- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
A Procedure for Precise Determination of Glutathione Produced by Saccharomyces cerevisiae
Published: Vol 8, Iss 12, Jun 20, 2018 DOI: 10.21769/BioProtoc.2887 Views: 7893
Reviewed by: Valentine V TrotterPushpendra SinghAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
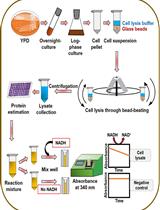
An Assay to Determine NAD(P)H: Quinone Oxidoreductase Activity in Cell Extracts from Candida glabrata
Anamika Battu [...] Rupinder Kaur
Nov 5, 2021 3089 Views
Isolation of Mitochondria from Ustilago maydis Protoplasts
Juan Pablo Pardo [...] Lucero Romero-Aguilar
Jan 5, 2022 3605 Views
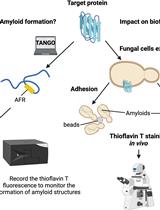
Use of the Fluorescent Dye Thioflavin T to Track Amyloid Structures in the Pathogenic Yeast Candida albicans
Thierry Mourer [...] Sophie Bachellier-Bassi
Feb 5, 2024 2375 Views
Abstract
In bioproduction, yields of products must be calculated precisely for accurate evaluation of various fermentation conditions. To evaluate productivity of microorganisms, product amounts per unit of medium volume (e.g., mg-product/L-broth), and/or product amounts per unit of a microorganism amount (e.g., mg-product/mg-dry cell weight) are often used. Nonetheless, detailed procedures for calculation of these production yields are often omitted in research articles, whereas methods for product quantification are described well. Here, we describe a detailed calculation procedure from our previous studies on glutathione production by Saccharomyces cerevisiae. This procedure can be applied to various other products and microorganisms, and therefore, may prove to be useful in various other bioproduction studies.
Keywords: Saccharomyces cerevisiaeBackground
Glutathione is the most abundant thiol-containing tripeptide in all living organisms and functions as a bioactive substance with varied roles in cells, e.g., as redox and antidotal agents. Therefore, glutathione is widely used in the medical, food, and cosmetic industries nowadays, and the demand has increased in recent years. Glutathione is industrially produced mainly by fermentation using Saccharomyces cerevisiae, which originally contains a high concentration of glutathione and has served as a safe, food-producing microorganism. Studies of microbial glutathione production in various microorganisms will become more important in the future. To evaluate the productivity in terms of glutathione via fermentation by various microorganisms, here we describe our detailed procedures of sample preparation, quantification of reduced and oxidized glutathione by high performance chromatography (HPLC), and calculations of two types of yield (Hara et al., 2012; Hara et al., 2015; Kiriyama et al., 2013; Kobayashi et al., 2017).
Materials and Reagents
- 200 μl pipette tips (FCR&Bio, catalog number: AG-200-FP-Y )
- 1,000 μl pipette tips (FCR&Bio, catalog number: AG-1000B )
- 1.5 ml microcentrifuge tubes (FUKAEKASEI and WATSON, catalog number: 131-815C )
- 50 ml centrifuge tubes (Corning, catalog number: 430291 )
- Filter unit with 0.22 μm pore size (Shimadzu, catalog number: GLCTD-MCE1322 )
- Saccharomyces cerevisiae
- Yeast cell culture medium
- Ultrapure water (Milli-Q)
- Ice
- Ethylenediaminetetraacetic acid (EDTA) disodium salt dihydrate (NACALAI TESQUE, catalog number: 15111-45 )
- Sodium hydroxide (NaOH) (NACALAI TESQUE, catalog number: 31511-05 )
- Potassium dihydrogen phosphate (KH2PO4) (Wako Pure Chemical Industries, catalog number: 163-04265 )
- 1-Heptanesulfonate (NACALAI TESQUE, catalog number: 31528-92 )
- Phosphoric acid (Wako Pure Chemical Industries, catalog number: 162-20492 )
- Methanol (NACALAI TESQUE, catalog number: 21929-23 )
- Yeast extract dried (NACALAI TESQUE, catalog number: 15838-45 )
- Hipolypepton (NIHON PHARMACEUTICAL, catalog number: 392-02115 )
- D-(+)-Glucose (NACALAI TESQUE, catalog number: 16805-35 )
- Aureobasidin A (Takara Bio, catalog number: 630499 )
- 99.5% Ethanol (NACALAI TESQUE, catalog number: 14713-95 )
- Agar powder (NACALAI TESQUE, catalog number: 01028-85 )
- Standard Buffer Solution (pH 4.01) (NACALAI TESQUE, catalog number: 37219-75 )
- Standard Buffer Solution (pH 6.86) (NACALAI TESQUE, catalog number: 37220-35 )
- Standard Buffer Solution (pH 9.18) (NACALAI TESQUE, catalog number: 37221-25 )
- YPD medium (see Recipes)
- Aureobasidin A stock solution (see Recipes)
- 500 mM EDTA (see Recipes)
- 100 mM EDTA (see Recipes)
- HPLC mobile phase (see Recipes)
Equipment
- Micropipetter (volume range 20-200 μl) (Nichiryo, model: NPX-200 )
- Micropipetter (volume range 100-1,000 μl) (Nichiryo, model: NPX-1000 )
- Block heater (ASTEC, model: BI-516C )
- Aluminum microtube rack (Bio Medical Science, model: BMS-D081 )
- Centrifuge (KUBOTA, model: 3740 )
- Centrifuge rotor for 1.5 ml microcentrifuge tubes (maximum capacity is 21,880 x g) (KUBOTA, model: AF-2236 )
- Centrifuge rotor for 50 ml centrifuge tubes (maximum capacity is 22,140 x g) (KUBOTA, model: AF-5004CH )
- Glass test tube (NICHIDEN RIKA GLASS, model: 101015 )
- Baffled Erlenmeyer flask (AGC TECHNO GLASS, model: 4551FK200R )
- Silicone sponge tapered plug closure (Shin-Etsu Polymer, model: T-19 )
- Silicone sponge closure (Shin-Etsu Polymer, model: C-40 )
- Spectrophotometer (Shimadzu, model: UVmini-1240 )
- Spectrophotometer cell (Hellma, model: 104-10-40 )
- High-performance liquid chromatography system (Shimadzu, model: Prominence)
- Octa decyl silyl (ODS) column (YMC, model: YMC-Pack ODS-A )
- Electronic scale (Mettler-Toledo International, model: XS105DU )
- Vortex mixer (Scientific Industries, model: Vortex-Genie 2 )
- Ice-making machine (HOSHIZAKI, model: CM-100K )
- Deep freezer (temperature range: from -20 to -30 °C) (Panasonic, model: MDF-U539 )
- Deep freezer (temperature range: from -50 to -85 °C) (PHC, model: MDF-U33V-PJ )
- pH meter (Horiba, model: F-52 )
- Forced air flow oven (Tokyo Rikakikai, model: WFO-1210 )
- Disposable loop (AS ONE, model: 6-488-02 )
- Incubator for solid preculture (Sanyo, model: MIR-153 )
- Incubator for liquid preculture (TAITEC, model: BR-43FL・MR )
- Incubator for glutathione production (TAITEC, model: G・BR-200 )
Procedure
- Glutathione production
- An S. cerevisiae strain was streaked with a disposable loop on the YPD solid medium and grown at 30 °C for 72 h.
- A single grown colony was inoculated into 5 ml of the YPD liquid medium in a 25 ml glass test tube closed with a tapered plug closure made of silicone sponge. The bacteria were aerobically grown with agitation at 200 rpm for 18 h at 30 °C.
- Transfer 50 μl of the grown yeast culture into a 1.5 ml microtube. Add 750 μl of ultrapure water and 200 μl of 500 mM EDTA to the 1.5 ml microtube and mix thoroughly by vortexing.
- Measure OD600 of this diluted yeast suspension in the EDTA solution.
- To determine OD600 of the original yeast preculture (ODPC), we multiplied the measured OD600 by 20 (dilution factor).
- The grown cells were inoculated into 20 ml of the YPD liquid medium in a 200 ml baffled Erlenmeyer flask plugged with a silicone sponge closure and were grown at 30 °C with agitation at 150 rpm for up to 48 h (Figure 1). The yeast cell culture volume (ml) for inoculation was calculated using the equation below. In the following equation, 20 and 0.03 indicate the total volume (μl) of the YPD liquid medium used for glutathione fermentation and the initial cell density (OD600), respectively.
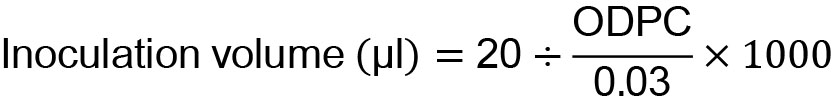
- The yeast culture (1 ml) grown in a baffled Erlenmeyer flask was sampled at 24 and 48 h.
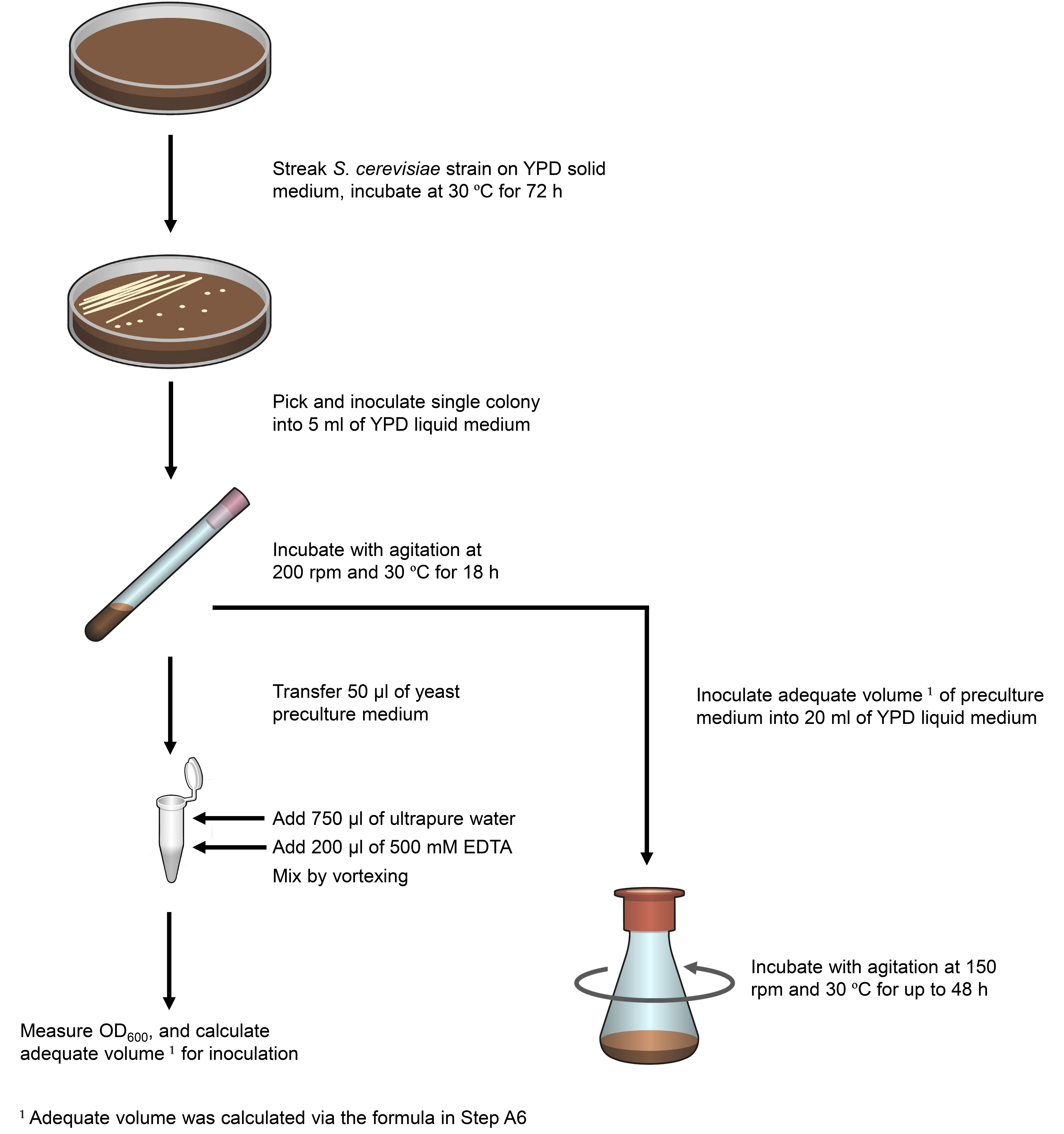
Figure 1. Schematic outline of glutathione production. A more detailed procedure is described in Procedure A.
- An S. cerevisiae strain was streaked with a disposable loop on the YPD solid medium and grown at 30 °C for 72 h.
- Determination of biomass concentration
- Cultivate yeast under the conditions described in Procedure A.
- Transfer 50 μl of the yeast culture (grown for glutathione production) into a 1.5 ml microtube. Add 200 μl of 500 mM EDTA and 750 μl of ultrapure water up to the 1.5 ml label in the microtube and mix thoroughly by vortexing.
- Measure OD600 of this diluted yeast suspension in an EDTA solution.
- To determine OD600 of the original yeast culture, multiply the measured OD600 by 20 (dilution factor).
- Centrifuge (16,000 x g, 10 min) all the yeast culture medium in 50 ml centrifuge tubes.
- Discard the supernatant and add 20 ml of room temperature ultrapure water to the cell pellet and thoroughly resuspend it by vortexing.
- Repeat this wash procedure one more time and re-pellet the cells.
- Add an adequate volume of 100 mM EDTA to the cell pellet to achieve optical density [at 600 nm (OD600) of the cell suspension] of approximately 1.0.
- Dilute the cell suspension with room temperature 100 mM EDTA to prepare 2-, 4-, 8-, and 16-fold diluted cell suspensions, and measure OD600 of these serial dilutions.
- Weigh empty open 50 ml centrifuge tubes and centrifuge (16,000 x g, 10 min) 20 ml of these cell suspensions in these tubes.
- Remove the supernatant and incubate the open centrifuge tube at 80 °C in a forced air flow oven overnight to dry the cells sufficiently.
- Weigh each 50 ml centrifuge tube with the dry cell pellet and calculate the weight of the dry cell pellet by subtracting the weight of the empty tube from the total weight (Figure 2).
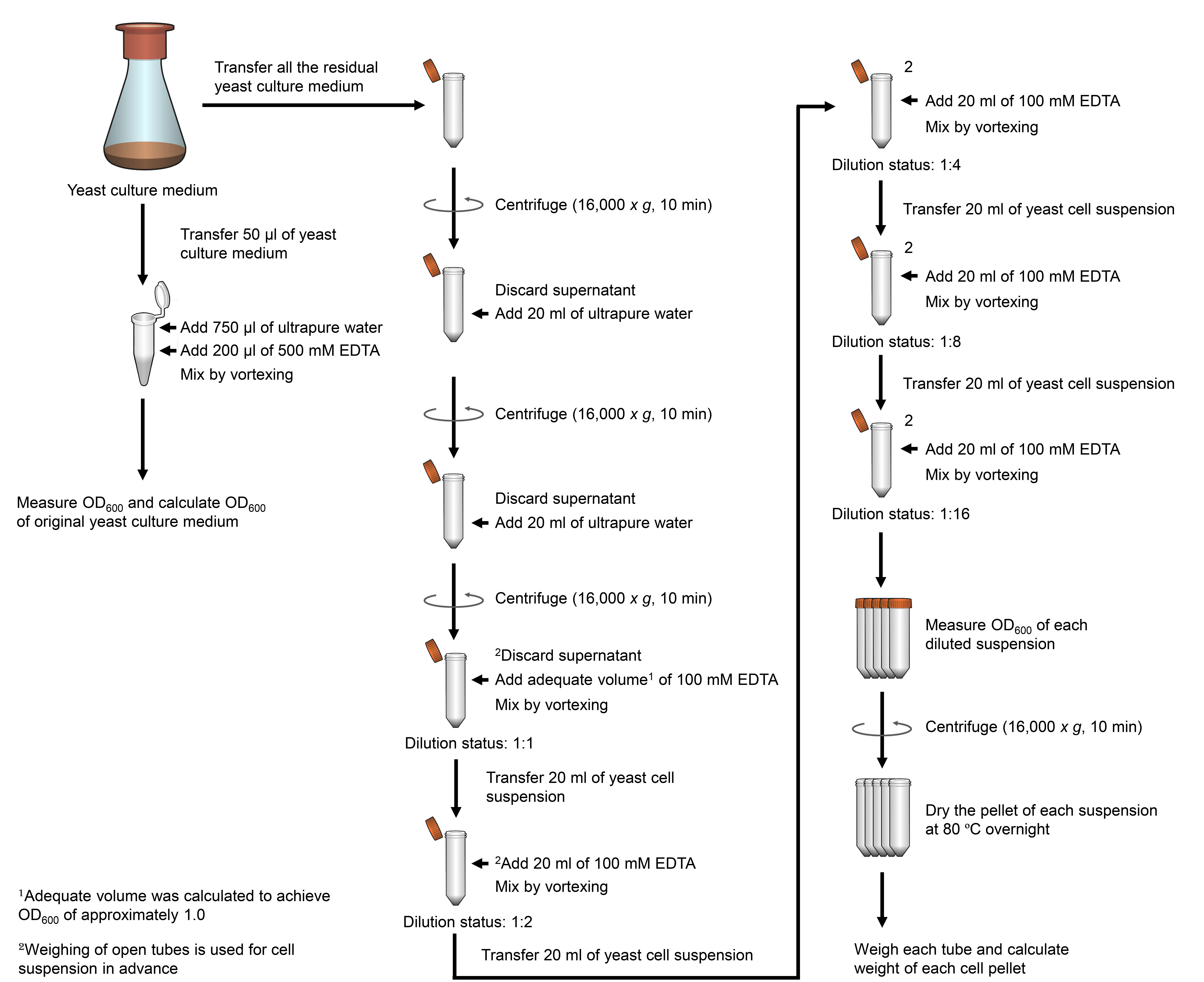
Figure 2. Schematic outline of determination of biomass concentration. More detailed procedure is described in Procedure B. - Construct a calibration curve for OD600 and biomass concentration ([g-dry cell weight]/[L-broth]). An example of a calibration curve is illustrated in Figure 3.
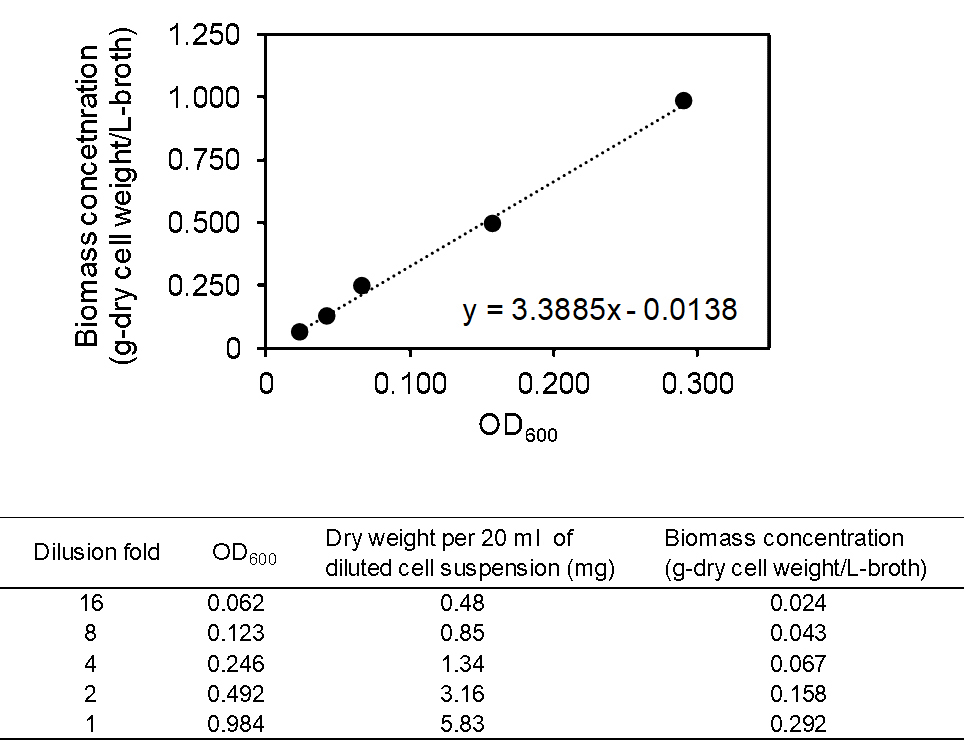
Figure 3. Example of calibration curve for OD600 and biomass concentration. Cells of S. cerevisiae GCI strain grown in YPD medium at 30 °C for 48 h were used for the construction of this calibration curve.
- Cultivate yeast under the conditions described in Procedure A.
- Measurement of cell concentration and cell disruption
- Transfer 1 ml of the yeast culture grown for glutathione production in a 1.5 ml microtube. Add 250 μl of 500 mM EDTA to the 1.5 ml microtube and mix thoroughly by vortexing.
- Transfer 50 μl of this yeast culture containing 100 mM EDTA into a fresh 1.5 ml microtube. Add 950 μl of ultrapure water into the 1.5 ml microtube and mix thoroughly by vortexing.
- Measure OD600 of this diluted yeast suspension in the EDTA solution.
- To determine OD600 of the original yeast culture (ODC), multiply the measured OD600 by 25 (dilution factor).
- Centrifuge (16,000 x g, 1 min) 1.2 ml of the residual yeast culture in a 100 mM EDTA solution and resuspend the pellet in 1 ml of ultrapure water by vortexing.
- Repeat this wash procedure one more time and re-pellet the cells.
- Add an adequate volume (μl) of ultrapure water calculated by means of the equation below for the cell pellet and resuspend it well. In the following equation, 960 and 35 mean residual yeast culture volume (μl) without the EDTA solution, and OD600 that is optimal for glutathione determination, respectively.
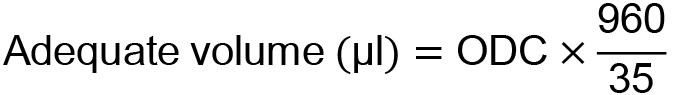
- Transfer 980 μl of distilled water and 20 μl of the concentrated yeast cell suspension to a different 1.5 ml microtube and mix thoroughly.
- Measure OD600 of this diluted yeast cell suspension and multiply the measured OD600 by 50 (dilution factor) to obtain OD600 of the concentrated resuspended cells (ODR).
- To disrupt yeast cells, incubate the concentrated cell suspension at 95 °C for 3 min and then cool on ice for 3 min.
- Centrifuge (16,000 x g, 10 min) the disrupted cell suspension, and transfer 50 μl of the supernatant (solution containing cell content) to a new 1.5 ml microtube. The cell content solution can be stored at -20 °C before HPLC analysis (Figure 4). At least three samples per strain are needed for accurate measurements.
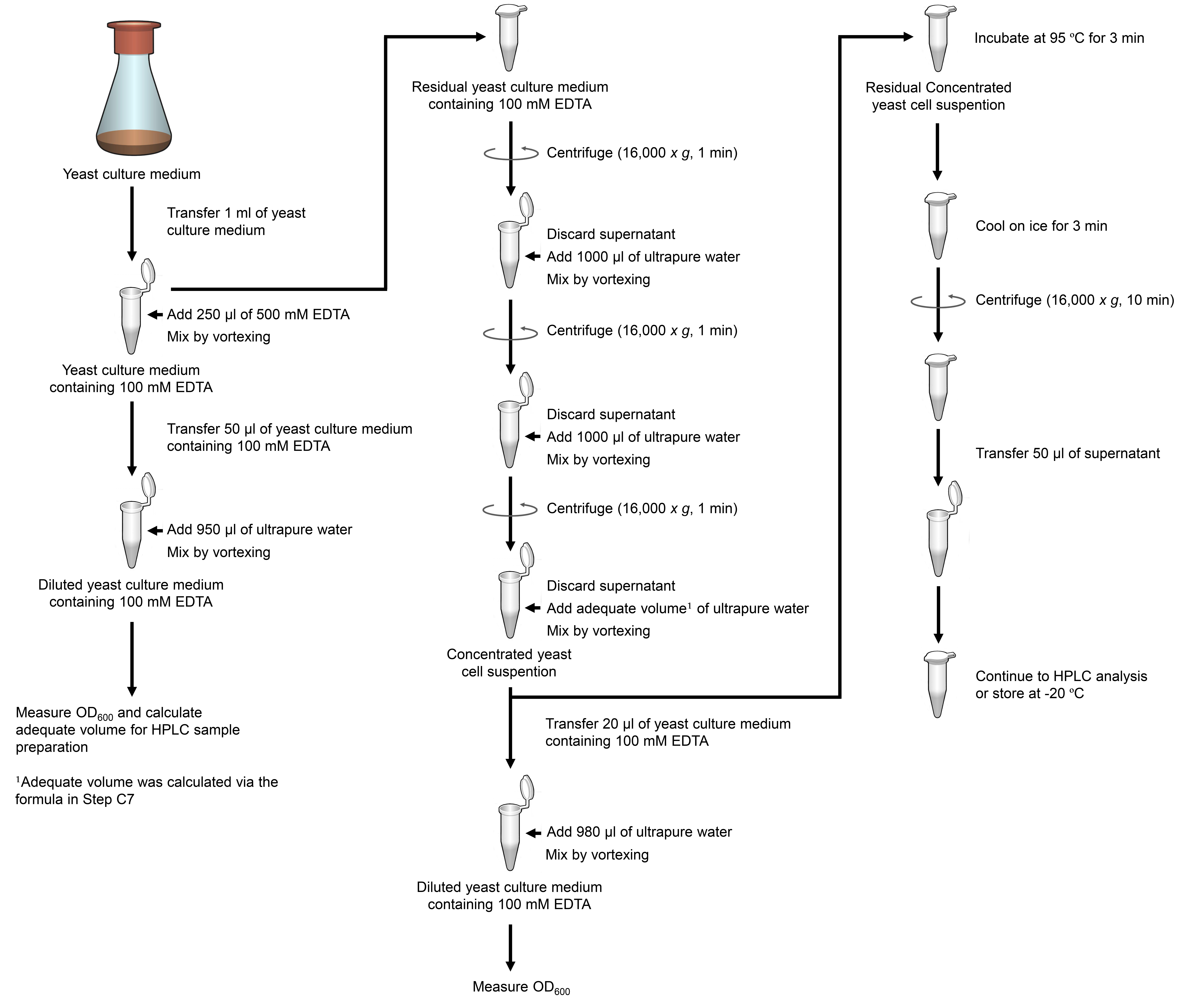
Figure 4. Schematic outline of measurement of cell concentration and cell disruption. A more detailed procedure is described in Procedure C.
- Transfer 1 ml of the yeast culture grown for glutathione production in a 1.5 ml microtube. Add 250 μl of 500 mM EDTA to the 1.5 ml microtube and mix thoroughly by vortexing.
- HPLC analysis
- Add 200 μl of the HPLC mobile phase to 50 μl of the cell content solution.
- Filter the 5-fold diluted cell content solution through the filter unit with 0.22 μm pore size to obtain an HPLC sample.
- Set each parameter as follows.
Pump: run time, 25 min; flow rate, 1 ml/min; maximum pressure, 20.0 MPa.
UV detector: deuterium arc lamp was used, response, 1.0 sec; internal temperature, 40 °C; wavelength, 210 nm.
Column oven: temperature, 30 °C; maximum temperature, 40 °C.
Auto sampler: injection volume, 20 μl; syringe speed, 15 μl/sec; flush volume, 200 μl; needle stroke, 52 mm; flush speed 35 μl/sec; temperature 10 °C. - Equilibrate the column to allow the mobile phase to flow for at least 90 min at a flow rate of 1 ml/min.
- Start analysis according to the HPLC manual to quantify reduced and oxidized glutathione in the HPLC sample. Examples of HPLC analysis are shown in Figure 5.
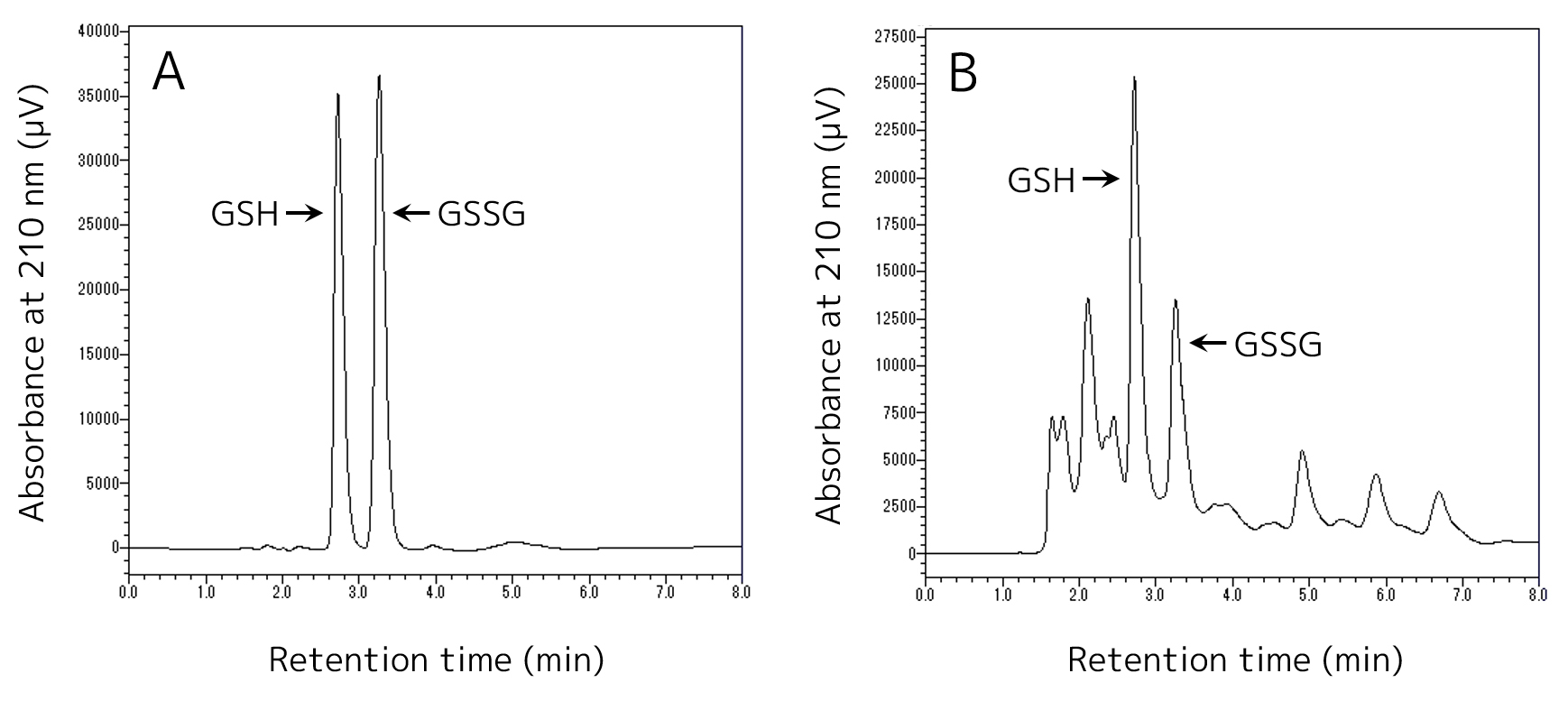
Figure 5. Examples of HPLC analysis. A. HPLC analysis of a sample containing 20 mg/L GSH and GSSG. B. HPLC analysis of a sample extracted from the S. cerevisiae GCI strain cells grown in the YPD medium at 30 °C for 48 h.
- Add 200 μl of the HPLC mobile phase to 50 μl of the cell content solution.
Data analysis
- Calculate cell concentration ([g-dry cell weight]/[L-broth]) from ODC and from the calibration curve of OD600 and biomass ([g-dry cell weight]/[L-broth]).
- Calculate the cell concentration rate by means of the following equation:

- Calculate volumetric glutathione production (VGP, mg/[L-broth]) via the following equation. In this equation, 5 means a dilution factor for the HPLC sample.
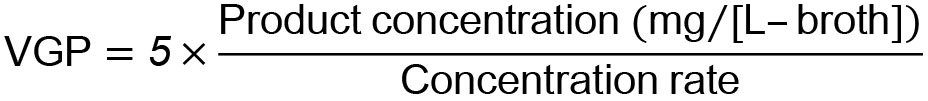
- Calculate glutathione content (GC) (mg/[mg-cells]) using the following equation:
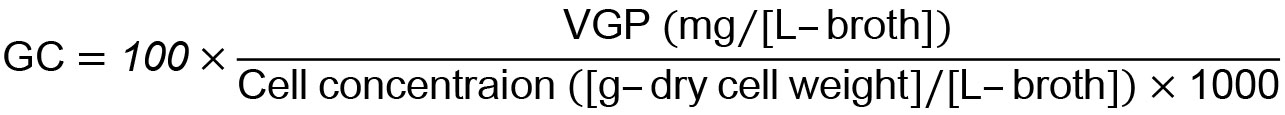
Notes
- If the resolution of an ODS column decreased, and HPLC peaks of GSH and GSSG became difficult to separate. An increase in methanol concentration in the HPLC mobile phase may improve the separation of the peaks.
- In this protocol, the addition of EDTA to yeast culture is needed to measure OD600 accurately. EDTA inhibits lectin activity on the cell surface and consequently prevents cell aggregation. If EDTA was not added, OD600 of yeast decreased via aggregation as compared with the original figure.
- When 250 μl of 500 mM EDTA was added to 1,000 μl of the yeast culture medium, and 50 μl of this mixture (total volume was 1,250 μl) was used to measure OD600; 1,200 μl of this residual mixture consisted of 960 μl of the yeast culture medium and 240 μl of 500 mM EDTA. The value of 960 in the equation in Step C7 is this residual volume of the original yeast culture medium.
- In many cases, GSH and GSSG concentrations in an HPLC sample were within the calibration curve, when OD600 of the cell suspension before disruption was ~35. Therefore, this protocol contained this value of 35 in the calculation of the adequate volume of ultrapure water.
Recipes
- YPD medium (1 L)
10 g of Yeast extract dried
20 g of hipolypeptone
20 g of D-(+)-glucose
1.0 ml of aureobasidin A stock solution (if needed)
20 g of agar powder (if needed)
Ultrapure water - Aureobasidin A stock solution (1 L)
0.5 mg of aureobasidin A
99.5% ethanol - 500 mM EDTA (1 L)
186.12 g of EDTA disodium salt dihydrate
Ultrapure water
Adjust pH to 8.0 with 1.0 M NaOH - 100 mM EDTA (1 L)
200 ml of 500 mM EDTA
Ultrapure water - HPLC mobile phase (1 L)
6.12 g of KH2PO4
1.818 g of 1-heptanesulfonate
212.6 ml of methanol
Ultrapure water
Adjust pH to 2.8 with 98% phosphoric acid
Acknowledgments
This work was supported by a Special Coordination Fund for Promoting Science and Technology, Creation of Innovative Centers for Advanced Interdisciplinary Research Areas (Innovative Bioproduction, Kobe) from the Ministry of Education, Culture, Sports, and Science and Technology (MEXT), Japan. This protocol was adapted from our previous works (Hara et al., 2012; Hara et al., 2015; Kiriyama et al., 2013; Kobayashi et al., 2017).
Competing interests
The authors have no conflict of interest or competing interest.
References
- Hara, K. Y., Aoki, N., Kobayashi, J., Kiriyama, K., Nishida, K., Araki, M. and Kondo, A. (2015). Improvement of oxidized glutathione fermentation by thiol redox metabolism engineering in Saccharomyces cerevisiae. Appl Microbiol Biotechnol 99(22): 9771-9778.
- Hara, K. Y., Kiriyama, K., Inagaki, A., Nakayama, H. and Kondo, A. (2012). Improvement of glutathione production by metabolic engineering the sulfate assimilation pathway of Saccharomyces cerevisiae. Appl Microbiol Biotechnol 94(5): 1313-1319.
- Kiriyama, K., Hara, K. Y. and Kondo, A. (2013). Oxidized glutathione fermentation using Saccharomyces cerevisiae engineered for glutathione metabolism. Appl Microbiol Biotechnol 97(16): 7399-7404.
- Kobayashi, J., Sasaki, D., Hara, K. Y., Hasunuma, T. and Kondo, A. (2017). Enzymatic improvement of mitochondrial thiol oxidase Erv1 for oxidized glutathione fermentation by Saccharomyces cerevisiae. Microb Cell Fact 16(1): 44.
Article Information
Copyright
© 2018 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Kobayashi, J., Sasaki, D. and Kondo, A. (2018). A Procedure for Precise Determination of Glutathione Produced by Saccharomyces cerevisiae. Bio-protocol 8(12): e2887. DOI: 10.21769/BioProtoc.2887.
Category
Microbiology > Microbial biochemistry > Protein
Biochemistry > Protein > Quantification
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










