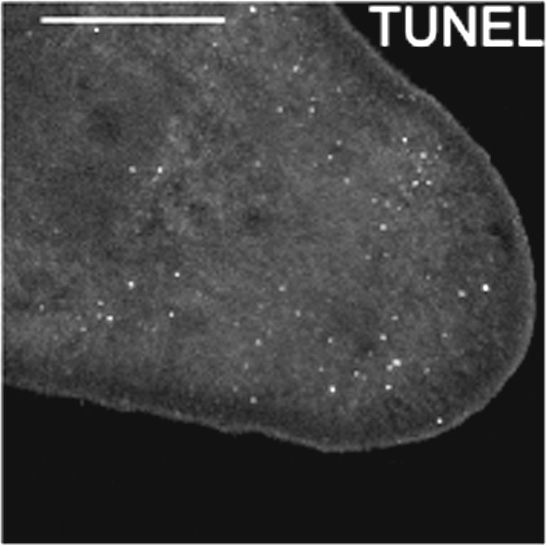
Figure 3. Result of a TUNEL assay. The image shows the tail of a planarian imaged by confocal microscopy. White dots correspond to nucleus of dying cells. Scale bar = 250 μm.
Data analysis
- Acquire images using a confocal microscope.
- Analyse the images in ImageJ software.
Recipes
- Lysis buffer
Tris-HCl (pH 8) 5 mM
EDTA (pH 8) 20 mM
Triton 0.5%
Reserve the lysis buffer on ice until it is necessary - Assay buffer
HEPES (pH 7.5) 20 mM
Glycerol 10%
DTT 2 mM
Reserve the assay buffer at RT until it is necessary - PBSTx
1x PBS with 0.3% Triton - PBSTB
PBSTx with 0.25% BSA
Acknowledgments
This work was supported by grant BFU2008-01544 and BFU2014-56055-P (Ministerio de Educación y Ciencia) and grant 2009SGR1018 (AGAUR). N.S. was supported by the APIF fellowship from the Universitat de Barcelona. Caspase-3 activity measurement and TUNEL assay protocol were adapted from the González-Estévez et al. (2007) and Pellettieri et al. (2010), respectively.
Competing interests
The authors declare no conflicts of interest or competing interests.
References
- Almuedo-Castillo, M., Crespo-Yanez, X., Seebeck, F., Bartscherer, K., Salo, E. and Adell, T. (2014). JNK controls the onset of mitosis in planarian stem cells and triggers apoptotic cell death required for regeneration and remodeling. PLoS Genet 10(6): e1004400.
- Baguñá, J. and Romero, R. (1981). Quantitative analysis of cell types during growth, degrowth and regeneration in the planarians Dugesia mediterranea and Dugesia tigrina. Hydrobiologia 84: 181-194.
- de Sousa, N. (2017). Ph.D. Thesis. Role of Hippo pathway in planarians. University of Barcelona.
- González-Estévez, C. and Salo, E. (2010). Autophagy and apoptosis in planarians. Apoptosis 15(3): 279-292.
- González-Estévez, C., Felix, D. A., Aboobaker, A. A. and Salo, E. (2007). Gtdap-1 promotes autophagy and is required for planarian remodeling during regeneration and starvation. Proc Natl Acad Sci U S A 104(33): 13373-13378.
- González-Estévez, C., Felix, D. A., Rodriguez-Esteban, G. and Aboobaker, A. A. (2012). Decreased neoblast progeny and increased cell death during starvation-induced planarian degrowth. Int J Dev Biol 56(1-3): 83-91.
- Miller, C. M. and Newmark, P. A. (2012). An insulin-like peptide regulates size and adult stem cells in planarians. Int J Dev Biol 56(1-3): 75-82.
- Pellettieri, J. and Sanchez Alvarado, A. (2007). Cell turnover and adult tissue homeostasis: from humans to planarians. Annu Rev Genet 41: 83-105.
- Pellettieri, J., Fitzgerald, P., Watanabe, S., Mancuso, J., Green, D. R. and Sanchez Alvarado, A. (2010). Cell death and tissue remodeling in planarian regeneration. Dev Biol 338(1): 76-85.
Article Information
Copyright
© 2018 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Sousa, N. D. and Adell, T. (2018). Detection of Cell Death in Planarians. Bio-protocol 8(19): e3039. DOI: 10.21769/BioProtoc.3039.
Category
Developmental Biology > Cell growth and fate > Proliferation
Cancer Biology > Cell death > Animal models
Cell Biology > Cell viability > Cell death
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link