- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Artificial Inoculation of Epichloë festucae into Lolium perenne, and Visualisation of Endophytic and Epiphyllous Fungal Growth
Published: Vol 8, Iss 17, Sep 5, 2018 DOI: 10.21769/BioProtoc.2990 Views: 10075
Reviewed by: Amey RedkarCynthia L. MonacoSatyabrata Nanda

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

In Vitro Hyphal Branching Assay Using Rhizophagus irregularis
Takaya Tominaga and Hironori Kaminaka
Aug 20, 2024 2385 Views
Silencing Arbuscular Mycorrhizal Fungal Gene Using Chitosan Nanoparticle-Mediated dsRNA Delivery System
Chumei Yan [...] Xianan Xie
Jun 5, 2025 2697 Views

A Reliable In Planta Inoculation and Antifungal Screening Protocol for Rhizoctonia solani-Induced Sheath Blight in Rice
Alinaj Yasin [...] Palash Deb Nath
Nov 5, 2025 1618 Views
Abstract
Natural hosts for the fungal endophyte Epichloë festucae include Festuca rubra (fine fescue) and Festuca trachyphylla (hard fescue). Some strains also form stable associations with Lolium perenne (perennial ryegrass). L. perenne is a suitable host to study fungal endophyte–grass interactions, such as endophytic fungal growth within the plant and epiphyllous growth on the plant surface. Here we provide a detailed protocol based on work by, for artificial inoculation of E. festucae into L. perenne, and newly developed staining and visualization techniques for observing endophytic and epiphyllous hyphae and the expressorium, an appressorium-like structure used by the fungus to exit the plant. The staining method uses a combination of glucan binding aniline blue diammonium salt (AB) and chitin binding wheat germ agglutinin-conjugated Alexa Fluor®488 -(WGA-AF488). This protocol will be a useful tool to study Epichloë-grass interactions, particularly the comparison of different Epichloë-grass associations, various endophyte-host developmental stages, as well as the analysis of mutant Epichloë strains.
Keywords: ExpressoriumBackground
Latch and Christensen (1985) developed a protocol for artificial infection of grasses with Epichloë endophytes. This work has been the basis for the extensive E. festucae Fl1–L. perenne symbiosis research (reviewed in Scott et al., 2012). In our recent work, we described a newly identified fungal structure, the expressorium that enables E. festucae to exit L. perenne (Becker et al., 2016). For visualization of fungal and plant structures, we used two different fluorophores. Wheat germ agglutinin-conjugated Alexa Fluor®488 (WGA-AF488) binds to sialic acid and N-acetylglucosamine residues and is commonly used as an indicator for chitin (Figueroa-Lopez et al., 2014). Aniline blue diammonium salt (AB) is commonly used as a stain for β-D-1,3-glucans, such as callose at plant cell walls and related glucans in fungal cell walls (Wood and Fulcher, 1984). Staining of ethanol and potassium hydroxide (KOH) cleared (i.e., fixed) plant samples with AB and WGA-AF488, as per our protocol described below, enables a comprehensive visualisation of (i) Epichloë endophytic and epiphytic hyphae, (ii) plant responses based on callose depositions, and (iii) the plant cuticle. By collecting a combination of light emitted from the dyes described here and autofluorescence from the host plant, we can observe a strong signal from the plant cuticle. This overcomes the need for a specific dye for lipophilic components, which might interfere with WGA-AF488 or aniline blue imaging. Interestingly, WGA-AF488 exclusively stains the cell wall of epiphyllous Epichloë hyphae and is restricted to hyphal septa and actively growing tips of endophytic hyphae, while AB is used to stain the endophytic hyphal cell wall and plant glucans. Propidium iodide (PI), which stains plant and fungal nuclei, can also be used in combination with WGA-AF488 and AB without the need for changing the confocal settings described below.
Materials and Reagents
Seed sterilization and inoculation:
- 1.5 ml reaction tubes (BRAND, catalog number: 780400 )
- Parafilm (Sigma-Aldrich, catalog number: P7793-1EA )
- Petri dishes (SARSTEDT, catalog number: 82.1472 )
- Household aluminum foil, or heavy duty foil for multiple uses
- Sterile filter paper (GE Healthcare, catalog number: 1001-090 )
- Potting mix (e.g., Klasmann and Deilmann, Substrat 1.090)
- Microscope slides (Carl Roth, catalog number: 0656.1 )
- Microscope slides coverslips (Carl Roth, catalog number: 0657.2 )
- Replacement blade No.11 for metal scalpel 10621 (DAHLE, catalog number: 10722 )
- Root trainers or pots (Gärtner Pötschke, catalog number: 280686 )
- Grass seeds (used here: Lolium perenne cv Samson seeds, endophyte free; Agricom, New Zealand)
- An Epichloë strain, E. festucae Fl1 (ATCC, catalog number: MYA-3407 )
- Autoclaved distilled water
- Murashige and Skoog Medium including vitamins (Duchefa Biochemie, catalog number: M0222 )
- Phyto agar (Duchefa Biochemie, catalog number: P1003 )
- Potato Dextrose Agar (Carl Roth, catalog number: X931 )
- Agar-Agar (Carl Roth, catalog number: 6494 )
- Sodium chloride (NaCl) (Sigma-Aldrich, catalog number: 793566 )
- Potassium chloride (KCl) (Sigma-Aldrich, catalog number: 746436 )
- Disodium hydrogen phosphate (Na2HPO4) (Sigma-Aldrich, catalog number: 795410 )
- Potassium dihydrogen phosphate (KH2PO4) (Sigma-Aldrich, catalog number: 795488 )
- Sulfuric acid (H2SO4) (Sigma-Aldrich, catalog number: 339741 )
- Sodium hypochlorite (NaClO + H2O; 12% Cl2) (Carl Roth, catalog number: 9062.3 )
- Potassium hydroxide (KOH) (Carl Roth, catalog number: 6751 )
- 95% Ethanol (Carl Roth, catalog number: 9065 )
- Tween 20 (Carl Roth, catalog number: 9127 )
- Glycerol (Carl Roth, catalog number: 7530 )
- Aniline Blue diammonium salt (C37H32N5O9S3) (Sigma-Aldrich, catalog number: 415049 )
- Wheat Germ Agglutinin, Alexa FluorTM 488 Conjugate (Thermo Fisher Scientific, catalog number: W11261 )
- Propidium iodide solution 0.5 mg/ml (Sigma-Aldrich, catalog number: P4864 )
- 3% water agar (see Recipes)
- 2.4% potato dextrose agar (see Recipes)
- Murashige and Skoog phyto agar (see Recipes)
- Aniline Blue diammonium salt stock solution (see Recipes)
- WGA-AF488 stock solution (see Recipes)
- PBS (pH 7.4) (see Recipes)
- Tween 20 stock (see Recipes)
- Staining Solution (see Recipes)
Equipment
- Fridge
- Incubator (Thermo Fisher Scientific, Heraeus, model: BK 600 )
- Stereomicroscope (Leica, model: Leica S6 D )
- Metal scalpel (DAHLE, catalog number: 10621 )
- Preserving jar
- Autoclave
- Stainless steel forceps (Sigma-Aldrich, catalog number: Z168777 )
- Thermos Scientific “Mr. Frosty” Freezing Container (Thermo Fisher Scientific, catalog number: 5100-0001 )
- Laminar air flow cabinet
- Confocal Laser Scanning Microscope, CLSM, (Leica, models: Leica TCS SP5 and Leica TCS SP8 )
- Desiccator Base (DWK Life Sciences, DURAN®, catalog number: 247714605 )
- Desiccator Lid (DWK Life Sciences, DURAN®, catalog number: 244204605 )
- Stopcock with PTFE Spindle (side) (DWK Life Sciences, DURAN®, catalog number: 247980306 )
- Stopcock with PTFE Spindle (top) (DWK Life Sciences, DURAN®, catalog number: 247990401 )
- Water jet filter pump (BRAND, catalog number: 159600 )
Procedure
- Seed surface sterilization (adapted from Latch and Christensen, 1985)
- Soak the seeds in sulfuric acid (H2SO4) 50% (v/v) for 30 min.
- Rinse the seeds in tap water 3 times for 2 min each time.
- Soak the seeds in NaClO 13% (w/v) for 30 min.
- Work under sterile conditions from now on.
- Rinse the seeds in sterile distilled water. 3 times for 2 min each time.
- Place the seeds on sterilized filter paper and air dry in a sterile Petri dish under the laminar flow
Note: This will take approximately 2 h. - Dry seeds can be stored in the sterile Petri dish sealed with Parafilm in the fridge at 4 °C for up to 3 months. However, note that the germination rate of the seeds may decline after longer storage (Figure 1A).
- Preparation of perennial ryegrass seedlings adapted from (Latch and Christensen, 1985)
- Place the surface sterilized seeds on Petri dishes containing 3% (w/v) water agar. Up to 10 seeds per plate can be added. Ensure the seeds are orientated in the same direction. Inoculation is easiest when the seeds are placed with the concave half facing up and the wedge-shaped rachilla segment at the bottom (Figure 1B). Each floret is born on a segmented central axis called the rachilla, and each seed retains its rachilla segment.
- Place the seeds at 20-22 °C in the dark by wrapping the Petri dishes in aluminum foil. Ensure that the plate is positioned on its side, with the seeds at the bottom and that the plates remain in that position throughout seed germination.
- Subculture the fungal strains for inoculation onto a fresh, thick PDA plate. Do not put more than 4 agar pieces on a 12 cm Petri dish. Seal the dishes with Parafilm and incubate them at 20 °C with the bottom facing downwards (Figure 1C). For storing the fungal strains long term, grind 1 cm2 of freshly grown mycelia in 2 ml PD Broth. Let the mycelia grow for 3-5 days at room temperature. Add 2.5 ml of 30% glycerol and dispense 750 µl aliquots of mycelia into Eppendorf tubes. Freeze cultures slowly and evenly using a freezing container (e.g., Mr. Frosty). Store tubes at -80 °C.
- After 7-8 days, seed germination will have taken place and shoot lengths will range from 4 cm to 7 cm (Figure 1D).
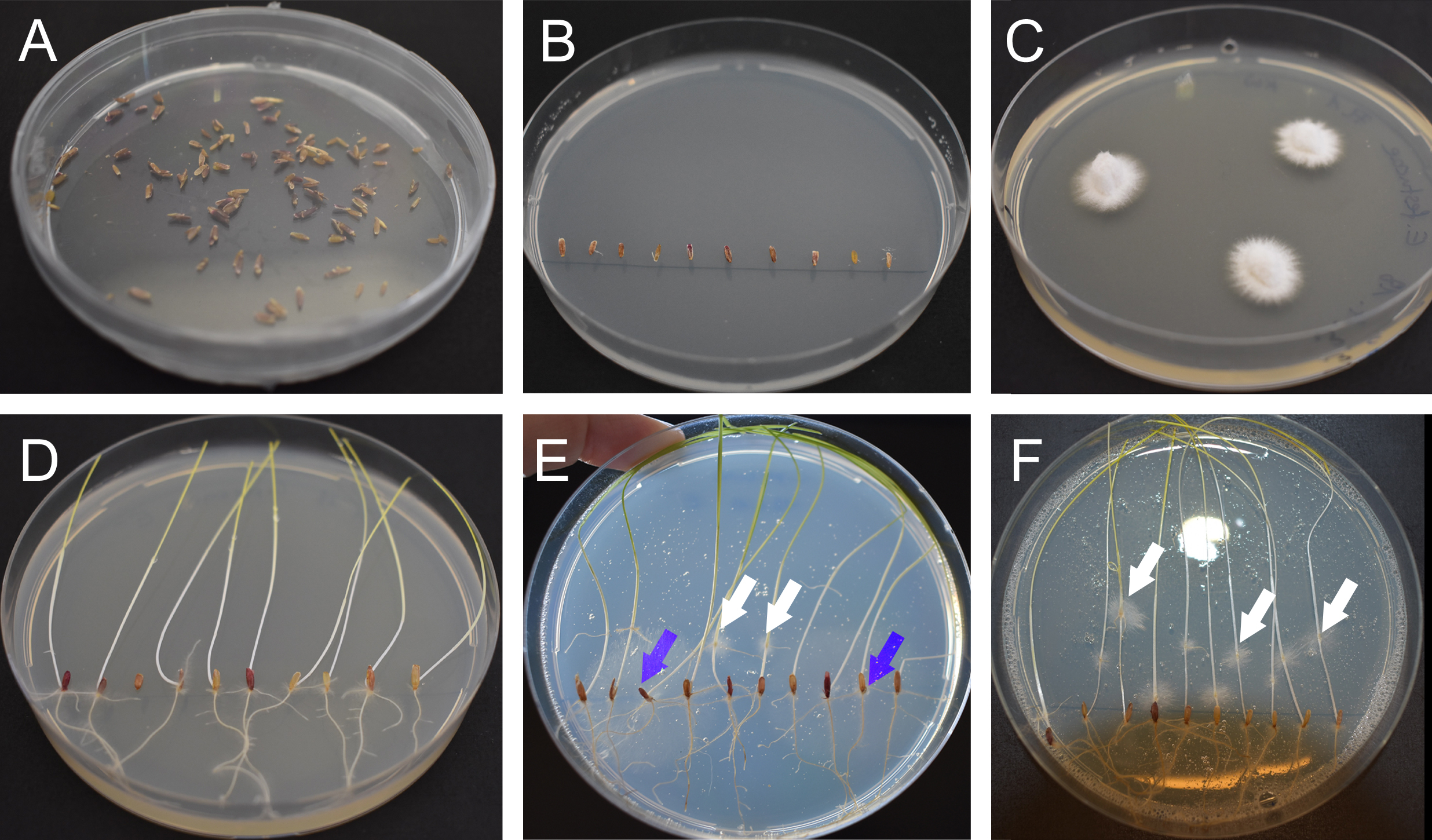
Figure 1. Preparation of perennial ryegrass seeds and Epichloë mycelium. A. Sterilized and dried seeds. B. Seeds placed on 3% water agar. C. E. festucae Fl1 mycelium on PDA agar grown for one week at 20 °C. D. Germinated ryegrass after one week on 3% water agar. E. Contaminated plate after inoculation. The white arrows indicate E. festucae Fl1 mycelium growing out from inoculation points, the blue arrows indicate contaminations growing out from the seeds. F. Non-contaminated plate one-week post inoculation. The white arrows indicate E. festucae Fl1 mycelium.
- Inoculation of perennial ryegrass seedlings (adapted from Latch and Christensen, 1985)
- Using a sterile scalpel and a dissecting microscope (Figure 2A), cut a 2-3 mm longitudinal slit at the developing apical meristem in between the seedling mesocotyl and coleoptile. Seedlings remain on water agar throughout this process. Use a sharp scalpel blade and replace this when it becomes blunt, approximately every 20-30 seedlings.
- Insert a small piece (approx. 2 mm x 2 mm) of freshly grown mycelia cut from the colony edge into the slit of the seedling (Figure 2B). Once you get used to the procedure and are able to work fast (mycelium should not dry out), you can place several pieces of mycelium onto the blade and insert them one by one into the slits of the seedlings. Steps C1 and C2 are shown in Videos 1 and 2.Video 1. Mycelia blocks cut from the colony edgeVideo 2. Mycelia is placed on a scalpel blade and inserted directly after cutting a small slit in the apical meristem
- Incubate the seedlings on the water agar plates at 20-22 °C for a further 7 days in the dark (Figure 1F). Check the seedlings for contamination after 2 days (Figure 1E). E. festucae colonies grow out from the inoculation point, while faster growing fungal contaminations grow out from the seeds. Transfer the seedlings to light (photoperiod of 16 h light) and incubate them for a further 7 days at 20-22 °C. Seal Petri dishes with Parafilm.
- Transfer the seedlings into sterile pots (e.g., preserving jar) containing Murashige and Skoog Phyto agar (Figure 2C). Using forceps gently push the seed and roots into the agar. Or: Transfer the seedlings to root trainers containing potting mix (Figure 2D).
- Place the plants in a greenhouse or an environmentally controlled growth room at 20-22 °C with a photoperiod of 16 h of light (approximately 100 µmol m-2 sec-1). Check the plants in sterile pots regularly for contamination.
- After 6-8 weeks (Figure 2E), or when more than one new grass shoot, a tiller (Figure 3A), has developed, check that the plants are infected by either aniline blue staining or by immunoblotting (Florea et al., 2015). Transplant the seedlings into single pots for further growth.
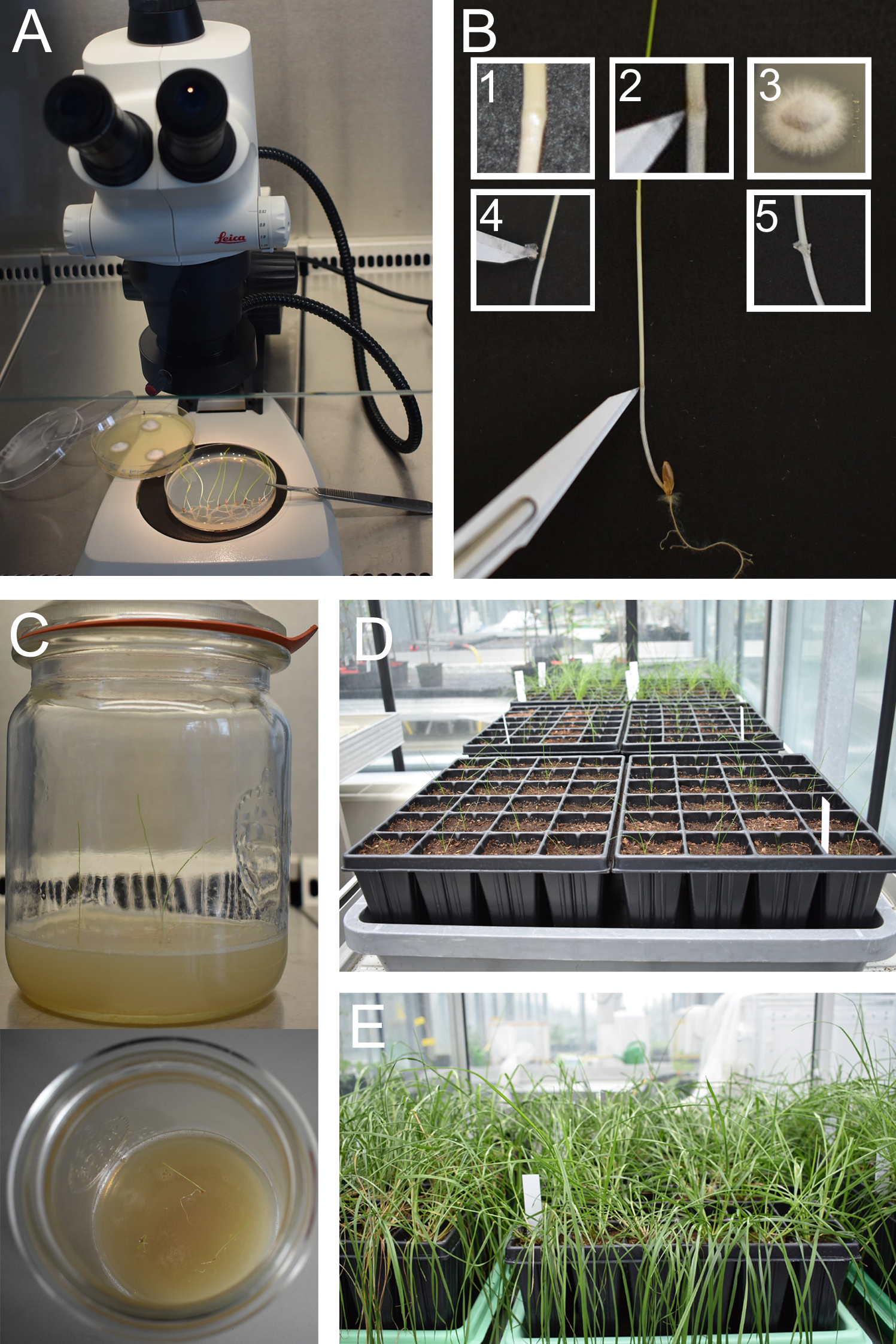

Figure 2. Inoculation of perennial ryegrass seedlings and plant cultivation. A. Set up a dissecting microscope under a laminar flow cabinet. B. Inoculate a small piece of E. festucae mycelium into a small incision made at the SAM, visible as a small knot. C and D. Two weeks after inoculation (one week in the dark, one week in light) plant seedlings in sterile jars or in root trainers in potting mix. E. Check for infection when several tillers have developed.
- Staining of endophytes in perennial ryegrass for confocal laser scanning microscopy (CLSM)
- Cut the tillers close to the root of infected ryegrass plants (Figure 3A).
- Peel the outermost leaf sheath from the tiller and discard (Figure 3B), cut an approx. 2 cm sized piece from the base of the pseudostem and discard this, too. Cut a 2-3 cm piece of the pseudostem (Figure 3C 1). Cut this section in half longitudinally (Figure 3D).
- Cut a 2 cm piece around the ligule (Figure 3C 2).
- Cut a 2 cm piece from the blade (Figure 3C 3).
- Place the plant tissue into an Eppendorf tube containing 95% EtOH and soak them overnight at 4 °C (Figure 3D). Steps D2-D5 are shown in Video 3. If the leaves are still dark green, transfer them into fresh 95% EtOH to clear the plant sample from excessive chlorophyll. Video 3. Preparing leaf samples for clearing and staining
- Remove the ethanol and treat with 10% KOH for 3 h. KOH solubilizes plant cell wall polysaccharides, this softens the plant cell wall and allows penetration of the WGA-AF488, AB, and PI.
- Thoroughly wash the tissue three times with 1 ml of PBS (pH 7.4); any remaining KOH will ruin the staining procedure.
- Incubate the samples in freshly prepared staining solution for 30 min (Figure 3E), including a vacuum infiltration step (Figures 3F and 3G; Video 4). Open the lid of the 1.5 ml cups during the vacuum infiltration procedure. In order to prevent damage to the desiccator, the lid should be carefully slid on and off instead of being directly placed onto the apparatus. Vacuum infiltrate the samples for 2 min at approximately 16 mbar. (According to the manufacturer, the ultimate pressure of 16 mbar [water temperature 12 °C] is reached across a wide range of water supply). You can check the vacuum by carefully trying to take off the lid of the chamber. After two min slowly ventilate the chamber, to avoid destroying the plant tissue. Repeat the vacuum infiltration and ventilation step two more times. The vacuum infiltration procedure is shown in Video 4. Note that we sped up incubation time of the samples in the video.
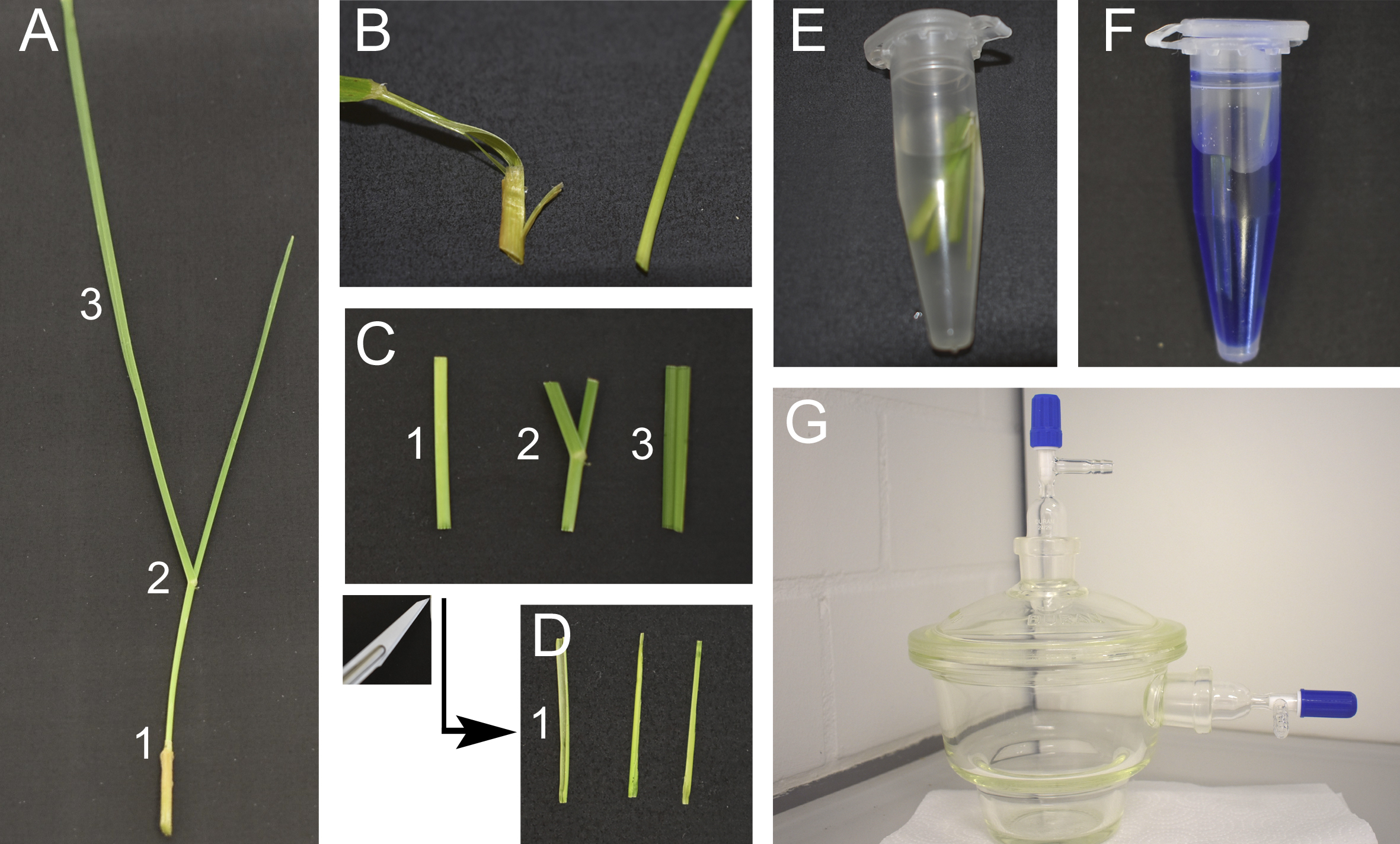

Figure 3. Staining of endophytes in perennial ryegrass for CLSM. A. Cut tillers close to the root. Tillers can be separated in three parts, the pseudostem (1), the area around the ligule containing the blade meristem (2) and the blade (3). B. Peel the outermost leaf sheath and discard. C. Cut a 2 cm sized piece from the base of the pseudostem (1), the area around the ligule (2) and from the blade (3). D. Cut the pseudostem section (1) in half longitudinally. E. Incubate the leaf samples in 95% ethanol overnight and treat with 10% KOH for 3 h, then wash them three times with PBS. F. Prepare the staining solution. G. Vacuum infiltrate the samples.Video 4. Vacuum infiltration for staining of endophytes in grass. Seal the lid by turning the upper stopcock. Open the side stopcock connecting the chamber to the water jet pump. Turn the water on, vacuum is now applied to the chamber. Check after a few seconds that vacuum is established, you cannot slide the lid to the left or right. Wait 2 min. For ventilation of the chamber, close the side stopcock in order to avoid water being sucked into the chamber. Turn the water off. Open the upper stopcock on the lid slowly. You should hear a hissing sound when air flows into the chamber. Repeat the procedure two more times.
- Visualization by CLSM
- Mount the samples on slides with coverslips for confocal microscopy. Add a droplet of 50% glycerol for long-term observations to prevent the samples from drying out. Glycerol has a different refractive index than water, for short-term observations use water instead of glycerol. Using forceps carefully place a leaf sample on the slide. Leaves tend to fold, so you might need to use two forceps to carefully spread them apart. Preparation under a dissecting microscope is advisable.
- Excite AB, WGA-AF488 (and PI) simultaneously with the 488 nm argon ion and 561 nm diode-pumped solid-state (DPSS) lasers.
- Use three detectors to capture the emission fluorescence from the dyes as well as plant autofluorescence. Use blue pseudocolor (PMT1, 498-551 nm) for emission fluorescence from WGA-AF488 excited with the 488 nm argon ion laser. Use green (PMT2 or HyD1, 593-625 nm) and red (PMT3, 661-796 nm) pseudocolors for emission fluorescence from aniline blue, plant autofluorescence and propidium iodide resulting from excitation with the 561 nm DPSS laser (Figure 4).
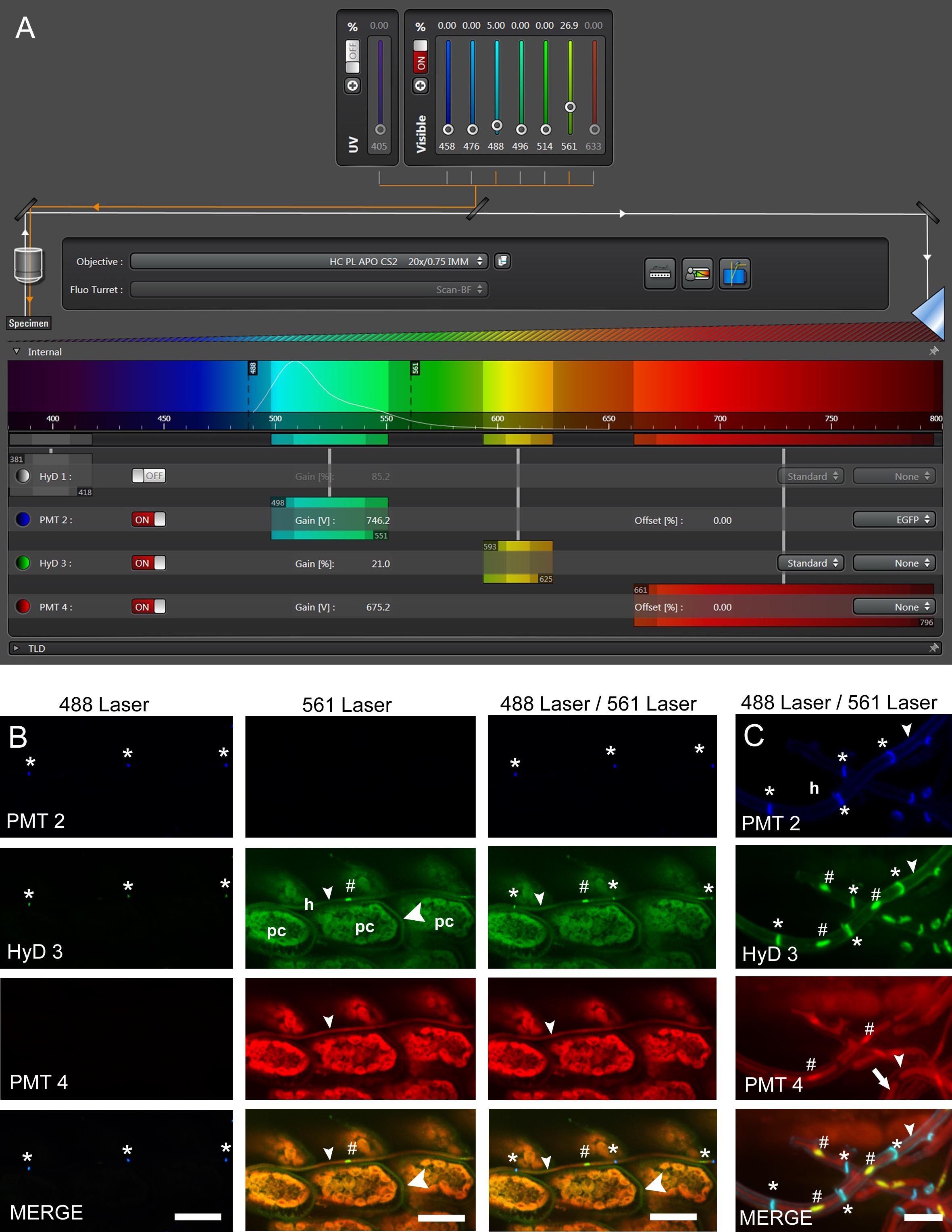
Figure 4. Confocal laser scanning microscopy (CLSM) of E. festucae in hard fescue. A. Settings for CLSM (here using a Leica TCS SP8 microscope) of samples stained with WGA-AF488, aniline blue and propidium iodide. B and C. CLS micrographs depicting the different and merged fluorescence signals captured with detectors (PMT 2, PMT 4 and HyD 3) that collect light of defined wave lengths (as shown in A). The laser used to excite the samples is also indicated. (B) Endophytic hyphae (h) of E. festucae Fl1 in between plant cells (pc) of Festuca trachyphylla leaf blade. The emission fluorescence from WGA-AF488 excited by the 488 nm argon ion laser is captured with the PMT 2 and also the HyD 3 detector by crossover of fluorescence emission. WGA-AF488 stains chitin only present in septa (asterisk) of endophytic hyphae, plant cells are not stained. The emission fluorescence from AB, PI and plant (chlorophyll) autofluorescence excited by the 561 nm DPSS laser is captured with the HyD 3 detector. PI stains nuclei (hash) of endophytic hyphae. AB stains β-D-1,3-glucans in fungal (small arrowhead) and plant cell walls (big arrowhead). Autofluorescence of chlorophyll and fungal cytoplasm is captured with the HyD 3 and PMT 4 detector. Note that the green (HyD 3) and red (PMT 4) pseudocolors are combined to orange in the merged images. (C) Epiphyllous hyphae (h) of E. festucae Fl1 on the Festuca trachyphylla blade. WGA-AF488 stains chitin in septa (asterisk) and the cell wall (small arrowhead) of epiphyllous hyphae. The weak WGA-AF488 signal captured with PMT 2 indicates that the hyphae shown here just start unmasking and/or accumulating chitin at the cell wall. The HyD 3 detector captures the PI nuclei signal (hash), the WGA-AF488 signal of septa and cell wall (asterisk) and the AB stained fungal cell wall (small arrowhead). The PMT 4 detector captures the AB stained fungal cell wall (small arrowhead) and the fungal cytoplasm (arrow). Scale bars: B = 20 µm, C = 5 µm.
Data analysis
- Image processing (LAS AF and ImageJ) is used to enhance the quality of images. We adjust the contrast and brightness of the merged image if necessary.
- The protocol can be used to visualize early infection stages (Figure 5), endophytic and epiphyllous hyphae as well as expressoria (Figure 6), and callose depositions of plants (Figure 7). Nuclei are visualized by addition of propidium iodide to the staining solution (Figures 6C and 6E).
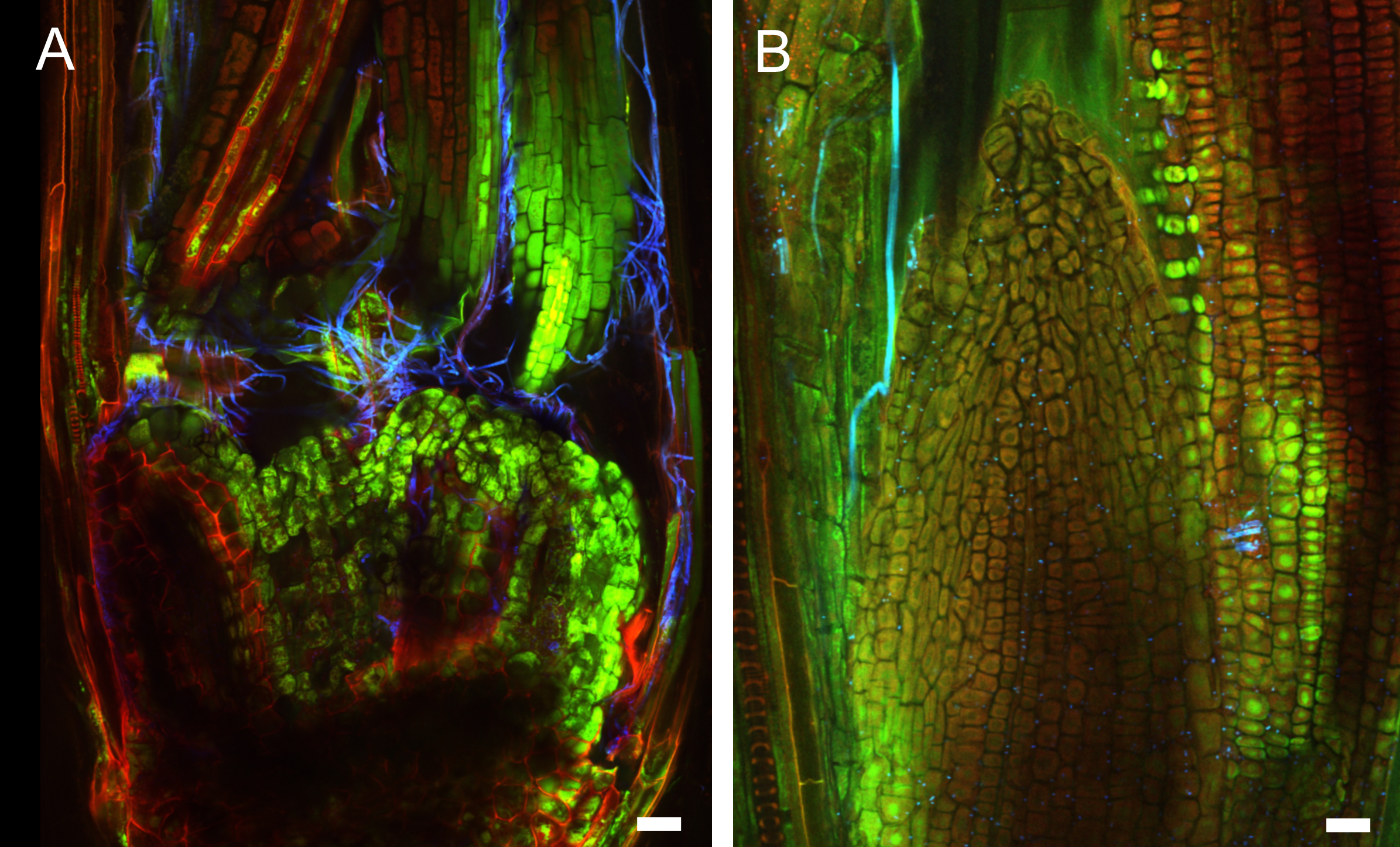
Figure 5. Early infection of perennial ryegrass by E. festucae Fl1. Samples were taken two weeks after inoculation and stained with WGA-AF488 and AB. A. Image taken at the inoculation side. WGA-AF488 stains chitin in epiphyllous hyphae as well as septa of endophytic hyphae of E. festucae. B. Endophytic hyphae colonizing a leaf primordium. Cell wall glucan of endophytic hyphae stained with AB is shown in orange, septa and epiphyllous hyphae are shown in blue. Scale bars = 20 µm.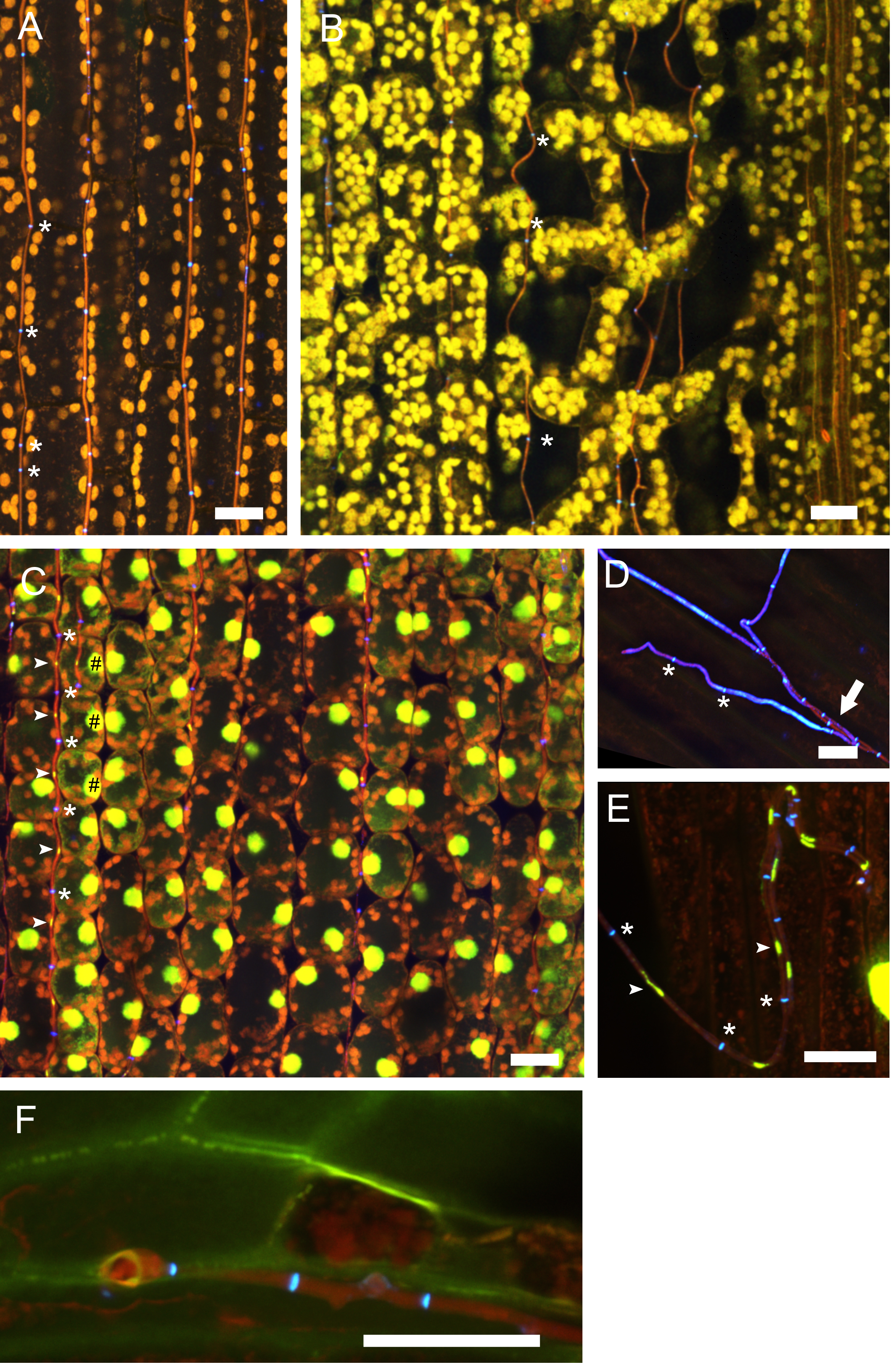
Figure 6. Endophytic and epiphyllous hyphae of E. festucae associated with perennial ryegrass. Samples were taken two weeks after inoculation and stained with WGA-AF488 and AB (A, B, D, F) and WGA-AF488, AB and PI (C, E). A. Endophytic hyphae in the leaf sheath. The cell walls of endophytic hyphae are stained with AB and the chitin in septa is stained with WGA-AF488, shown in blue, and marked by asterisks. Autofluorescence of the plant chloroplasts is shown in orange. B. Endophytic hyphae in the leaf blade; note that the hyphae are not straight between the spongy mesophyll. Staining and pseudocolors are as in A. C. Endophytic hyphae in the blade, stained additionally with PI for visualization of nuclei. Fungal nuclei are in yellow and marked by arrows. Plant nuclei are shown in yellow and marked by hash symbols. D. Cell walls of the endophytic hyphae are not stained by WGA-AF488 and so are hyphae after emergence from the plant stained by AB, shown in red pseudocolor and marked by an arrow. Cell walls and septa of epiphyllous hyphae on the leaf sheath contain chitin stained with WGA-AF488, shown in blue, septa are marked by asterisks. E. Epiphyllous hyphae on the leaf blade, fungal nuclei are stained additionally with PI (arrows). Note that the hyphal cell walls are chitin-free shortly after emerging on the outer surface and therefore only septa bind WGA-AF488 and appear blue (asterisks), while the fungal cell wall is stained by AB and shown in red. F. Expressorium near blade meristem rupturing the cuticle, which shows green autofluorescence. Scale bars = 20 µm.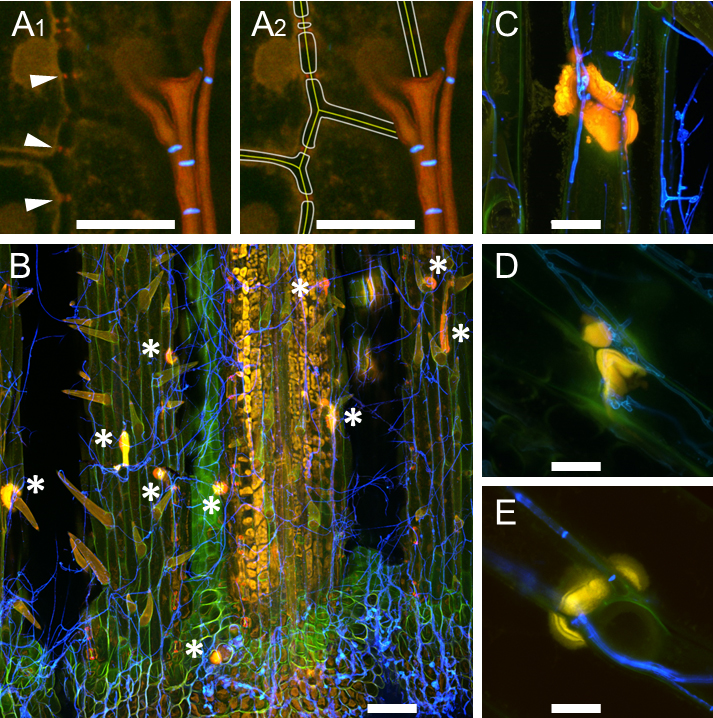
Figure 7. Callose production at cell walls of perennial ryegrass. Samples were taken eight weeks after inoculation and stained with WGA-AF488 and AB. A. E. festucae-independent callose production at plasmodesmata; endophytic hyphae of E. festucae wild type Fl1 are shown in orange, septa in blue; A1. Arrowheads pointing to callose; A2. Artist’s view of A1, in which plant middle lamella and cell walls were added graphically. B. E. festucae ∆noxA mutant inducing formation of callose papilla (asterisk) where a hypha exits the plant; epiphyllous hyphae are shown in blue. C-E. Callose papillae in E. festucae ∆noxA mutant (C), in E. festucae ∆noxB mutant (D), in E. festucae ∆noxR mutant (E). L. perenne reacts with induction of callose production to all nox mutants in a similar way. Scale bar in A = 10 µm, in B = 100 µm, in C-E = 20 µm.
Notes
- When inoculating endophyte mutants, use mycelia grown on PD without antibiotics as antibiotics will kill the seedlings.
- Infection numbers will depend on your inoculation skill and plant genotype as well as Epichloë strain. We usually get 50-80% infected plants using E. festucae Fl1 and L. perenne Samson. We recently succeeded in inoculating E. festucae Fl1 in Festuca trachyphylla.
Recipes
- 3% water agar
Add 30 g agar to 1 L distilled water
Autoclave for 20 min at 121 °C and pour plates
Plates can be stored in the fridge for up to two weeks - 2.4% potato dextrose agar
Dissolve 24 g potato dextrose agar in 1 L distilled water
Autoclave for 20 min at 113 °C and pour plates
Plates can be stored in the fridge for up to two weeks - Murashige and Skoog phyto agar (for 1 L)
Make up to 1 L in distilled waterMurashige and Skoog Medium including vitamins 4.3 g Phyto agar 7.5 g
Autoclave for 20 min at 113 °C and pour in preservation jars approx. 4 to 5 cm in height. - Aniline Blue diammonium salt stock solution
Dissolve 0.01 g Aniline Blue diammonium salt in 1 ml Milli-Q water
Store at room temperature - WGA-AF488 stock solution
Dissolve 0.001 g WGA-AF488 in 1 ml Milli-Q water
Store at -20 °C in the dark - PBS (pH 7.4) (for 1 L)
Make up to 1 L in distilled waterNaCl 8.0 g KCl 0.2 g Na2HPO4 1.44 g KH2PO4 0.24 g
Adjust pH to 7.4 - Tween 20 stock
Make 1 ml stock by adding 20 µl Tween 20 to 980 µl Milli-Q water
Store at room temperature - Staining Solution (for 1 ml)
Make up to 1 ml in PBS (pH 7.4)Aniline Blue diammoium salt stock solution 20 µl WGA-AF488 stock solution 5 µl Propidium iodide solution 2 µl Tween 20 stock solution 10 µl
Make fresh on the day of use
Acknowledgments
The protocol as adapted from Latch and Christensen (1985) and Becker et al. (2016). We would like to thank Karolin Warnecke for excellent technical assistance. We gratefully acknowledge support by grants from the Bioprotection Research Centre and Marsden Fund of the Royal Society Te Apārangi.
Competing interests
The authors declare no conflict of interests.
References
- Becker, M., Becker, Y., Green, K. and Scott, B. (2016). The endophytic symbiont Epichloë festucae establishes an epiphyllous net on the surface of Lolium perenne leaves by development of an expressorium, an appressorium-like leaf exit structure. New Phytol 211(1): 240-254.
- Figueroa-Lopez, A. M., Cordero-Ramirez, J. D., Quiroz-Figueroa, F. R. and Maldonado-Mendoza, I. E. (2014). A high-throughput screening assay to identify bacterial antagonists against Fusarium verticillioides. J Basic Microbiol 54 Suppl 1: S125-133.
- Florea, S., Schardl, C. L. and Hollin, W. (2015). Detection and isolation of Epichloë species, fungal endophytes of grasses. Curr Protoc Microbiol 38: 19A 11 11-19A 11 24.
- Latch, G. C. M. and Christensen, M. J. (1985). Artificial infection of grasses with endophytes. Ann Appl Biol 107(1): 17-24.
- Scott, B., Becker, Y., Becker, M. and Cartwright, G. (2012). Morphogenesis, growth, and development of the grass symbiont Epichlöe festucae. In: Martín, J. P. and Pietro, A. D. (Eds.). Morphogenesis and Pathogenicity in Fungi. Springer, 243-264.
- Wood, P. J. and Fulcher, R. G. (1984). Specific interaction of Aniline Blue with (1→3)-β-D-Glucan. Carbohydrate Polymers 4: 49-72.
Article Information
Copyright
© 2018 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Becker, Y., Green, K. A., Scott, B. and Becker, M. (2018). Artificial Inoculation of Epichloë festucae into Lolium perenne, and Visualisation of Endophytic and Epiphyllous Fungal Growth. Bio-protocol 8(17): e2990. DOI: 10.21769/BioProtoc.2990.
Category
Microbiology > Microbe-host interactions > Fungus
Plant Science > Plant cell biology > Cell staining
Cell Biology > Cell staining > Chitin
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link











