- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Expansion of Airway Basal Cells and Generation of Polarized Epithelium
Published: Vol 8, Iss 11, Jun 5, 2018 DOI: 10.21769/BioProtoc.2877 Views: 14471
Reviewed by: Meenal SinhaAntoine de MorreeAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Primary Mouse Choroidal Endothelial Cell Culture
Qiuhua Yang [...] Yuqing Huo
Jun 20, 2025 2221 Views
Isolation and Culture of Ferret Airway Stem Cells
Ziying Yan [...] Feng Yuan
Jul 20, 2025 2439 Views
Optimization of Adipogenic Differentiation Protocol for Murine and Human Cell Culture Models
Junwan Fan [...] Wenyan He
Jan 20, 2026 284 Views
Abstract
Airway basal stem cells are the progenitor cells within the airway that exhibit the capacity to self-renew and give rise to multiple types of differentiated airway epithelial cells. This stem cell-derived epithelium displays organized architecture with functional attributes of the airway mucosa. A protocol has been developed to culture and expand human airway basal stem cells while preserving their stem cell properties and capacity for subsequent mucociliary differentiation. This achievement presents a previously unrealized opportunity to maintain a durable supply of progenitor cells derived from healthy donors to differentiate into human primary airway epithelium for cellular and molecular-based studies. Further, basal stem cells can be harvested from patients with a specific airway disease, such as cystic fibrosis, enabling investigation of potentially altered behavior of disease-specific cells in the appropriate context of the airway mucosa. Here we describe, in detail, a protocol for the serial expansion of airway basal stem cells to enable the generation of nearly unlimited airway basal cells that can be stored and readily available for subsequent culturing and differentiation. In addition, we describe culturing and differentiation of airway basal stem cells on permeable transwell filters at air-liquid interface to create functional mucociliary pseudostratified polarized airway epithelial mucosa.
Keywords: Airway basal stem cellsBackground
Airway disease modeling and drug discovery have benefited greatly from the development and use of primary airway epithelial cultures grown on permeable transwell filters at air-liquid interface (ALI). This model has several advantages over immortalized cell lines in that the primary epithelial cells can differentiate into an airway mucosa that features multiple epithelial cell types including ciliated, serosal, and basal cells and their arrangement is quite reflective of in vivo cellular organization. Primary ALI models exhibit functional micro-physiological processes including beating cilia and the ability to secrete mucus, features which are notably absent in the cell line-derived epithelial monolayer. Further, primary cells do not rely on artificial immortalization or transformation, as cell line-derived epithelial cells do, and therefore are unencumbered by the potential deranged signaling seen in cell lines, which can misrepresent processes occurring in airway epithelium. Despite these notable limitations, immortalized cell lines are widely employed to model and investigate the airway epithelium because primary airway epithelial cells present their own set of challenges. Primary epithelial cells fail to replicate after a few passages and must be continuously harvested and isolated to complete each set of studies. In addition, molecular biology techniques to alter or delete the expression of genes of interest are difficult to achieve and sustain with primary epithelial cells. These disadvantages, creating both cost and technical hurdles, have hindered the widespread usage of primary ALI cultures despite their obvious advantages for investigating the airway mucosa.
Recently, transforming growth factor-β superfamily signaling (BMP/TGFβ/SMAD signaling) activity was found to be suppressed in p63+ basal cell compartments, but highly active in differentiated apically positioned cells (Mou et al., 2016). These observations facilitated the establishment of a novel culture platform with the use of dual SMAD inhibition to overcome the growth arrest and irreversible differentiation encountered in standard culturing of primary cells. Importantly, epithelial basal cell culture no longer relies upon co-culture of primary epithelial cells with mitotically inactive fibroblast feeder layers, a technique established in the 1970’s to enhance epithelial cell proliferation by improving the capacity of cultured cells to escape senescence (Rheinwald and Green, 1975). With this newly discovered ability to expand airway basal stem cells, patient-specific cells can be isolated and preserved from small biopsies of the airway, generating a virtually limitless supply of patient-specific airway epithelium in vitro (Mou et al., 2016). When differentiated on air-liquid interface, airway basal stem cell-derived epithelium forms a polarized mucociliary layer that exhibits proper epithelial architecture, physiology and response to clinically relevant pharmacologic agents (Mou et al., 2016; Yonker et al., 2017a and 2017b). Further, airway disease can be studied by generating basal stem cell-derived epithelium from individuals with airway disease, such as cystic fibrosis (CF), a genetic disease caused by a mutation in the CFTR gene, resulting in progressive respiratory failure. Primary ALI mucosa can be used to explore and understand cellular and molecular mechanisms that contribute to CF disease pathology.
Chronic infection and aberrant inflammation are hallmarks of CF (Yonker et al., 2015). Persistent infection with Pseudomonas aeruginosa, a typical pathogen in the CF airway, contributes to overzealous inflammation marked by pathological neutrophilic breach of the airway mucosa. A better understanding of cellular and molecular mechanisms mediating neutrophil trans-epithelial migration has the potential to inform and improve the efficacy of anti-inflammatory therapeutic strategies critically needed to treat inflammation mediated lung damage associated with CF. Recently, we have applied our advanced culture method in a co-culture system with neutrophils to investigate neutrophil-airway epithelium interaction and P. aeruginosa-induced neutrophil trans-epithelial migration (Yonker et al., 2017a and 2017b), Our findings from cultured human airway basal stem cells differentiated on ALI are consistent with prior molecular mechanistic studies implicating epithelial-derived neutrophil chemoattractant hepoxilin A3, an arachidonic acid metabolite synthesized by 12-lipoxygenase, as key in driving P. aeruginosa-induced trans-epithelial migration (Yonker et al., 2017a and 2017b). Additionally, these studies revealed previously unknown cellular mechanisms associated with neutrophilic breach of the airway mucosal barrier. Neutrophils applied to the basolateral aspect of the infected airway mucosal ALI organize in clusters before migrating through the mucosal barrier in response to epithelial infection. These organized clusters of neutrophils persist along the apical surface following trans-epithelial migration before individual neutrophils begin detaching from clusters into the airspace compartment (Yonker et al., 2017a). These results demonstrate that our expandable airway basal stem cell culturing and ALI differentiation system has tremendous potential to be exploited in a variety of ways to better understand micro-anatomy, physiology, interaction with pathogens, and innate immunity at the airway mucosa.
Below, we describe, in detail, a protocol for the generation of unlimited airway basal cells for the development of functional airway mucosal models that exhibit relevant physiological functioning. This protocol overcomes several barriers associated with investigations involving primary cells and represents a robust cellular platform for human disease modeling and drug development.
Materials and Reagents
- For airway basal cell expansion
- 6-well tissue culture plate (Corning, Falcon®, catalog number: 353046 )
- 150 mm tissue culture-treated culture dish (Corning, catalog number: 430599 )
- T175 culture flask (Thermo Fisher Scientific, catalog number: 159910 )
- 500 ml filter units (Merck, catalog number: SCGPU05RE )
- 804G cells
Note: 804G is a rat bladder epithelial cell line. It secretes matrix proteins enriched in laminin and collagen. The 804G-conditioned medium can be used as a cost-efficient coating medium. - Epithelial basal cell culture medium (Lonza, catalog number: CC-3118 or PromoCell, catalog number: C-21170 )
- Airway culture medium LHC-9 (Thermo Fisher Scientific, GibcoTM, catalog number: 12680013 )
- A8301 (antagonist of TGF/SMAD signaling, Tocris Bioscience, catalog number: 2939 )
- DMH-1 (antagonist of BMP4/SMAD signaling, Tocris Bioscience, catalog number: 4126 )
- CHIR99021 (agonist of WNT signaling, Tocris Bioscience, catalog number: 4423 )
- Y27632 (ROCK Inhibitor, Tocris Bioscience, catalog number: 1254 )
- Trypsin (Thermo Fisher Scientific, GibcoTM, catalog number: 25200056 )
- Penicillin-Streptomycin (Thermo Fisher Scientific, GibcoTM, catalog number: 15140122 )
- Dulbecco's phosphate-buffered saline (Thermo Fisher Scientific, GibcoTM, catalog number: 14190144 )
- RPMI 1640 medium, GlutaMAXTM supplement (Thermo Fisher Scientific, GibcoTM, catalog number: 61870036 )
- Fetal bovine serum (Thermo Fisher Scientific, GibcoTM, catalog number: 16000-044 )
- Complete Airway Basal Cell Culturing Medium (see Recipes)
- 6-well tissue culture plate (Corning, Falcon®, catalog number: 353046 )
- For airway-liquid-interface differentiation
- 0.22 μm filter (EMD Millipore, catalog number: SLGP033RS )
- 6.5 mm2 Transwell® with 0.4 µm Pore Polyester Membrane Insert, Sterile (Corning, catalog number: 3470 ), packaged 12 inserts in a 24-well plate
- 12 mm2 Transwell® with 0.4 µm Pore Polyester Membrane Insert, Sterile (Corning, catalog number: 3460 ), packaged 12 inserts in a 12-well plate
- PneumaCultTM-ALI Medium (STEMCELL Technologies, catalog number: 05001 )
- Hydrocortisone (STEMCELL Technologies, catalog number: 07904 )
- Heparin solution (STEMCELL Technologies, catalog number: 07980 )
- Sodium chloride (Thermo Fisher Scientific, InvitrogenTM, catalog number: AM9759 )
- 100% ethanol (Sigma-Aldrich, catalog number: E7023 )
- Complete ALI Medium (see Recipes)
- Hydrocortisone Solution (see Recipes)
- 0.22 μm filter (EMD Millipore, catalog number: SLGP033RS )
- For immunofluorescence
- Slides Thermofrost (Fisher Scientific, FisherbrandTM, catalog number: 12-550-15 )
- FisherbrandTM Cover Glasses: Rectangles (Fisher Scientific, FisherbrandTM, catalog number: 12-544-E )
- 2 ml Eppendorf tubes (Eppendorf, catalog number: 022431048 )
- ImmEdge Hydrophobic Barrier PAP Pen (Vector Laboratories, catalog number: H-4000 )
- Fluoromount-G (SouthernBiotech, catalog number: 0100-01 )
- DAPI (4',6-diamidino-2-phenylindole) (Thermo Fisher Scientific, Thermo ScientificTM, catalog number: 62247 )
- Triton X-100 (Sigma-Aldrich, catalog number: X100-500ML )
- 4% paraformaldehyde (Santa Cruz Biotechnology, catalog number: sc-281692 )
- Bovine serum albumin (Sigma-Aldrich, catalog number: A2153-100G )
- Sucrose (Sigma-Aldrich, catalog number: S7903 )
- OCT compound (Fisher Scientific, catalog number: 23730571 )
- Primary and secondary antibodies (see Table 1 below)
Table 1. Primary and Secondary antibodies for immunofluorescenceCatalog Number Vendor Name Target Primary antibody ab76013 Abcam Rabbit monoclonal, anti-NKX2.1, 1:200
Reacts with: Mouse, Rat, HumanAirway epithelium Primary antibody ab23630 Abcam Rabbit polyclonal, anti-FOXA2, 1:200
Reacts with: Mouse, Rat, HumanAirway epithelium Primary antibody ab53121 Abcam Rabbit polyclonal, anti-Cytokeratin 5, 1:500-1:1,000
Reacts with: Mouse, Rat, HumanBasal cells Primary antibody GTX102425 GeneTex Rabbit polyclonal, anti-p63, 1:300
Reacts with: Mouse, Rat, HumanBasal cells Primary antibody T6793 Sigma-Aldrich Mouse monoclonal (clone 6-11B-1), anti-Acetylated tubulin, 1:2,000-5,000
Reacts with: plant, pig, human, monkey, hamster, invertebrates, chicken, bovine, rat, frog, protista, mouseCiliated cells Primary antibody 14-9965-82 Thermo Fisher Scientific Mouse monoclonal (clone 2A5), anti-FOXJ1, 1:100
Reacts with: Mouse, HumanCiliated cells Primary antibody HPA031828 Sigma-Aldrich Rabbit polyclonal, anti-SCGB1A1 (CCSP), 1:200
Reacts with: HumanClub cells Primary antibody MA5-12178 Thermo Fisher Scientific Mouse monoclonal (clone 45M1), anti-Mucin 5AC antibody, 1:200-500
Reacts with: HumanGoblet cells Secondary antibody A-21202 ; A-21203 ; A-31571 Thermo Fisher Scientific Donkey anti-Mouse IgG (H+L), Alexa Fluor® 488, 594, 647 donkey anti-mouse IgG (H+L), 1:200 Mouse primary Ab Secondary antibody A-21206 ; A-21207 ; A-31573 Thermo Fisher Scientific Donkey anti-Mouse IgG (H+L), Alexa Fluor® 488, 594, 647 donkey anti-rabbit IgG (H+L), 1:200 Rabbit primary Ab Secondary antibody A-11055 ; A-11058 ; A-21447 Thermo Fisher Scientific Donkey anti-Mouse IgG (H+L), Alexa Fluor® 488, 594, 647 donkey anti-goat IgG (H+L), 1:200 Goat primary Ab
Equipment
- 250 ml bottles (Corning, PYREX®, catalog number: 1395-250 )
- Incubator at 37 °C with 5% CO2 (CO2 water jacketed incubator, Thermo Fisher Scientific, Thermo ScientificTM, catalog number: 3111 )
- Pipettes (PIPETMAN L Starter Kit, Gilson, catalog number: F167350 )
- Centrifuge with swing bucket rotor for 15 ml and 50 ml centrifuge tubes (Beckman Coulter, model: CS-6 )
- Vacuum pump
- Sterile laminar flow hood (The Baker Company, Sterilgard II )
- Hemostat or tweezers
- Scalpel
- 37 °C water bath (VWR, catalog number: 89501-468 )
- 4 °C cold room
- Inverted Light microscope (Nikon, diaphot TMD)
- Olympus Fluoview FV10i confocal microscope
- Olympus IX81 inverted fluorescence microscope (Olympus, model: IX81 )
Software
- ImageJ software (downloaded from https://imagej.nih.gov/ij/)
Procedure
Part I: Serial expansion of human airway basal stem cells
- Preparation of the ‘Complete Airway Basal Cell Culturing Medium’ (see Recipe 1)
- Preparation of 804G-conditioned medium
- Culture 804G cells in RPMI + 10% FBS + 1:100 penicillin/streptomycin (optional) on 15-cm2 dishes or T175 flasks until they reach confluence. Change medium daily.
- Once the cells reach confluence, add a large volume of culture medium (60-100 ml) to each dish or flask.
- From this point, the medium will be collected to make 804G-conditioned medium. Change medium every other day. Collect the supernatant that is enriched with matrix proteins. The collection can be performed up to 4 times. Combine all collected supernatants, filter through 0.4 μm filter units and aliquot in 250 ml bottles. The 804G-conditioned medium can be stored at 4 °C for up to 2 months and -20 °C for up to 1 year.
- Culture 804G cells in RPMI + 10% FBS + 1:100 penicillin/streptomycin (optional) on 15-cm2 dishes or T175 flasks until they reach confluence. Change medium daily.
- Culture and serial expansion of human airway basal cells (Figures 1A and 1B)
- Human airway basal cells are isolated from fresh discarded lung tissues using a protocol reported previously (Fulcher et al., 2005). Ensure that you have obtained proper authorization from your institution review board to obtain human samples for experimentation.
- Pre-coat culture plates for all airway basal cell culture. To coat the plates, add sufficient 804G-conditioned medium to cover culturable areas (see Table 2), incubate the plates at 37 °C overnight, or for at least 2 h. Aspirate as much 804G-conditioned medium as possible or briefly rinse with PBS. It is our experience, human primary airway basal cells prefer 804G coating over collagen, fibronectin, matrigel and other combinations of coating media. Morphology of human airway basal cells cultured on 804G-coated plates is better than those cultured on collagen-coated plates. However, if you are culturing mouse cells rather than human cells, collagen-coated plates or uncoated plates are preferred.
Culture the airway basal cells in ‘Complete Airway Basal Cell Culturing Medium’ on 804G-coated plates until they reach 80-90% confluence. Change medium daily. Based on our experience, the initial cell density has minimal effects on cell morphology and cell proliferation rate. You can start the cell culture at a confluence from 3% to 30%. For the routine cell maintenance, we start the cells at 10-20% confluence; the cells will reach 80-90% confluence after 4 days.
Table 2. The approximate volume of 804G-conditioned medium that is needed to coat the culture plates
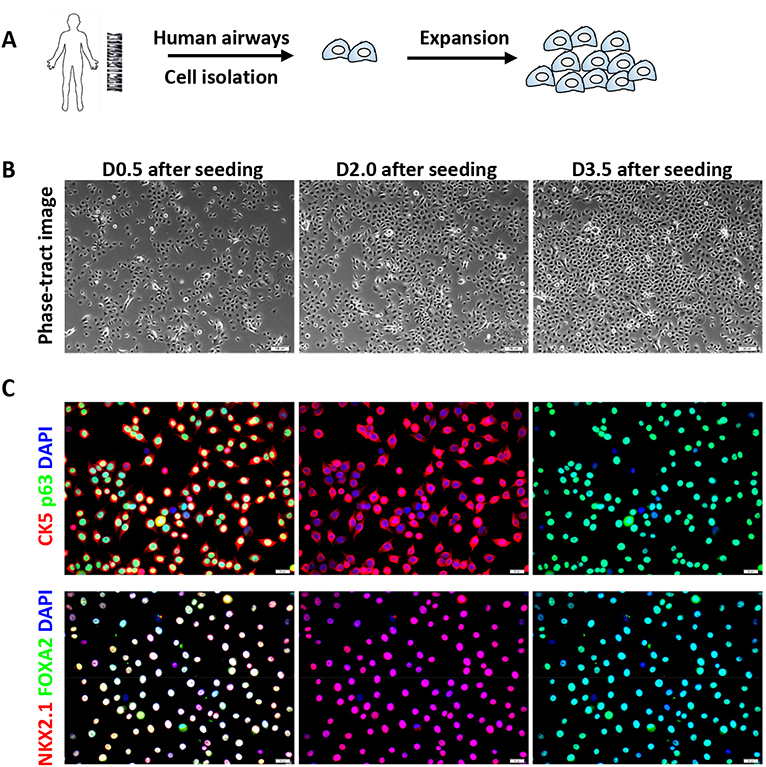
Figure 1. Serial expansion and immunofluorescence characterization of human airway basal cells. A. Schematic of serial expansion of human airway basal cells. B. Phase contrast images of the human basal cells at various days after cell seeding (initial seeding density is 10-15%). Scale bars = 50 μm. C. Immunofluorescence of basal stem cell markers (CK5 and p63) and airway-specific transcription factors (NKX2.1 and FOXA2) on cultured airway basal cells (Passage 6). Scale bars = 20 μm. - To split the cells, wash the cells with PBS, then add enough volume of 0.5% Trypsin to cover the cells and incubate at 37 °C. Monitor the cell dissociation under a microscope after 3 min. When the cells are loosened, stop trypsin digestion by adding DMEM (or RPMI) + 10% FBS. Gently transfer the cell suspension to a centrifuge tube.
- Centrifuge the cells at 300 x g for 5 min at room temperature.
- Aspirate the supernatant and resuspend the cell pellet in an appropriate volume of ‘complete Airway Basal Cell Culturing Medium’. Seed the airway basal cells at the desired density (from 5-20% confluence).
- Change the medium every day to maintain good morphology of the cells and their exponential proliferation rate. For a 6-well plate, add 1.5 ml to 2 ml per well.
- When the cells reach 80-90% confluence, split them again at 1:10 with the same procedure as described above. Human airway basal cells have a population doubling time around 30-35 h (Mou et al., 2016), so it takes ~4-5 days for another passage if the cells are split at 1:10.
- Human airway basal cells are isolated from fresh discarded lung tissues using a protocol reported previously (Fulcher et al., 2005). Ensure that you have obtained proper authorization from your institution review board to obtain human samples for experimentation.
- Immunofluorescence characterization of cultured airway basal cells (Figure 1C)
- Assess the stemness of airway basal cells every 3-4 passages along the course of their serial expansion. The stemness of airway basal cells is scored with the following criteria: (a) Expression of stem cell markers (p63 and CK5), and airway-specific markers (NKX2.1 and FOXA2); (b) Possession of robust proliferation potential, meaning that cells can self-renew at an exponential rate; (c) Possession of differentiation potential, meaning that cells can differentiate into main types of airway epithelium including basal cells, ciliated cells, club cells, and mucin-secreting goblet cells (see Part II and Figure 2).
Note: The maximal passage number of a given airway basal cell line, until complete depletion of its proliferative potential, is variable and individual cell line-dependent. The airway basal cells can be expanded for a long time under the established conditions described herein. We are generally able to passage our cells to passage 25 or beyond, while maintaining the overall differentiation efficiency based on the staining of mucus-producing goblet cells and cilia-bearing ciliated cells. However, this potential likely declines with each passage. We have reported that the prolonged culture would influence some physiological functions such as CFTR expression and chloride conductance (Mou et al., 2016). The speed of the declination in channel function over cell passage is faster than the speed of the declination in the ability to proliferate and generate airway epithelium. Therefore, the recommendation of “working maximal passage number” is dependent on the cellular analysis to be performed. For example, if the one intends to study CFTR channel biology, we would suggest using the cells “younger” than passage 6-8 at 1:10 splitting ratio. Whereas, if one’s goal is to generate airway epithelium or to study the response of airway epithelium to cytokines and pathogens, he/she can expand the cells for more passages. Based on our data, airway epithelium differentiated from airway basal cells even at passage 20 still respond robustly to IL-13 to induce goblet cell hyperplasia.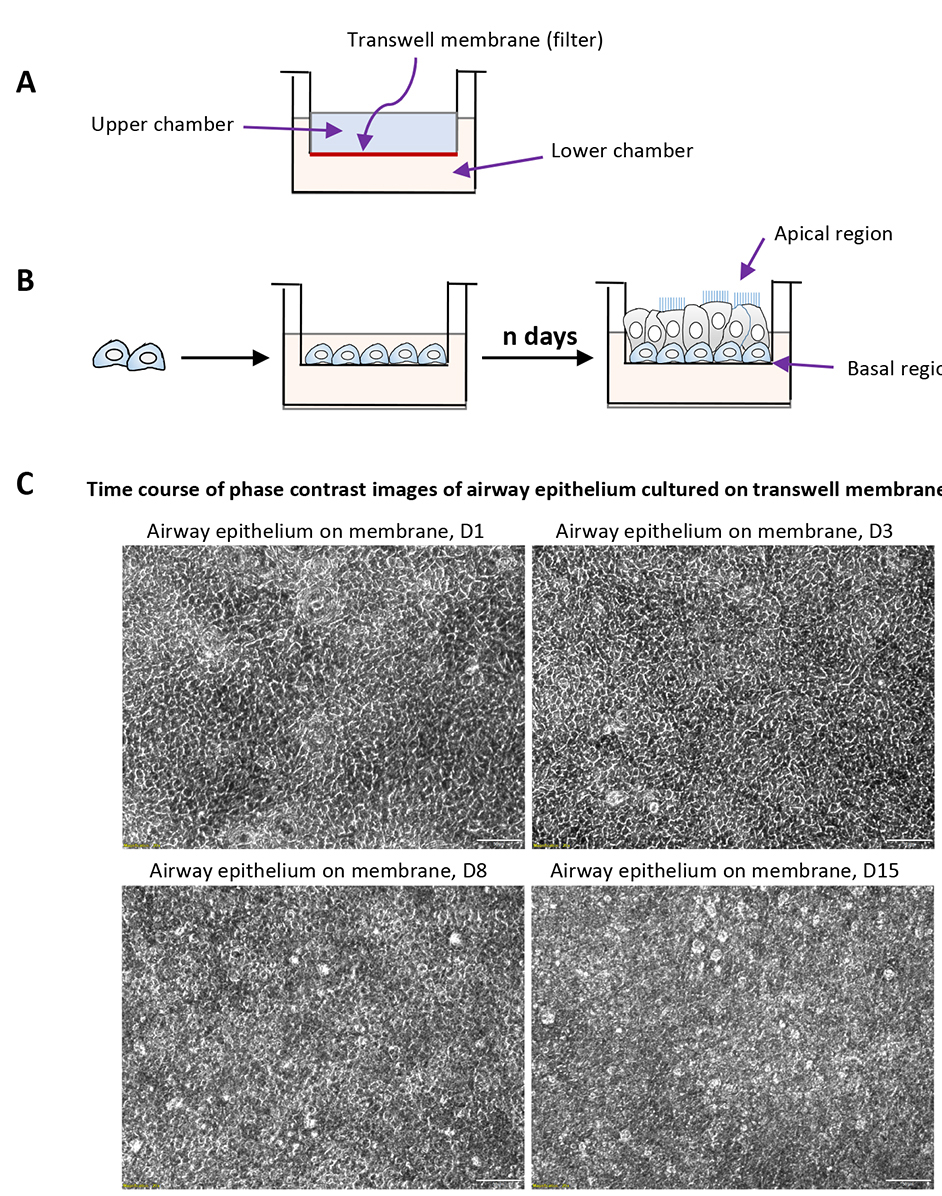
Figure 2. Mucociliary differentiation of human airway basal cells cultured on transwell membrane in airway-liquid interface. A. Schematic description of Transwell® permeable supports used for differentiation of airway basal cells; B. Schematic strategy of air-liquid interface differentiation; C. The phase contrast images of primary airway epithelial culture of ALI culture at variable time points. - Fix the cells in culture with 4% paraformaldehyde (PFA) for 5 min at room temperature.
- Wash the cells three times with PBS + 0.2% Triton X-100. Incubate the cells with washing buffer for 30 sec at room temperature for each interval of washing.
- Cover the cells with primary antibodies diluted in PBS + 1% BSA and incubate at room temperature for 2 h or at 4 °C overnight. For example, add 100 μl of antibody solution to each well of 48-well plate, or 500 μl of antibody solution to each well of 12-well plates.
- Wash the cells four times with PBS + 0.2% Triton X-100. Incubate the cells with washing buffer for 30 sec at room temperature for each interval of washing.
- Cover the cells with fluorescent-conjugated secondary antibodies diluted in PBS + 1% BSA and incubate at room temperature for 1 h (protect from the light). For example, add 100 μl of antibody solution to each well of 48-well plate, or 500 μl of antibody solution to each well of 12-well plates.
- Wash the cells four times with PBS + 0.2% Triton X-100. For most markers listed in Table 1, incubate the wash buffer for 10-30 sec.
- Cover the cells with enough 1 μg/ml DAPI solution in (Milli-Q®)-H2O. Incubate for 1 min and then image the cells with the Olympus IX81 inverted fluorescence microscope.
- Assess the stemness of airway basal cells every 3-4 passages along the course of their serial expansion. The stemness of airway basal cells is scored with the following criteria: (a) Expression of stem cell markers (p63 and CK5), and airway-specific markers (NKX2.1 and FOXA2); (b) Possession of robust proliferation potential, meaning that cells can self-renew at an exponential rate; (c) Possession of differentiation potential, meaning that cells can differentiate into main types of airway epithelium including basal cells, ciliated cells, club cells, and mucin-secreting goblet cells (see Part II and Figure 2).
Part II: Mucociliary differentiation of airway basal stem cells on the air-liquid interface
- Preparation of ‘Complete ALI Medium’ (see Recipe 2)
- Mucociliary differentiation of human airway basal cells on the air-liquid interface (Figure 2)
- Cell seeding (up to 1 day)
- A day prior seeding, pre-coat the transwell membranes with 804G-conditioned medium (for 24-well transwell, we add 150 μl coating medium/well; for 12-well transwell, we add 400 μl coating medium/well). Split the cells from the culture dishes and seed the cells at a density of ~5 x 103/mm2 in ‘Complete Airway Basal Cell Culturing Medium’ (see Part I).
- Add sufficient ‘Complete Airway Basal Cell Culturing Medium’ to both upper and lower chambers (for example, for a 12-well transwell plate, add 300-500 µl of culture medium to the upper chamber and 500-1,000 µl of culture medium to the lower chamber). At this seeding density, the cells should immediately reach 100% confluence upon attachment. Cell expansion on transwell membrane is not generally recommended since it will cause heterogeneity in differentiation and result in lower differentiation efficiency. If the cell number is not enough, briefly expand the cells until they reach 100% confluence (we suggest no more than 2 days of cell expansion).
- A day prior seeding, pre-coat the transwell membranes with 804G-conditioned medium (for 24-well transwell, we add 150 μl coating medium/well; for 12-well transwell, we add 400 μl coating medium/well). Split the cells from the culture dishes and seed the cells at a density of ~5 x 103/mm2 in ‘Complete Airway Basal Cell Culturing Medium’ (see Part I).
- Pre-ALI treatment (up to 1 day)
Remove ‘Complete Airway Basal Cell Culturing Medium’ from upper and bottom chambers, replace with ‘Complete ALI Medium’, incubate the cells at 37 °C overnight or at least for 2 h. - Mucociliary differentiation on ALI (> 15 days)
Remove the ‘Complete ALI Medium’ from the upper chamber to initiate an air-liquid interface. Add the same ALI medium only to the bottom chamber. Change medium every 2 days until the differentiation is well established. - Ciliogenesis can be monitored by inverted-phase microscopy (Video 1). The appearance of ciliated cells can be observed after 11-14 days depending on cell passage. We usually culture ALI for 16-20 days as a standard protocol. For cells at later passages, a longer time of ALI differentiation may be required. To facilitate the cell stratification (the basal cells need to stratify from a single cell layer to more layers to differentiate) and mucociliary differentiation efficiency especially from the aged airway basal cells, we recommend adding 1 μM A8301 + 5 μM ROCKi to ‘Complete ALI Medium’ for use during the first week of differentiation.Video 1. Human airway stem cells (P6) were differentiated on ALI for 20 days. The cilia beating was visualized under the Olympus IX81 inverted microscope. Scale bar = 20 μm.
- Cell seeding (up to 1 day)
- Whole mount immunofluorescence characterization of differentiated airway epithelium (Figure 3A)
- Aspirate the ALI medium in the lower chamber.
- Fix the cells on transwell membranes by adding PFA 4% to both upper chamber and bottom chamber. Place the plates at 4 °C for 10 min.
- Aspirate the PFA and wash the membranes 3 times with PBS. At this point, transwell membranes can be stored in PBS for up to one week at 4 °C until the immunofluorescence staining can be performed.
- Excise the membrane from the transwell using a sharp scalpel.
- Cut the membranes into 4 pieces (if 12 mm2 membranes are used) or cut the membranes into 2 pieces (if 6.5 mm2 membranes are used).
- Place each membrane piece in a 2 ml Eppendorf tube and add PBS + 0.2% Triton to permeabilize the cells. Incubate for 5 min.
- Aspirate the PBS + 0.1% Triton solution. Cover the membrane piece with 150 μl primary antibodies diluted in PBS + 1% BSA and incubate at room temperature for 2 h or at 4 °C for overnight.
- Wash the membrane piece four times with PBS + 0.2% Triton X-100.
- Cover the membrane piece with 150 μl fluorescent-conjugated secondary antibodies diluted in PBS + 1% BSA and incubate at room temperature for 1 h (protect from the light).
- Wash the cells four times with PBS + 0.2% Triton X-100.
- Cover the membrane piece with 150 μl 1 μg/ml DAPI solution in (Milli-Q®)-H2O. Incubate for 1 min.
- Add a small drop of Fluoromount-G on a microscope slide. Take out the membrane piece and put it (with the cell side face up) in the Fluromount-G drop. Gently add the coverslip, avoiding any bubble formation.
- Image the immunofluorescence using an Olympus Fluoview FV10i Confocal Microscope or a Nikon A1 Confocal Laser Microscope.
- Aspirate the ALI medium in the lower chamber.
- Immunofluorescence characterization of differentiated airway epithelium on transverse membrane sections (Figure 3B)
- Fix and excise transwell membranes as described above.
- Transfer the excised membranes in any containers to process the membranes for cryosectioning. We routinely use 12-well plates for this purpose. Incubate the excised membrane with 30% sucrose in PBS overnight at 4 °C.
- Transfer the membranes to 30% sucrose:OCT (1:1) and incubate for 1-2 h at room temperature.
- Transfer the membranes to 100% OCT and incubate for 15 min at room temperature.
- Cut the membranes into 2-4 pieces and embed the membrane in a vertical position in 100% OCT on a cold metal platform on dry ice.
- Cut 4-7 μm frozen sections using a standard cryo-sectioning protocol. Store the sections at -20 °C for up to 1 year or at -80 °C for a longer time.
- To stain the airway epithelial cells on transverse sections, remove the OCT by rinsing the slides briefly with tap water. Circle the tissue with ImmEdge Hydrophobic Barrier PAP Pen.
- Stain the slides sequentially with primary and secondary antibodies using a similar protocol described above.
- Image the immunofluorescence using an Olympus Fluoview FV10i Confocal Microscope or a Nikon A1 Confocal Laser Microscope.
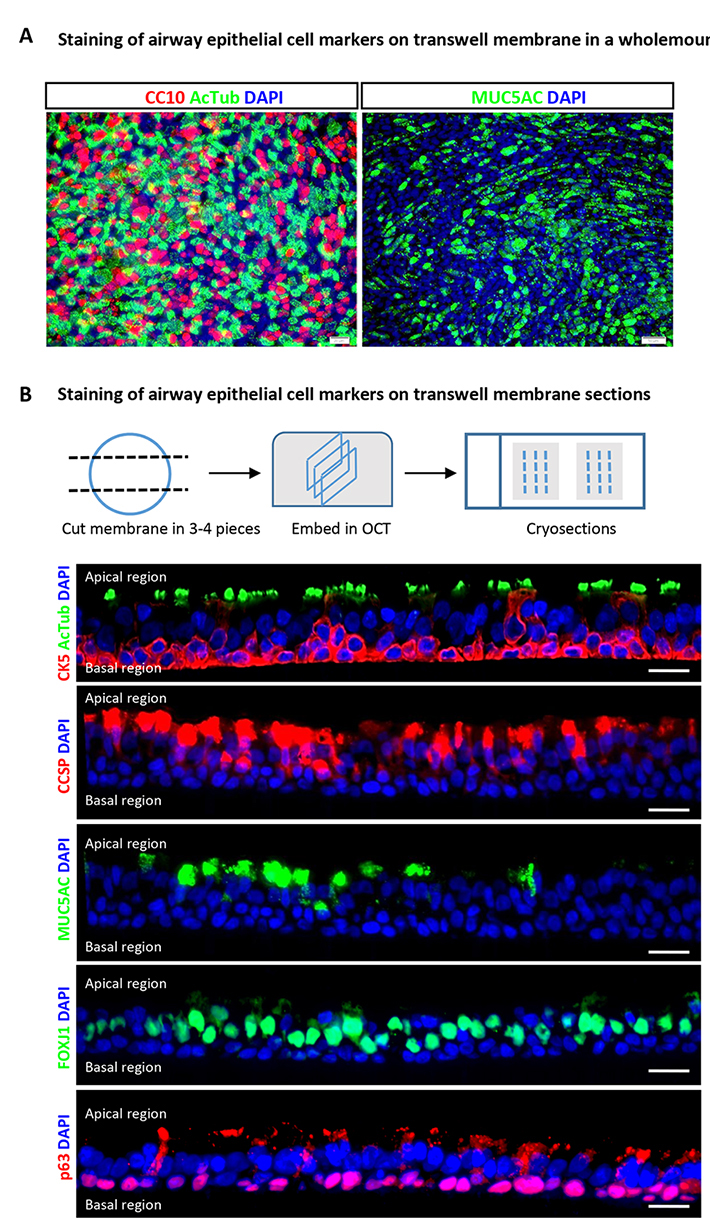
Figure 3. Immunofluorescence characterization of human airway basal cells cultured on airway-liquid interface. A. Immunofluorescence for markers of differentiation (acetylated-tubulin for ciliated cells, CC10 for club cells and MUC5AC for goblet cells) using wholemount staining. Scale bars = 50 μm. B. Immunofluorescence for airway epithelial cell markers (acetylated-tubulin and FOXJ1 for ciliated cells, CC10 for club cells and MUC5AC for goblet cells, and p63 for basal stem cells) on transverse ALI membrane sections. Scale bars = 20 μm.
- Fix and excise transwell membranes as described above.
Notes
ALI culture can be dissociated into single cell population for further cellular analyses. To dissociate the cells, aspirate the ALI medium from the lower chamber, rinse the cells gently with PBS, then add enough dissociation medium (0.25% trypsin + 0.5 mg/ml DNase I) to the upper and lower chamber to dissociate the cells from the membrane. For example, for the 12-well plate, add 0.5 ml dissociation medium to the upper chamber and 1 ml of dissociation medium to the bottom chamber. The goal of DNase I is to digest the DNA fragment released from the dead cells to decrease the viscosity of cell suspension. Monitor the cell dissociation under a microscope after 3-5 min. When the cells are loosened, stop trypsin digestion by adding DMEM (or RPMI) + 10% FBS. Transfer the cell suspension to a centrifuge tube and gently pipette several times to generate the single cell suspension. Centrifuge the cell suspension at 400 x g for 5 min and re-suspend the cell pellets in the desired solution for analysis. For example, the cells can be fixed in 4% PFA for 5 min for single-cell immunofluorescence assay after cytospinning. Alternatively, the cells can be suspended in PBS + 1% FBS for flow cytometry-based quantification assay. The success of flow cytometry is dependent on the availability of surface antibodies suitable for sorting. The authors suggest that readers optimize their sorting protocol independently to separate their particular cells of interest.
Data analysis
Image document and data analysis:
The staining on slides is visualized using an Olympus Fluoview FV10i Confocal Microscope or Olympus IX81 inverted fluorescence microscope. The staining of cells on culture dishes and the whole-mount staining on ALI transwell membranes are visualized with the Olympus IX81 inverted fluorescence microscope. Images are captured at multiple focal planes and combined using MicroSuite FIVE (Olympus Soft Imaging Solutions) and Extended Focal Imaging (EFI) module to create a single in-focus image, capturing the cellular complexities of the thicker ALI cultures.
The acquired images from the fluorescence microscope can be processed using the ImageJ software. The quantification will be performed by counting at least 5 random fields of view with a 20x or a 60x objective, and calculating the average and standard deviation.
Recipes
- Complete Airway Basal Cell Culturing Medium
Airway basal cell culture medium, 500 ml*
1.0 μM A-83-01 (50 μl of 10 mM stock) – TGFβ antagonist**
0.2 μM DMH-1 (5 μl of 10 mM stock) – BMP4 antagonist**
0.5 μM CHIR99021 (25 μl of 10 mM Stock) – WNT agonist**
5 μM Y27632 (250 μl of 10 mM stock) – ROCK inhibitor**
1:100 Penicillin-Streptomycin (Pen/Strep) – optional
Notes:- The medium can be prepared in advance in 500 ml rather than to add compounds freshly for each medium utility. To keep the activity of compounds, growth factors and other components in the complete medium, we suggest removing the desired amount of medium into a 50 ml centrifuge tube rather than warming up the whole bottle for each medium usage. The bottle of complete medium should be stored at 4 °C. The pre-prepared complete medium can remain effective for at least 1 month when stored at 4 °C.
- *For convenience, we routinely use commercial airway basal cell culture medium such as the medium from Lonza or Promocell. However, you can use the lab-made medium such as hTEC medium (Fulcher et al., 2005; You and Brody, 2013) or other commercially-available culture medium such as LHC-9 (Thermo Fisher).
- **The complete culture medium contains a combination of chemical compounds (TGFβ antagonist A-83-01, BMP4 antagonist DMH-1, WNT agonist CHIR99021, and ROCK inhibitor Y-27632). Other TGFβ antagonists, BMP4 antagonists, WNT agonists, and ROCK inhibitors can be used instead of A8301, DMH-1, CHIR99021 and Y-27632. If other compounds are used, the dose of the compound needs to be optimized separately. To determine the optimal dose of compounds, one can titrate the compound and measure the cell proliferation rate at the various concentrations (see Mou et al., 2016, Figure S2A). The best dose or the range of dose is the one that can promote the cell proliferation while causing minimal cell toxicity. In addition, the resulting cells should be homogenous, small, and tightly packed as expected for stem cells.
- The medium can be prepared in advance in 500 ml rather than to add compounds freshly for each medium utility. To keep the activity of compounds, growth factors and other components in the complete medium, we suggest removing the desired amount of medium into a 50 ml centrifuge tube rather than warming up the whole bottle for each medium usage. The bottle of complete medium should be stored at 4 °C. The pre-prepared complete medium can remain effective for at least 1 month when stored at 4 °C.
- Complete ALI Medium
500 ml PneumaCultTM-ALI Medium (Stemcell)
1 ml of Heparin sodium Salt
0.5 ml Hydrocortisone (see below) - Hydrocortisone Solution
200 μl of hydrocortisone stock (dissolve 120 mg hydrocortisone in 10 ml (Milli-Q®-H2O)
150 μl of 5 N NaCl
10 μl 100% ethanol
4620 μl (Milli-Q®)-H2O
Total 5 ml
Filter through 0.22 μm filter to sterilize
Aliquot and store at -20 °C
Add 0.5 ml each aliquot for 500 ml ALI medium
Final hydrocortisone in ALI is: 0.48 μg/ml
Acknowledgments
This protocol was adapted from the previously published work (Mou et al., 2016). The authors thank the entire Rajagopal Laboratory for their constructive assistance in the initial culture methodology development. The authors wish to thank the Cystic Fibrosis Foundation for a research funding (MOU16G0) and (HURLEY16G0) that supported the work in this manuscript, BPH was also supported by NIH R01 AI095338. We also wish to extend our thanks to members of the Hurley Laboratory and Mou Laboratory for their reading of the text, their constructive criticisms and comments and for valuable discussion and support.
Competing interests
All authors declare that they have no conflicts of interest or competing interests.
References
- Fulcher, M. L., Gabriel, S., Burns, K. A., Yankaskas, J. R. and Randell, S. H. (2005). Well-differentiated human airway epithelial cell cultures. Methods Mol Med 107: 183-206.
- Mou, H., Vinarsky, V., Tata, P. R., Brazauskas, K., Choi, S. H., Crooke, A. K., Zhang, B., Solomon, G. M., Turner, B., Bihler, H., Harrington, J., Lapey, A., Channick, C., Keyes, C., Freund, A., Artandi, S., Mense, M., Rowe, S., Engelhardt, J. F., Hsu, Y. C. and Rajagopal, J. (2016). Dual SMAD signaling inhibition enables long-term expansion of diverse epithelial basal cells. Cell Stem Cell 19(2): 217-231.
- Rheinwald, J. G. and Green, H. (1975). Serial cultivation of strains of human epidermal keratinocytes: the formation of keratinizing colonies from single cells. Cell 6(3): 331-343.
- Yonker, L., Cigana, C., Hurley, B. and Bragonzi, A. (2015). Host-pathogen interplay in the respiratory environment of cystic fibrosis. J Cyst Fibros 14: 431-439.
- Yonker, L. M., Mou, H., Chu, K. K., Pazos, M. A., Leung, H., Cui, D., Ryu, J., Hibbler, R. M., Eaton, A. D., Ford, T. N., Falck, J. R., Kinane, T. B., Tearney, G. J., Rajagopal, J. and Hurley, B. P. (2017a). Development of a primary human co-culture model of inflamed airway mucosa. Sci Rep 7(1): 8182.
- Yonker, L. M., Pazos, M. A., Lanter, B. B., Mou, H., Chu, K. K., Eaton, A. D., Bonventre, J. V., Tearney, G. J., Rajagopal, J. and Hurley, B. P. (2017b). Neutrophil-derived cytosolic PLA2α contributes to bacterial-induced neutrophil transepithelial migration. J Immunol 199(8): 2873-2884.
- You, Y. and Brody, S. L. (2013). Culture and differentiation of mouse tracheal epithelial cells. Methods Mol Biol 945: 123-143.
Article Information
Copyright
© 2018 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Levardon, H., Yonker, L. M., Hurley, B. P. and Mou, H. (2018). Expansion of Airway Basal Cells and Generation of Polarized Epithelium. Bio-protocol 8(11): e2877. DOI: 10.21769/BioProtoc.2877.
Category
Stem Cell > Adult stem cell > Epithelial stem cell
Cell Biology > Cell isolation and culture > Cell growth
Cell Biology > Cell isolation and culture > Cell differentiation
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link