- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Detection of Catalase Activity by Polyacrylamide Gel Electrophoresis (PAGE) in Cell Extracts from Pseudomonas aeruginosa
Published: Vol 8, Iss 11, Jun 5, 2018 DOI: 10.21769/BioProtoc.2869 Views: 9417
Reviewed by: Dennis NürnbergMichael EnosDeena Jacob

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
H2 Production from Methyl Viologen–Dependent Hydrogenase Activity Monitored by Gas Chromatography
Nuttavut Kosem
Dec 5, 2023 1799 Views
Monitoring Protein Stability In Vivo Using an Intein-Based Biosensor
John S. Smetana [...] Christopher W. Lennon
Apr 20, 2025 1620 Views
Endo-1,4-β-D-xylanase Assay Using Azo-Xylan and Variants Thereof
Luca Bombardi [...] Salvatore Fusco
Apr 20, 2025 1965 Views
Abstract
Bacteria in nature and as pathogens commonly face oxidative stress which causes damage to proteins, lipids and DNA. This damage is produced by the action of reactive oxygen species (ROS) such as hydrogen peroxide (H2O2), singlet oxygen, superoxide anion and hydroxyl radical. ROS are generated by antimicrobials, environmental factors (e.g., ultraviolet radiation, osmotic stress), aerobic respiration, and host phagocytes during infective processes. Pseudomonas aeruginosa, a versatile bacterium, is a prevalent opportunistic human pathogen which possesses several defense strategies against ROS. Among them, two catalases (KatA and KatB) have been well characterized by their role on the defense against multiple types of stress. In this protocol, KatA and KatB activities are detected by polyacrylamide gel electrophoresis (PAGE). It is also suggested that the detection of KatB is elusive.
Keywords: Pseudomonas aeruginosaBackground
P. aeruginosa is a ubiquitous bacterium that can be found in a free form in terrestrial and aquatics habitats or as an opportunistic human pathogen causing fatal infections in immunocompromised individuals, patients with skin damage or cystic fibrosis. To defend itself from ROS generated by its strong aerobic metabolism, host phagosomal vacuoles and environmental factors, this microorganism possesses several antioxidative strategies. Among them, two monofunctional catalases (KatA and KatB) are responsible for decomposing H2O2 to water and O2. KatA is the main catalase and has unique characteristics: it is unusually stable and essential to H2O2 resistance, osmoprotection and virulence (Hassett et al., 2000; Lee et al., 2005). It has been suggested that the stability of KatA is one of the main factors for the high level activity under normal growth conditions, and for this reason, katA has been regarded as a constitutively expressed gene in P. aeruginosa (Heo et al., 2010). However, it has been reported that KatA activity is induced in the stationary growth phase (up to 10-fold) and by increased levels of H2O2 (Brown et al., 1995; Suh et al., 1999; Heo et al., 2010). Moreover, katA expression has been demonstrated to be modulated by the global regulator OxyR and Quorum Sensing, whose activation depends on increased levels of H2O2 and high cell density, respectively (Hassett et al., 1999; Heo et al., 2010). KatB is only detected in the presence of H2O2 or paraquat and is partially involved in resistance to oxidative stress (Brown et al., 1995; Lee et al., 2005).
Solar ultraviolet-A (UVA) radiation is one of the main environmental stress factors for P. aeruginosa. Given the oxidative nature of UVA damage, we studied the role of catalases in defense of this microorganism against radiation. We demonstrated that KatA is essential in the optimal response against lethal doses of UVA, both in planktonic cells and biofilms (Costa et al., 2010; Pezzoni et al., 2014). In addition, we reported that low doses of UVA increase KatA and KatB activity and that this regulation occurs at the transcriptional level (Pezzoni et al., 2016). This phenomenon is relevant since it constitutes an adaptive mechanism that prevents cell damage by subsequent exposure to lethal doses of UVA, H2O2, or sodium hypochlorite (Pezzoni et al., 2016).
In the course of our studies, it became necessary to do an in-depth analysis of catalase activity. The total catalase activity in cell extracts was quantified by following spectrophotometrically the decomposition of H2O2, according to Aebi (1984). However, this assay cannot distinguish between KatA and KatB activities. To analyze individual catalase activity, we implement the method proposed by Wayne and Díaz (1986). In brief, crude cell extracts are loaded onto non-denaturing polyacrylamide gels (PAGE), and both catalases are separated by their differential electrophoretic motility; colorless bands of catalase activity are revealed by incubation of the gel with H2O2 and subsequent addition of a ferric chloride-potassium ferricyanide solution. The principle of this method involves the reaction of H2O2 with potassium ferricyanide (III) by reducing it to ferrocyanide (II); the peroxide is oxidized to O2. Ferric chloride reacts with ferrocyanide (II) to form an insoluble blue pigment. Because of the action of catalase on H2O2 decomposition, areas where this enzyme is active develop as clear bands in a blue gel (Patnaik et al., 2013). Additional papers were consulted to fine-tune this technique (Brown et al., 1995; Hassett et al., 1999; Elkins et al., 1999). The studies were performed with the prototypical P. aeruginosa strain PAO1 and isogenic derivatives PW8190 (katA::IslacZ/hah) and PW8769 (katB::IslacZ/hah) carrying mutations into katA and katB, respectively. Mutant strains devoid for KatA or KatB are useful to analyze the role of each enzyme in response to stress and as controls in PAGE catalase assays.
In this protocol, we describe how to detect individual catalase activity by PAGE using cell extracts from P. aeruginosa. Because of the particular characteristics of KatA (high abundance and stability), its detection does not present major difficulties. On the contrary, KatB detection is elusive, so that two changes were applied to the conventional technique: a protein extraction reagent was used instead of sonication to prepare the cell extracts, and the electrophoresis was performed at 4 °C. Based on these assays, it was concluded that KatB is an unstable enzyme, a fact that should be taken into account in quantitative or qualitative catalase assays under inducing (oxidative) conditions.
Materials and Reagents
- Pipette tips
- 50 ml sterile conical Falcon tubes (Nunc® EZ FlipTM, Thermo Fisher Scientific, catalog number: 362696 )
- 1.5 ml sterile Eppendorf centrifuge tubes (Eppendorf, catalog number: 022364111 )
- Sonication device
Note: This was assembled in our laboratory by attaching four plastic tubes (3 cm diameter, 3 cm high) to a plastic box (Figure 1).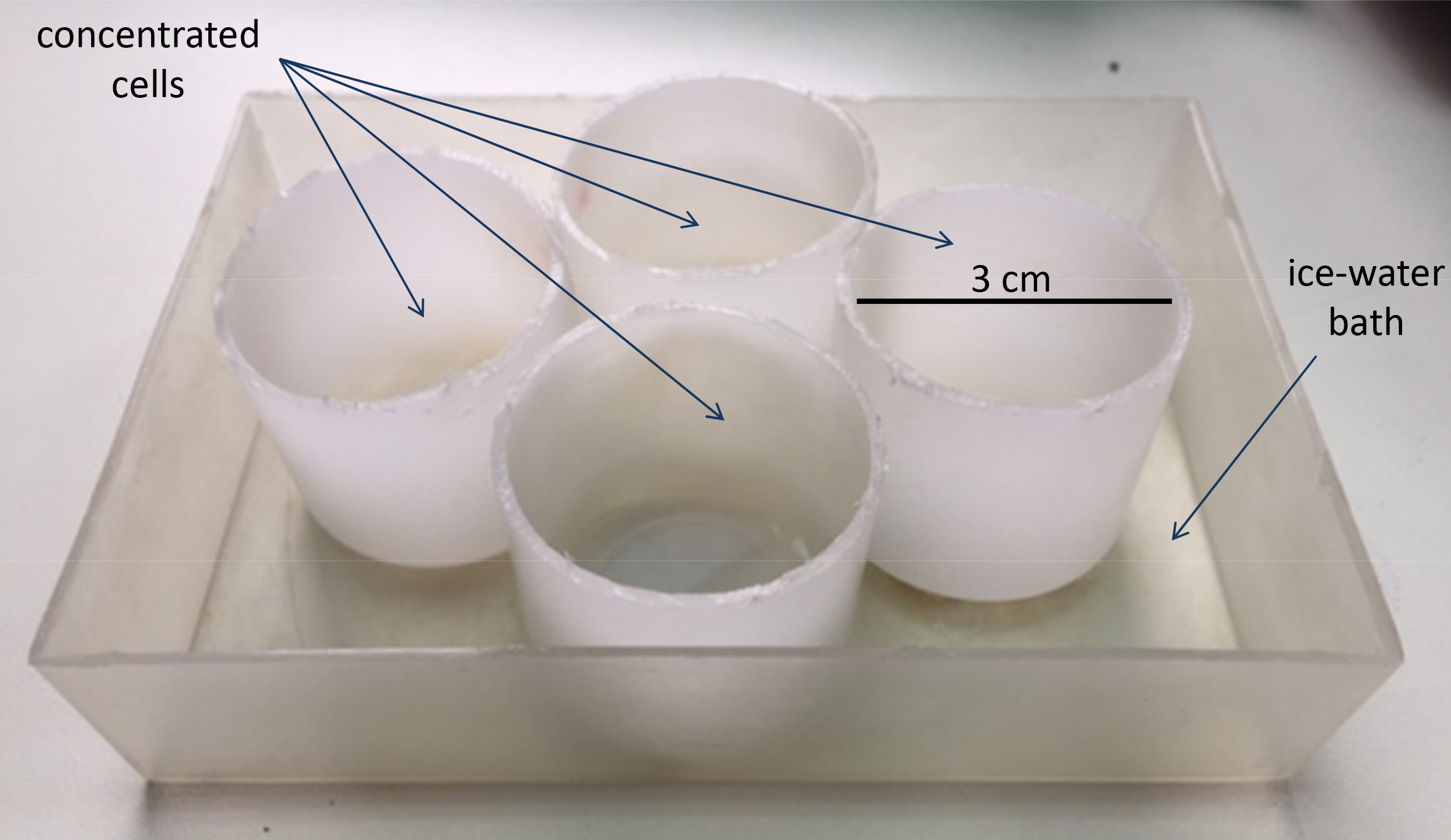
Figure 1. Sonication device - Paper towel (WypAll* X 60 Jumbo Roll, KCWW, Kimberly-Clark, catalog number: 30218593 )
- Spatula
- Pyrex tray (Pyrex® Storage 13 x 18 cm)
- P. aeruginosa strains
Note: PAO1, referred to as the wild-type, and catalase mutants PW8190 and PW8769 were obtained from the Washington Genome Center. Catalase mutants were constructed by insertion of IslacZ/hah transposon into katA (PW8190, hereinafter KatA-less strain) or katB (PW8769, hereinafter KatB-less strain) (Jacobs et al., 2003). - Distilled water
- Tryptone (Oxoid, catalog number: LP0042 )
- Yeast extract (Merck, catalog number: 103753 )
- Sodium chloride (NaCl) (Biopack, catalog number: 1646.08 )
- Albumin from bovine serum (Sigma-Aldrich, catalog number: A4378 )
- Sodium phosphate dibasic (Na2HPO4) (Sigma-Aldrich, catalog number: S3264 )
- Sodium phosphate (NaH2PO4) (Sigma-Aldrich, catalog number: S0751 )
- Hydrogen peroxide (H2O2) 30% (Merck, catalog number: 107210 )
- Bugbuster Protein Extraction Reagent (Merck, Novagen, catalog number: 70584-4 )
- Sodium thiosulfate (Na2S2O3) (Avantor Performance Materials, MacronTM, catalog number: 8100-04 )
- Ammonium persulfate ((NH4)2S2O8) (MP Biomedicals, catalog number: 04802811 )
- TEMED (MP Biomedicals, catalog number: 02195516 )
- Trizma base (Tris[hydroxymethyl]aminomethane) (C4H11NO3) (Sigma-Aldrich, catalog number: T1503 )
- Glycine (Sigma-Aldrich, catalog number: G7126 )
- Hydrochloric acid fuming 37% (HCl) (Merck, catalog number: 100317 )
- Acrylamide (Sigma-Aldrich, catalog number: A8887 )
- Bisacrylamide N,N’-methylene-bis-acrylamide (Sigma-Aldrich, catalog number: M7256 )
- EDTA (Merck, Calbiochem, catalog number: 324503 )
- Sodium hydroxide (NaOH) (Avantor Performance Materials, MacronTM, catalog number: 7708 )
- Glycerol (Merck, catalog number: 104094 )
- Bromophenol blue (VWR, DBH, catalog number: 20015 )
- Ferric chloride (FeCl3) (Avantor Performance Materials, MacronTM, catalog number: 5029-04 )
- Potassium ferricyanide (K3Fe (CN)6) (UCB, catalog number: b1599 )
- LB medium (see Recipes)
- 4 M NaCl (see Recipes)
- Saline solution (see Recipes)
- 50 mM sodium phosphate buffer, pH 7 (see Recipes)
- 30 mM H2O2 (see Recipes)
- 4 mM H2O2 (see Recipes)
- 10% ammonium persulfate (see Recipes)
- 1.5 M Tris-HCl buffer pH 8.8 (see Recipes)
- 1.5 M Tris-HCl buffer pH 6.8 (see Recipes)
- 30% acrylamide mix solution (acrylamide bisacrylamide ratio 37.5:1) (see Recipes)
- 6% resolving gel solution (see Recipes)
- 5% stacking gel solution (see Recipes)
- 1 M Tris-HCl buffer pH 8 (see Recipes)
- 0.5 M EDTA pH 8 (see Recipes)
- Loading sample buffer (see Recipes)
- Running buffer (see Recipes)
- Ferric chloride/potassium ferricyanide solution (see Recipes)
Equipment
- 50, 125 and 150 ml sterile Erlenmeyer flasks (DWK Life Sciences, Duran®, catalog numbers: 21 216 17 , 21 216 28 , 21 990 27 )
- 2-20 µl, 20-100 µl, 100-1,000 µl Kartell pluripet micropipettes (Kartell LABWARE, catalog numbers: 13000 , 13210 , 13220 ) and 1-10 ml Acura® manual micropipette (Socorex, model: Acura® manual 825 / Acura® manual 835 )
- 50, 100 and 1,000 ml borosilicate measuring cylinders (VILABO, catalog number: 3501114 , 3501115 , 3501118 )
- Conventional incubator shaker (New Brunswick Scientific, model: G25 )
- Gyratory water bath shaker (New Brunswick Scientific, model: G76 )
- Ice maker (Brema, model: TB 551 )
- UV-Vis Spectrophotometer (Biotraza, model: 752 )
- Refrigerated centrifuge (Hanil Scientific, model: Combi 514R )
- Vibra-Cell sonicator (Sonics & Materials, model: VC500 )
- Electrophoresis cell (Bioamerica, model: DYCZ-24DNBA )
- Power supply (Bioamerica, model: DYY-6CBA )
- Freezer ultra-low temperature (Sanyo, model: MDF-U76VC )
- Autoclave (HIRAYAMA, HICLAVETM, model: HVE-50 )
- Hot air oven sterilizer (Dalvo Intrumentos, model: OHR/T )
Procedure
- Preparation of cell extracts
Non-inducing conditions- Grow Pseudomonas aeruginosa strains (PAO1, KatA-less and KatB-less) overnight in 30 ml of LB medium in 150 ml Erlenmeyer flasks at 37 °C with shaking (200 rpm).
- Centrifuge the cultures (20 ml) in 50 ml Falcon tubes for 10 min, 10,000 x g, 4 °C. Discard the supernatants.
- Resuspend the cells with 20 ml of cold saline solution and keep them on ice.
- Centrifuge for 10 min, 10,000 x g, 4 °C. Discard the supernatants.
- Resuspend the cells in ice-cold 50 mM sodium phosphate buffer, pH 7 up to OD650 1 (about 7 ml).
- Sonicate 5 ml of concentrated cells in plastic tubes in an ice-water bath (Figure 1) under the following conditions: 2 min in pulsed mode, 18 mm tip diameter, 50% duty cycle, microtip limit 2. Keep the samples on ice.
- Centrifuge the extracts for 10 min, 10,000 x g, 4 °C. Discard the pellets carrying unlysed cells and cellular debris. Keep the supernatants on ice for a few hours until using them or store at -80 °C.
- Determine protein concentration of the extracts according to Lowry et al. (1951); bovine serum albumin is used as a standard.
Inducing conditions- Grow Pseudomonas aeruginosa strains (PAO1 and KatB-less) overnight in 30 ml of LB medium in 150 ml Erlenmeyer flasks at 37 °C with shaking (200 rpm).
- Dilute overnight cultures in 30 ml of fresh LB medium in 125 ml Erlenmeyer flasks to OD650 0.01 and grow in a gyratory water bath shaker at 37 °C until the cultures reach an OD650 0.3. This OD is reached in about 2 h.
- Divide the cultures into two fractions of 10 ml each in 50 ml Erlenmeyer flasks. Maintain one of them untreated (control) and add 2.26 µl of 30 mM H2O2 to the other fraction every 10 min for 1 h under sterile conditions, while shaking both fractions at 37 °C.
- Take 5 ml of the treated cultures at the end of 1 h and add 10 μl of 1 mg/ml sodium thiosulfate to neutralize the effect of H2O2; it is not necessary to keep the samples on ice.
- Centrifuge the neutralized cultures and 5 ml of the control cultures for 10 min, 10,000 x g, 4 °C. Discard the supernatants.
- Resuspend the cells with 10 ml of ice-cold 50 mM sodium phosphate buffer, pH 7.
- Centrifuge for 10 min, 10,000 x g, 4 °C. Discard the supernatants.
- Resuspend the cells in 1 ml of BugBuster Protein Extraction Reagent and keep for 15 min at room temperature.
- Centrifuge for 10 min, 16,000 x g, 4 °C. Keep the supernatants on ice for a few hours until using them or store at -80 °C.
- Determine protein concentration of the extracts according to Lowry et al. (1951); bovine serum albumin is used as a standard.
Notes:- Antibiotics are not added to culture media.
- BugBuster Protein Extraction Reagent was employed instead of sonication to detect KatB activity. This reagent is capable of cell wall perforation without denaturing soluble proteins. It provides an alternative to mechanical methods such as French Press or sonication for releasing proteins.
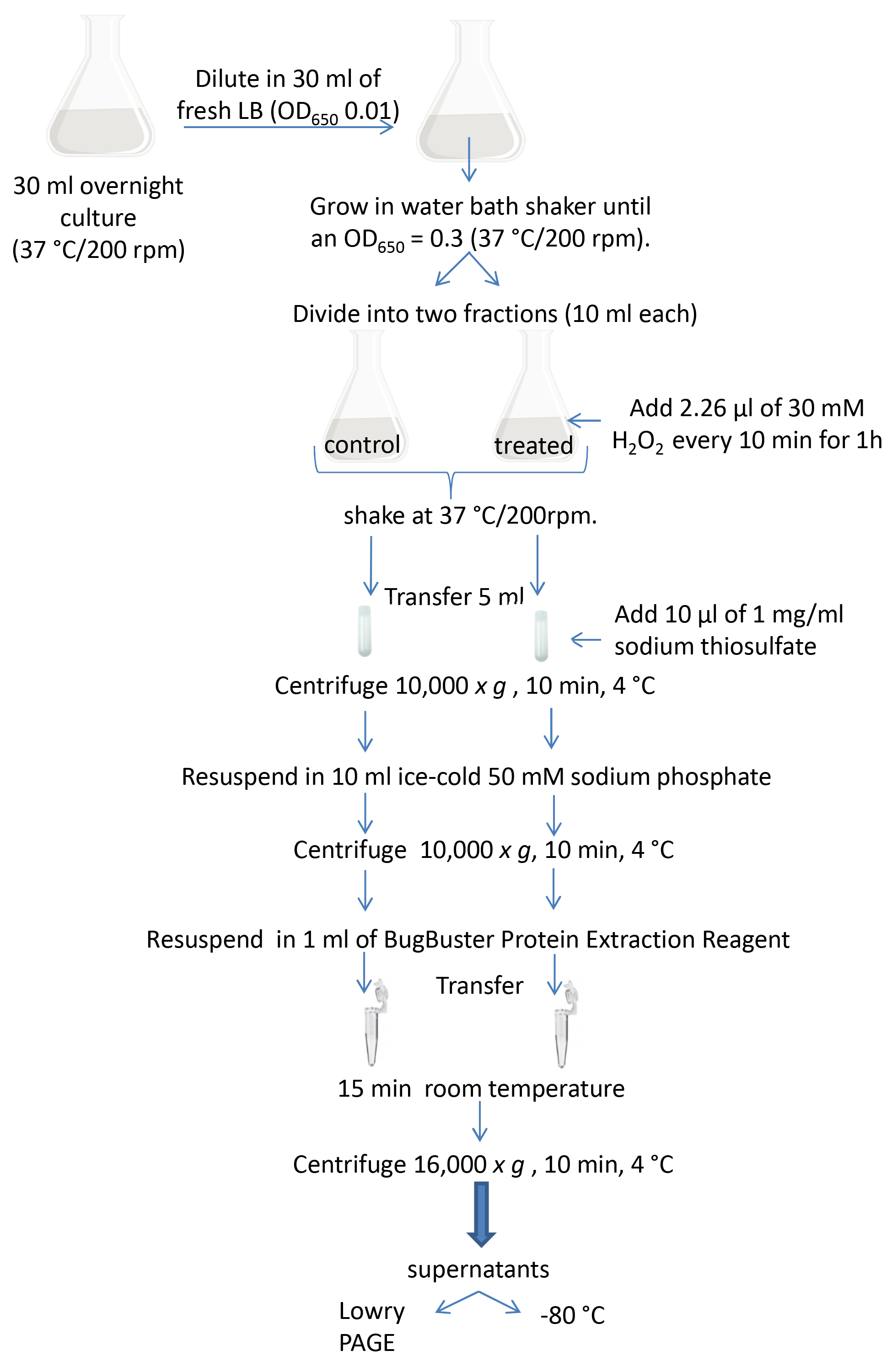 Figure 2. Schematic diagram of the experimental procedure
Figure 2. Schematic diagram of the experimental procedure - Grow Pseudomonas aeruginosa strains (PAO1, KatA-less and KatB-less) overnight in 30 ml of LB medium in 150 ml Erlenmeyer flasks at 37 °C with shaking (200 rpm).
- PAGE
- Set the two glass plates with the special wedge frames in the gel casting stand (see Figure 3).
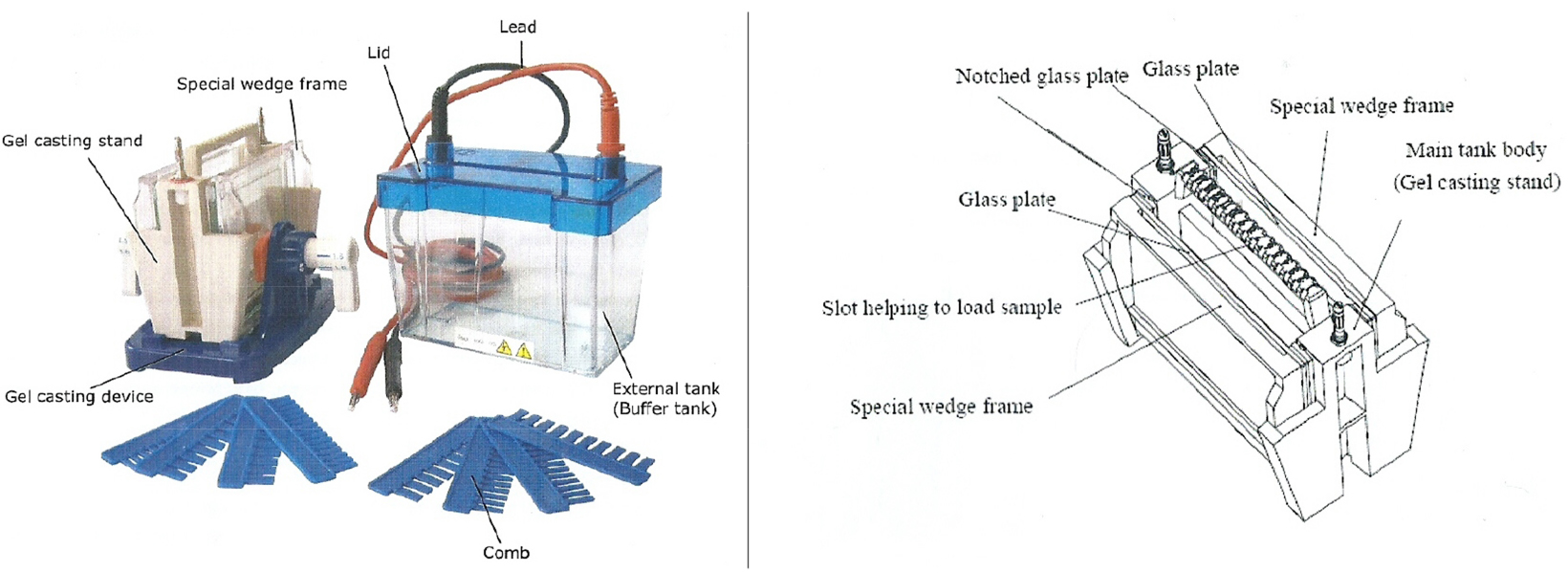
Figure 3. Electrophoresis cell (Bioamerica) - Pipet the 6% resolving gel solution into the gap between the glass plates.
- Wait for 15-30 min until it solidifies.
- Pipet the 5% stacking gel solution until overflow.
- Insert the comb without trapping air under the teeth. Wait for 15-30 min until it solidifies.
- Take the glass plates out of the gel casting stand and set them into the electrophoresis cell.
- Pour running buffer into the electrophoresis cell until the buffer level is higher than the top of the (shorter) inner gel plate (i.e., until the buffer covers the wells).
- Mix 50 µl of the cell extracts with 2 µl of loading buffer in 1.5 ml Eppendorf tubes and heat them in boiling water for 10 min.
- Load 10 µg of protein per sample into each well. Cover the cell with the lid and connect the electrodes to the power supply.
- Run the electrophoresis at 15 mA at room temperature (non-inducing conditions, KatA activity) or at 4 °C (inducing conditions, KatB activity).
Note: Electrophoresis is run at 4 °C for detecting KatB activity. This can be done in a cold room or in a refrigerator. However, this is not necessary for the detection of KatA activity. - Stop PAGE running when the dye front almost reaches the foot line of the glass plate. The run generally takes 4 h.
- Set the two glass plates with the special wedge frames in the gel casting stand (see Figure 3).
- Gel development
- Remove the glass plates from the electrophoresis tank and place them on a paper towel. Separate the plates by using a spatula.
- Soak the gel in distilled water in a Pyrex tray for 5 min at room temperature. Discard the water.
- Incubate the gel with 100 ml of a solution containing 4 mM H2O2 for 10 min at room temperature.
- Remove the solution and wash the gel with 100 ml of distilled water at room temperature.
- Soak the gel in 100 ml of a solution containing 1% (w/v) ferric chloride and 1% (w/v) potassium ferricyanide at room temperature.
- As soon as the gel turns dark green, remove the ferric chloride/potassium ferricyanide solution and rinse with distilled water to prevent overdevelopment.
- Once the dye has been removed, photograph the gel immediately. Storage in distilled water for a few days at 4 °C is possible. Areas of catalase activity show up as clear bands.
- Remove the glass plates from the electrophoresis tank and place them on a paper towel. Separate the plates by using a spatula.
Data analysis
According to this procedure described above, Figure 4 shows representative images of non-denaturing polyacrylamide gels stained for catalase activity employing non-inducing conditions (A) or inducing conditions with H2O2 (B).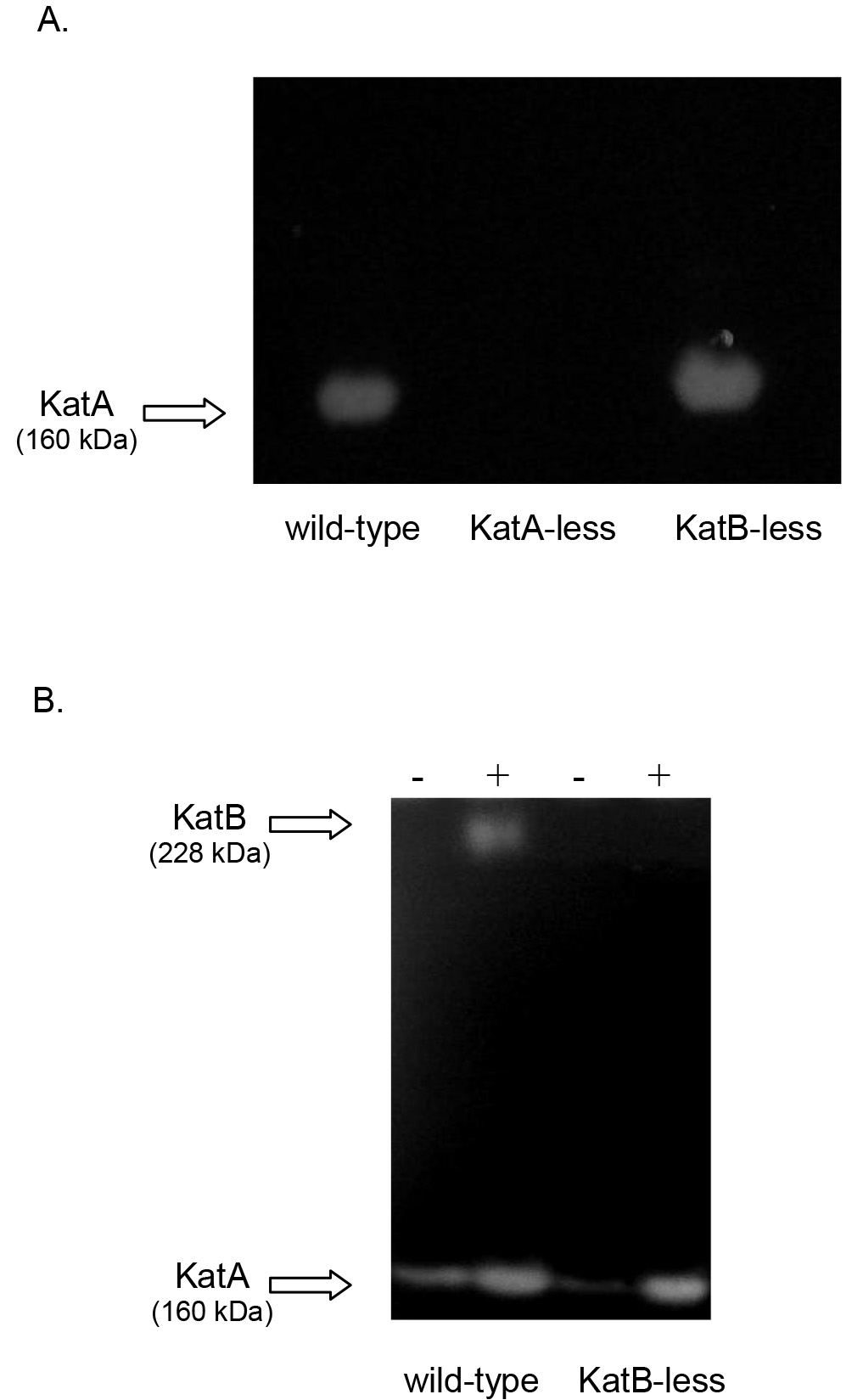
Figure 4. Catalase native PAGE analysis of P. aeruginosa wild type (PAO1) and its derivatives KatA-less and KatB-less strains. A. Extracts of PAO1 (wild-type), KatA-less and KatB-less strains grown under non-inducing conditions. The only activity detected corresponds to KatA since KatB is not expressed under this condition. B. Cultures of PAO1 and KatB-less strains, untreated (-) or treated with sublethal concentrations of H2O2 (+), were analyzed for catalase activity to demonstrate detection of KatB activity in non-denaturing polyacrylamide gels.
Note: The images were originally reported in Pezzoni et al., (2014). Protective role of extracellular catalase (KatA) against UVA radiation in Pseudomonas aeruginosa biofilms. J Photochem Photobiol B: Biol 131, 53-64.
Recipes
- LB medium
Dissolve:
10 g tryptone
5 g yeast extract
5 g NaCl
Bring the volume up to 1,000 ml in distilled water
Autoclave at 1 atm for 20 min - 4 M NaCl
Dissolve 46.7 g NaCl in 200 ml of distilled water
Autoclave at 1 atm for 20 min - Saline solution
Mix 7.5 ml sterile 4 M NaCl with 300 ml of sterile distilled water - 50 mM sodium phosphate buffer, pH 7
Solution A: dissolve 70.99 g Na2HPO4 in 500 ml of distilled water
Solution B: dissolve 59.98 g NaH2PO4 in 500 ml of distilled water
Mix 10.6 ml of solution A with 14.4 ml of solution B and add 475 ml distilled water - 30 mM H2O2
Add 0.340 ml of 30% H2O2 to 100 ml 50 mM sodium phosphate buffer, pH 7
Prepare fresh for every activity assay
The solution can be kept at room temperature during the experiment - 4 mM H2O2
Add 0.045 ml of 30% H2O2 to 100 ml distilled water
Prepare fresh for every activity assay - 10% ammonium persulfate
Dissolve 1 g ammonium persulfate in 10 ml distilled water
Store at -20 °C (shelf life 3 months) - 1.5 M Tris-HCl buffer pH 8.8
Dissolve 18.5 g Trizma base in 80 ml distilled water
Adjust to pH 8.8 with concentrated HCl and make up the volume to 100 ml - 1 M Tris-HCl buffer pH 6.8
Dissolve 12.114 g Trizma base in 80 ml distilled water
Adjust to pH 6.8 with concentrated HCl and make up the volume to 100 ml - 30% acrylamide mix solution
Dissolve 60 g acrylamide and 1.6 g bis acrylamide in 200 ml distilled water (acrylamide bisacrylamide ratio 37.5:1)
- Avoid directly contacting with polyacrylamide, ferricyanide and gels; they need to be handled with care.
- The order of adding solutions in the resolving and stacking gels solutions is important to avoid an early polymerization before pouring them between the glass plates.
- 6% resolving gel solution
5.4 ml distilled water
2 ml 30% acrylamide mix
2.5 ml 1.5 M Tris (pH 8.8)
0.1 ml 10% ammonium persulfate
0.008 ml TEMED - 5% stacking gel solution
2.1 ml distilled water
0.5 ml 30% acrylamide mix
0.38 ml 1.5 M Tris (pH 6.8)
0.03 ml 10% ammonium persulfate
0.003 ml TEMED - 1 M Tris-HCl buffer pH 8
Dissolve 12.114 g Trizma base in 80 ml distilled water
Adjust to pH 8 with concentrated HCl and make up the volume to 100 ml - 0.5 M EDTA pH 8
Dissolve 47 g EDTA in 200 ml distilled water
Adjust to pH 8 with concentrated NaOH and make up the volume to 250 ml - Loading sample buffer
1 ml 1 M Tris-HCl (pH 8)
4 ml 0.5 M EDTA (pH 8)
4 ml glycerol
25 mg bromophenol blue
1 ml distilled water
Store at -20 °C - Running buffer
Dissolve 3.03 g Trizma base and 14.4 g glycine in 1,000 ml distilled water
Store at 4 °C (shelf life 3 months) - Ferric chloride/potassium ferricyanide solution
Dissolve 1 g ferric chloride and 1 g potassium ferricyanide in 100 ml distilled water
Prepare fresh for every activity assay
Acknowledgments
The excellent technical assistance of Ms. P. Pereyra Schuth is gratefully acknowledged. This protocol is based on the work by Wayne and Díaz (1986). Financial support for this research was received from the Comisión Nacional de Energía Atómica (Argentina). M.P. is investigator at Consejo Nacional de Investigaciones Científicas y Técnicas (CONICET, Argentina).
Competing interests
The authors declare no conflict of interest.
References
- Aebi, H. (1984). Catalase in vitro. In: Methods in Enzymology. Parker, L. (Ed.). London, Academic Press 121-126.
- Brown, S. M., Howell, M. L., Vasil, M. L., Anderson, A. J. and Hassett, D. J. (1995). Cloning and characterization of the katB gene of Pseudomonas aeruginosa encoding a hydrogen peroxide-inducible catalase: purification of KatB, cellular localization, and demonstration that it is essential for optimal resistance to hydrogen peroxide. J Bacteriol 177(22): 6536-6544.
- Costa, C. S., Pezzoni, M., Fernandez, R. O. and Pizarro, R. A. (2010). Role of the quorum sensing mechanism in the response of Pseudomonas aeruginosa to lethal and sublethal UVA irradiation. Photochem Photobiol 86(6): 1334-1342.
- Elkins, J. G., Hassett, D. J., Stewart, P. S., Schweizer, H. P. and McDermott, T. R. (1999). Protective role of catalase in Pseudomonas aeruginosa biofilm resistance to hydrogen peroxide. Appl Environ Microbiol 65(10): 4594-4600.
- Hassett, D. J., Ma, J. F., Elkins, J. G., McDermott, T. R., Ochsner, U. A., West, S. E., Huang, C. T., Fredericks, J., Burnett, S., Stewart, P. S., McFeters, G., Passador, L. and Iglewski, B. H. (1999). Quorum sensing in Pseudomonas aeruginosa controls expression of catalase and superoxide dismutase genes and mediates biofilm susceptibility to hydrogen peroxide. Mol Microbiol 34(5): 1082-1093.
- Hassett, D. J., Alsabbagh, E., Parvatiyar, K., Howell, M. L., Wilmott, R. W. and Ochsner, U. A. (2000). A protease-resistant catalase, KatA, released upon cell lysis during stationary phase is essential for aerobic survival of a Pseudomonas aeruginosa oxyR mutant at low cell densities. J Bacteriol 182(16): 4557-4563.
- Heo, Y. J., Chung, I. Y., Cho, W. J., Lee, B. Y., Kim, J. H., Choi, K. H., Lee, J. W., Hassett, D. J. and Cho, Y. H. (2010). The major catalase gene (katA) of Pseudomonas aeruginosa PA14 is under both positive and negative control of the global transactivator OxyR in response to hydrogen peroxide. J Bacteriol 192(2): 381-390.
- Jacobs, M. A., Alwood, A., Thaipisuttikul, I., Spencer, D., Haugen, E., Ernst, S., Will, O., Kaul, R., Raymond, C., Levy, R., Chun-Rong, L., Guenthner, D., Bovee, D., Olson, M. V. and Manoil, C. (2003). Comprehensive transposon mutant library of Pseudomonas aeruginosa. Proc Natl Acad Sci U S A 100(24): 14339-14344.
- Lee, J. S., Heo, Y. J., Lee, J. K. and Cho, Y. H. (2005). KatA, the major catalase, is critical for osmoprotection and virulence in Pseudomonas aeruginosa PA14. Infect Immun 73(7): 4399-4403.
- Lowry, O. H., Rosebrough, N. J., Farr, A. L. and Randall, R. J. (1951). Protein measurement with the Folin phenol reagent. J Biol Chem 193(1): 265-275.
- Patnaik, S. C., Sahoo, D. K. and Chainy, G. B. (2013). A comparative study of catalase activities in different vertebrates. WebmedCentral Zoology 4(6):WMC004270.
- Pezzoni, M., Pizarro, R. A. and Costa, C. S. (2014). Protective role of extracellular catalase (KatA) against UVA radiation in Pseudomonas aeruginosa biofilms. J Photochem Photobiol B 131: 53-64.
- Pezzoni, M., Tribelli, P. M., Pizarro, R. A., Lopez, N. I. and Costa, C. S. (2016). Exposure to low UVA doses increases KatA and KatB catalase activities, and confers cross-protection against subsequent oxidative injuries in Pseudomonas aeruginosa. Microbiology 162(5): 855-864.
- Suh, S. J., Silo-Suh, L., Woods, D. E., Hassett, D. J., West, S. E. and Ohman, D. E. (1999). Effect of rpoS mutation on the stress response and expression of virulence factors in Pseudomonas aeruginosa. J Bacteriol 181(13): 3890-3897.
- Wayne, L. G. and Díaz, G. A. (1986). A double staining method for differentiating between two classes of mycobacterial catalase in polyacrylamide electrophoresis gels. Anal Biochem 157(1): 89-92.
Article Information
Copyright
© 2018 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Pezzoni, M., Pizarro, R. A. and Costa, C. S. (2018). Detection of Catalase Activity by Polyacrylamide Gel Electrophoresis (PAGE) in Cell Extracts from Pseudomonas aeruginosa. Bio-protocol 8(11): e2869. DOI: 10.21769/BioProtoc.2869.
Category
Microbiology > Microbial biochemistry > Protein > Activity
Biochemistry > Protein > Activity
Biochemistry > Protein > Electrophoresis
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










