- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Structural Analysis of Bordetella pertussis Biofilms by Confocal Laser Scanning Microscopy
Published: Vol 8, Iss 15, Aug 5, 2018 DOI: 10.21769/BioProtoc.2953 Views: 6895
Reviewed by: Emily CopeJuan Facundo Rodriguez AyalaSofiane El-Kirat-Chatel

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
An Inexpensive Imaging Platform to Record and Quantitate Bacterial Swarming
Weijie Chen [...] Jay X. Tang
Sep 20, 2021 3669 Views
Purification of the Bacterial Amyloid “Curli” from Salmonella enterica Serovar Typhimurium and Detection of Curli from Infected Host Tissues
Murugesan Sivaranjani [...] Aaron P. White
May 20, 2022 3252 Views
A Guideline for Assessment and Characterization of Bacterial Biofilm Formation in the Presence of Inhibitory Compounds
Bassam A. Elgamoudi and Victoria Korolik
Nov 5, 2023 3199 Views
Abstract
Biofilms are sessile communities of microbial cells embedded in a self-produced or host-derived exopolymeric matrix. Biofilms can both be beneficial or detrimental depending on the surface. Compared to their planktonic counterparts, biofilm cells display enhanced resistance to killing by environmental threats, chemicals, antimicrobials and host immune defenses. When in biofilms, the microbial cells interact with each other and with the surface to develop architecturally complex multi-dimensional structures. Numerous imaging techniques and tools are currently available for architectural analyses of biofilm communities. This allows examination of biofilm development through acquisition of three-dimensional images that can render structural features of the sessile community. A frequently utilized tool is Confocal Laser Scanning Microscopy. We present a detailed protocol to grow, observe and analyze biofilms of the respiratory human pathogen, Bordetella pertussis in space and time.
Keywords: BiofilmBackground
Bordetella pertussis is an obligate human pathogen of the upper respiratory tract that causes whooping cough or pertussis (Mooi, 2010; Dorji et al., 2018). Biofilms of B. pertussis form on a variety of artificial surfaces and under static, shaking, and fluid flow conditions (Mishra et al., 2005; Sloan et al., 2007; Serra et al., 2011). Microscopic evaluation of these biofilms shows that this bacterium produces irregularly shaped microcolonies separated by fluid channels, embedded in an exopolymeric matrix composed by extracellular DNA (eDNA), proteins and polysaccharides (Parise et al., 2007; Sloan et al., 2007; Serra et al., 2008; Conover et al., 2011; Nicholson et al., 2012; Ganguly et al., 2014; Cattelan et al., 2017). In addition to forming biofilms in the laboratory setting, B. pertussis forms multi-dimensional organ-adherent biofilms on the nose and trachea during experimental infections of mice. Development of these mammalian biofilms is characterized by an extracellular polymeric matrix composed of eDNA, the Filamentous hemagglutinin protein and the Bps polysaccharide (Conover et al., 2010 and 2011; Serra et al., 2011; Dorji et al., 2018). Based on these results, biofilm formation has been proposed by us and others as a possible strategy adopted by B. pertussis to infect, persist and continually circulate in the community (Cattelan et al., 2016). Consistent with this hypothesis, we found that currently circulating strains from Argentina and USA produce significantly higher levels of biofilms when compared to a laboratory reference strain and colonize the mouse nose and trachea at higher numbers than the prototype laboratory strain (Cattelan et al., 2017). These results also provide evidence that hyperbiofilm growth is a strategy employed by circulating organisms to infect and survive inside their host.
In order to study the mechanisms involved in biofilm development, microscopic evaluation is a key technique that allows differentiation of the steps of the process, from adhesion to maturation and dispersion. In particular, confocal laser scanning microscopy (CLSM) is frequently used because it allows visualization of architectural complexities of intact and hydrated biofilms. In this protocol, we describe how to grow and process samples of B. pertussis biofilms, as demonstrated in our recent publication (Cattelan et al., 2017).
Materials and Reagents
- 0.2 μm filter
- Bacteriological Petri plates (Fisher Scientific, catalog number: FB0875712 )
- Sterile test tubes (17 x 100 mm, RPI, Research Products International, catalog number: 168599 )
- Serological pipettes:
5 ml pipette (SARSTEDT, catalog number: 86.1253.001 )
25 ml pipette (SARSTEDT, catalog number: 86.1685.001 ) - NalgeneTM 2 ml cryogenic tubes (Thermo Fisher Scientific, Thermo ScientificTM, catalog number 5000-0020 )
- Coverglasses, 22 x 22 mm (Fisher Scientific, FisherbrandTM, catalog number: 12-542B )
- NuncTM 6-wells plates (Thermo Fisher Scientific, Thermo ScientificTM, catalog number: 150239 )
- Aluminum foil
- Microscope slides (Fisher Scientific, FisherbrandTM, catalog number: 12-552-3 )
- Sterile disposable plastic material:
Posi-Click tubes (Denville Scientific, catalog number: C2170 )
P200 and P1000 tips (USA Scientific, catalog numbers: 1111-0706 and 1112-1720 ) - B. pertussis strain harboring a pGB5P1-GFP plasmid
- Defibrinated sheep’s blood (Hemostat Laboratories, catalog number: DSB100 )
- 50% glycerol solution (Fisher Scientific, catalog number: G33-1 )
- Kanamycin (Sigma-Aldrich, catalog number: K1377 )
- Neutralized buffered formalin (Fisher Scientific, catalog number: SF100-4 )
- Stainer-Scholte medium (Stainer et al., 1970; Nicholson et al., 2012)
- Sterile PBS (Thermo Fisher Scientific, GibcoTM, catalog number: 14190144 )
- ProLongTM Gold Antifade Mountant (Thermo Fisher Scientific, InvitrogenTM, catalog number: P36934 )
- DifcoTM Bordet-Gengou agar plates (BD, BD Biosciences, catalog number: 248200 )
- L-Proline (Sigma-Aldrich, catalog number: 131547 )
- KH2PO4 (Sigma-Aldrich, catalog number: V000225 )
- KCl (Sigma-Aldrich, catalog number: P3911 )
- MgCl2·6H2O (Sigma-Aldrich, catalog number: M2393 )
- CaCl2 (Sigma-Aldrich, catalog number: C1016 )
- Tris base (Sigma-Aldrich, catalog number: T1378 )
- NaCl (Sigma-Aldrich, catalog number: S7653 )
- FeSO4·7H2O (Sigma-Aldrich, catalog number: F8633 )
- L-cystine (Sigma-Aldrich, catalog number: C8755 )
- L-ascorbic acid (Sigma-Aldrich, catalog number: A5960 )
- Nicotinic acid (Sigma-Aldrich, catalog number: PHR1276 )
- Reduced L-glutathione (Sigma-Aldrich, catalog number: G4251 )
- SS medium (see Recipes)
- SS supplement (filter-sterilized) (see Recipes)
- Bordet-Gengou (BG) medium (see Recipes)
Equipment
- Kimax® Erlenmeyer flasks (125 ml) (DWK Life Sciences, KIMBLE®, catalog number: 26500 )
- Sterile forceps
- PIPETMAN® Classic (Gilson, models: P20, P200, P1000, catalog numbers: F123600 , F123601 , F123602 )
- Humid chamber (an appropriately sized Tupperware container with either a weigh boat containing water or paper towels soaked with water)
- Chemical fume hood (Hamilton Laboratory Solutions, model: PL-822 )
- Eclipse Ti-E inverted Confocal Microscope (Nikon, model: D-Eclipse C1si )
- Vortex (Fisher Scientific, catalog number: 02-215-365 )
- pH meter (Denver Instrument, model: UB-10 )
- Incubator (Thermo Fisher Scientific, Thermo ScientificTM, model: FormaTM Steri-CycleTM )
- Benchtop centrifuge (Eppendorf, model: 5418 )
- Refrigerator (4 °C) (Whirlpool, model: WRR56X18FW02 )
- -20 °C freezer (Kenmore, model: 253-26722103 )
- SPECTRONIC TM Spectrophotometer (Thermo Fisher Scientific, model: GENESYS 20 )
- Weigh Balance (Ohaus, model: Scout Pro SP202 and VWR, model: VWR-164AC )
- Roller Drum (Eppendorf, New BrunswickTM, model: TC-2 )
Software
- ImageJ (Schneider et al., 2012)
- COMSTAT2 plugin (Heydorn et al., 2000)
Procedure
Notes:
- The entire protocol is depicted in Figure 1.
- Samples should be protected from light during the experiment. In all cases, medium is supplemented with appropriate antibiotic to maintain the plasmid coding for GFP.
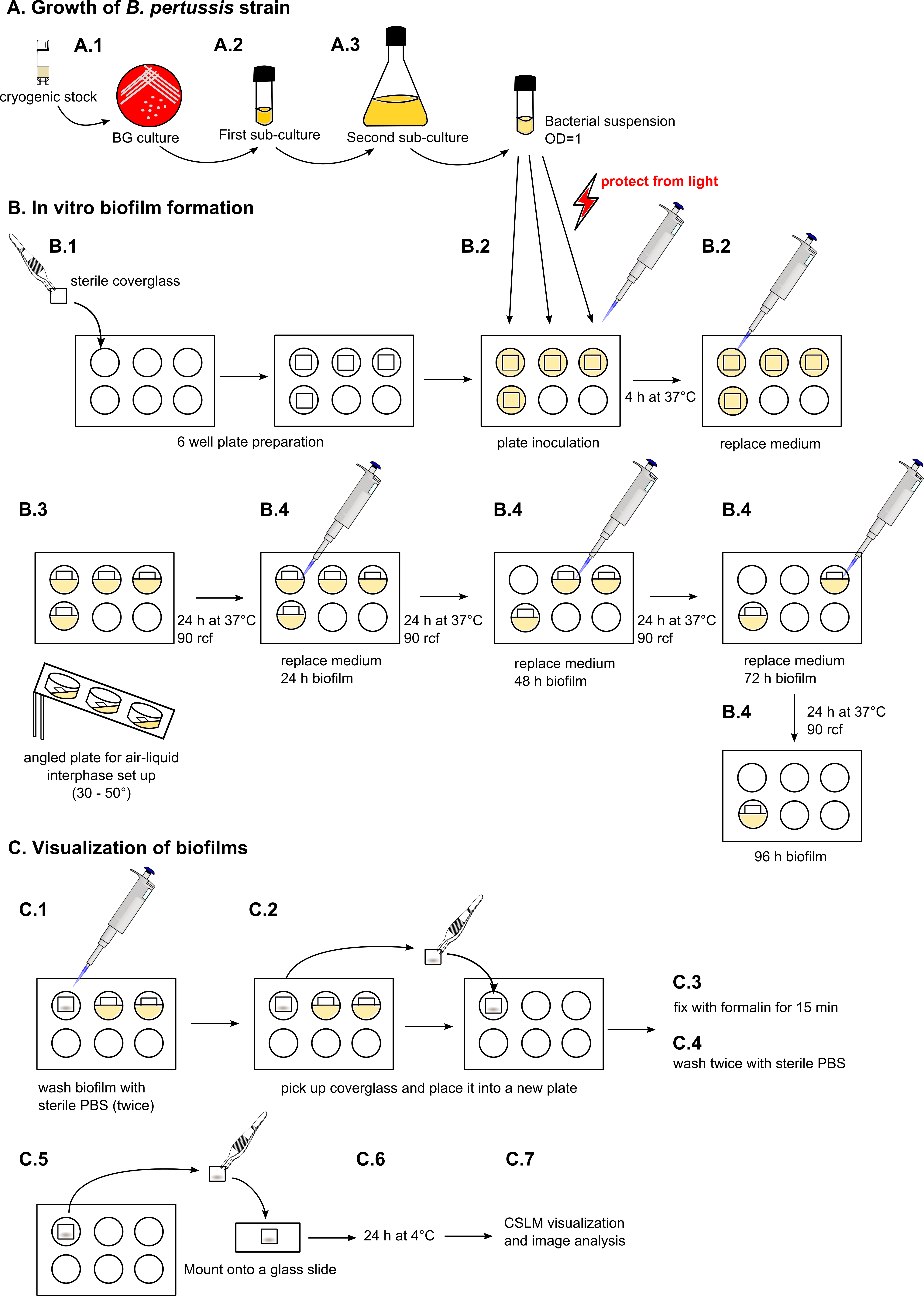
Figure 1. Schematic step-by-step protocol for growing B. pertussis biofilms for CLSM evaluation
- Growth of B. pertussis strains
- From frozen stock stored at -78 °C, streak B. pertussis strains harboring a GFP-coding plasmid, pGB5P1-GFP was used in this protocol (Weingart et al., 1999), on Bordet-Gengou agar plates supplemented with 10% defibrinated sheep’s blood and 25 μg/ml of kanamycin. Incubate the plates at 37 °C for 4-5 days.
- Pick individual colonies (~8-10) and inoculate 2 ml of Stainer-Scholte (SS) liquid medium supplemented with 25 μg/ml of kanamycin. Incubate at 37°C in a roller drum spinning at top speed for 24 h.
- Inoculate 30 ml of SS medium supplemented with 25 μg/ml of kanamycin (in a 100 ml Erlenmeyer flask), with the cultures from Step A2 to achieve an OD650 of 0.2. Incubate for 12 h at 37 °C with shaking at 160 rcf.
- From frozen stock stored at -78 °C, streak B. pertussis strains harboring a GFP-coding plasmid, pGB5P1-GFP was used in this protocol (Weingart et al., 1999), on Bordet-Gengou agar plates supplemented with 10% defibrinated sheep’s blood and 25 μg/ml of kanamycin. Incubate the plates at 37 °C for 4-5 days.
- In vitro biofilm formation
- Insert a sterile coverglass (flame sterilized) at the bottom of sterile 6-well plates.
- Prepare inoculum by diluting B. pertussis culture (Procedure A, Step 3) to an OD650 of 1.0 in SS medium. Pipet 1.5 ml of inoculum into individual wells of the 6-well plates. Make sure the bacterial suspension is covering the entire surface of the coverslip. In case other substrates are used (i.e., polycarbonate or aluminum coupons), volume should be adjusted according to the thickness of the material.
- Cover the plate and incubate statically in a humidified chamber for 4 h at 37 °C.
Note: This step allows the adhesion of bacteria to the surface. - Remove inocula (to allow removal of unattached planktonic bacteria) carefully without disturbing attached cells and add approximately 800 μl of fresh SS medium, dispensing along the wall of the well to prevent biofilm disruption. Place the plate at an angle of 30°-50° relative to horizontal, forming an air-liquid interface in the coverglass where biofilms will develop. Cover the plate with aluminum foil and incubate in a humidified chamber at 37 °C on a platform shaker at 90 rcf.
- If the biofilm formation is going to be examined for longer than 24 h, change medium every 24 h as described above. Samples can be processed at 24, 48, and 72 up to 96 h.
- Insert a sterile coverglass (flame sterilized) at the bottom of sterile 6-well plates.
- Visualization of biofilms
- After each time point, aspirate medium from each well and carefully wash the biofilms twice with sterile PBS.
- Pick the coverglass up with forceps and place it into a new 6-well plate. A needle can be used to aid in lifting the coverglass followed by use of forceps to pick the coverglass without touching the area where the biofilm developed.
- Carefully add 50 μl of neutral buffered formalin, or enough to cover the glass surface. Special attention must be taken in order to not disturb the biofilm and incubate for 15 min at room temperature.
- Remove fixative and wash twice with PBS.
- Pick the coverglass up with forceps and mount onto a glass slide with ProLongTM Gold mounting media. Alternatively, if bacteria are not labeled with a fluorescent protein, cells can be stained at this point with a fluorescent dye and then mounted.
- Allow the mounting media to harden by incubating at 4 °C for 24 h.
- Slides can be stored (weeks to months) in the dark at room temperature for later visualization. Examine the samples with a confocal microscope, taking images in a z-stack configuration, making sure to cover the entire biofilm structure from bottom to top.
- Analyze the images by using microscopy image visualization software (Figure 2).

Figure 2. Micrographs of a B. pertussis biofilm. Biofilms were grown from 24 up to 96 h and images were acquired in an inverted confocal microscope in a z-stack setting. Images are presented in xy, xz and yz planes.
- After each time point, aspirate medium from each well and carefully wash the biofilms twice with sterile PBS.
Data analysis
Quantitative data from images can be obtained with COMSTAT2 plug-in (Heydorn et al., 2000) run in ImageJ (Schneider et al., 2012). Structural features that can be analyzed are maximum and average thicknesses, substrate coverage, biomass and roughness coefficient (see structural data in Table 1). Detailed instructions for COMSTAT2 use are described in Heydorn et al. (2000). Appropriate statistical analysis should be carried out on the obtained data.
Table 1. Biofilm structural features of the images presented in Figure 2, obtained with COMSTAT2 software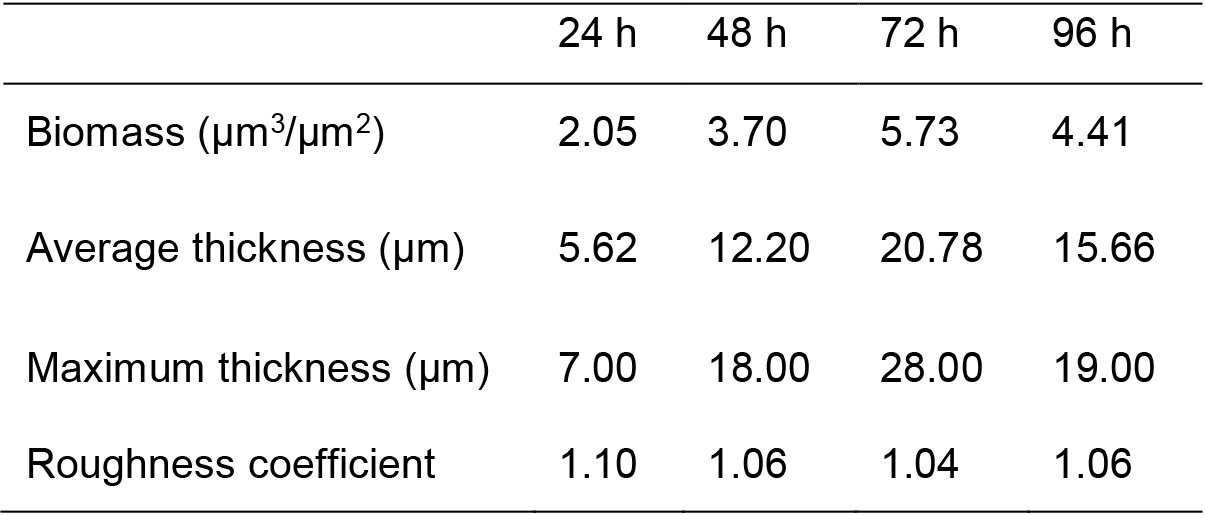
Notes
- Due to variations observed with this assay, there is need to be performed with duplicates of each sample and at least three independent replicates. For each sample, at least three representative images should be acquired.
- A critical consideration is not to disrupt the biofilm attached to the glass surface. Depending on the microorganism, cells may attach either strongly or loosely to surfaces. Thus it is important to be extremely careful while dispensing liquid to avoid strong shear forces.
- BG-agar medium and SS medium are sterilized by autoclaving at 121 °C for 15 min.
- To prepare a cryogenic stock, cultures grown to exponential phase (set up as explained in Step A2) should be used. Use sterile 2 ml cryogenic tubes and mix 500 μl of bacterial culture with 500 μl of sterile 50% glycerol solution. Store at -78 °C.
Recipes
Note: Unless otherwise indicated, all stock solutions are prepared using Milli Q water.
- SS medium
Monosodium glutamate: 10.70 g/L
L-Proline: 0.24 g/L
KH2PO4: 0.5 g/L
KCl: 0.20 g/L
MgCl2·6H2O: 0.20 g/L
CaCl2: 0.2 g/L
Tris base: 1.52 g/L
NaCl: 2.50 g/L
Adjust to pH 7.3-7.4 with HCl - SS supplement (0.2 μm filter-sterilized)
Dissolve in HCl concentrated, and then filter sterilize
FeSO4·7H2O: 10 mg/L
L-cystine: 40 mg/L
L-ascorbic acid: 20 mg/L
Nicotinic acid: 4 mg/L
Reduced L-glutathione: 100 mg/L - Bordet-Gengou (BG) medium
DifcoTM Bordet-Gengou agar (30 g/L)
20 ml of 50% glycerol
Autoclave at 121 °C for 15 min
Aseptically add 15% sterile, defibrinated blood to the medium at 45-50 °C
Mix well and plate
Acknowledgments
This project has been funded in part with Federal funds from the National Institute of Allergy and Infectious Diseases, National Institutes of Health, Department of Health and Human Services by grants 1R21AI123805-0 and R01AI125560.
This protocol was adapted from Cattelan, N., Jennings-Gee, J., Dubey, P., Yantorno, O. M. and Deora, R. (2017). Hyperbiofilm Formation by Bordetella pertussis Strains Correlates with Enhanced Virulence Traits. Infect Immun 85(12).
The authors declare no conflict of interest.
References
- Cattelan, N., Dubey, P., Arnal, L., Yantorno, O. M. and Deora, R. (2016). Bordetella biofilms: a lifestyle leading to persistent infections. Pathog Dis 74(1): ftv108.
- Cattelan, N., Jennings-Gee, J., Dubey, P., Yantorno, O. M. and Deora, R. (2017). Hyperbiofilm formation by Bordetella pertussis strains correlates with enhanced virulence traits. Infect Immun 85(12).
- Conover, M. S., Mishra, M. and Deora, R. (2011). Extracellular DNA is essential for maintaining Bordetella biofilm integrity on abiotic surfaces and in the upper respiratory tract of mice. PLoS One 6(2): e16861.
- Conover, M. S., Sloan, G. P., Love, C. F., Sukumar, N. and Deora, R. (2010). The Bps polysaccharide of Bordetella pertussis promotes colonization and biofilm formation in the nose by functioning as an adhesin. Mol Microbiol 77(6): 1439-1455.
- Dorji, D., Mooi, F., Yantorno, O., Deora, R., Graham, R. M. and Mukkur, T. K. (2018). Bordetella pertussis virulence factors in the continuing evolution of whooping cough vaccines for improved performance. Med Microbiol Immunol 207(1): 3-26.
- Ganguly, T., Johnson, J. B., Kock, N. D., Parks, G. D. and Deora, R. (2014). The Bordetella pertussis Bps polysaccharide enhances lung colonization by conferring protection from complement-mediated killing. Cell Microbiol 16(7): 1105-1118.
- Heydorn, A., Nielsen, A. T., Hentzer, M., Sternberg, C., Givskov, M., Ersboll, B. K. and Molin, S. (2000). Quantification of biofilm structures by the novel computer program COMSTAT. Microbiology 146 (Pt 10): 2395-2407.
- Mishra, M., Parise, G., Jackson, K. D., Wozniak, D. J. and Deora, R. (2005). The BvgAS signal transduction system regulates biofilm development in Bordetella. J Bacteriol 187(4): 1474-1484.
- Mooi, F. R. (2010). Bordetella pertussis and vaccination: the persistence of a genetically monomorphic pathogen. Infect Genet Evol 10(1): 36-49.
- Nicholson, T. L., Conover, M. S. and Deora, R. (2012). Transcriptome profiling reveals stage-specific production and requirement of flagella during biofilm development in Bordetella bronchiseptica. PLoS One 7(11): e49166.
- Parise, G., Mishra, M., Itoh, Y., Romeo, T. and Deora, R. (2007). Role of a putative polysaccharide locus in Bordetella biofilm development. J Bacteriol 189(3): 750-760.
- Schneider, C. A., Rasband, W. S. and Eliceiri, K. W. (2012). NIH Image to ImageJ: 25 years of image analysis. Nat Methods 9(7): 671-675.
- Serra, D. O., Conover, M. S., Arnal, L., Sloan, G. P., Rodriguez, M. E., Yantorno, O. M. and Deora, R. (2011). FHA-mediated cell-substrate and cell-cell adhesions are critical for Bordetella pertussis biofilm formation on abiotic surfaces and in the mouse nose and the trachea. PLoS One 6(12): e28811.
- Serra, D. O., Lucking, G., Weiland, F., Schulz, S., Gorg, A., Yantorno, O. M. and Ehling-Schulz, M. (2008). Proteome approaches combined with Fourier transform infrared spectroscopy revealed a distinctive biofilm physiology in Bordetella pertussis. Proteomics 8(23-24): 4995-5010.
- Sloan, G. P., Love, C. F., Sukumar, N., Mishra, M. and Deora, R. (2007). The Bordetella Bps polysaccharide is critical for biofilm development in the mouse respiratory tract. J Bacteriol 189(22): 8270-8276.
- Stainer, D. W. and Scholte, M. J. (1970). A simple chemically defined medium for the production of phase I Bordetella pertussis. J Gen Microbiol 63(2): 211-220.
- Sukumar, N., Love, C. F., Conover, M. S., Kock, N. D., Dubey, P. and Deora, R. (2009). Active and passive immunizations with Bordetella colonization factor A protect mice against respiratory challenge with Bordetella bronchiseptica. Infect Immun 77(2): 885-895.
- Weingart, C. L., Broitman-Maduro, G., Dean, G., Newman, S., Peppler, M. and Weiss, A. A. (1999). Fluorescent labels influence phagocytosis of Bordetella pertussis by human neutrophils. Infect Immun 67(8): 4264-4267.
Article Information
Copyright
© 2018 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Cattelan, N., Yantorno, O. M. and Deora, R. (2018). Structural Analysis of Bordetella pertussis Biofilms by Confocal Laser Scanning Microscopy. Bio-protocol 8(15): e2953. DOI: 10.21769/BioProtoc.2953.
Category
Microbiology > Microbial biofilm > Biofilm culture
Cell Biology > Cell imaging > Confocal microscopy
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










