- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Determination of Intracellular Osmolytes in Cyanobacterial Cells
Published: Vol 8, Iss 8, Apr 20, 2018 DOI: 10.21769/BioProtoc.2812 Views: 6962
Reviewed by: Valentine V TrotterAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Analysis of Heterocyst and Akinete Specific Glycolipids in Cyanobacteria Using Thin-layer Chromatography
Ritu Garg [...] Iris Maldener
Mar 20, 2022 2485 Views
Detecting Photoactivatable Cre-mediated Gene Deletion Efficiency in Escherichia coli
Yuta Koganezawa [...] Miki Umetani
Jun 5, 2023 1967 Views
Surface Plasmon Resonance for the Interaction of Capsular Polysaccharide (CPS) With KpACE
Zhe Wang [...] Chao Cai
Jun 20, 2025 3590 Views
Abstract
Most of the cyanobacteria accumulate osmolytes including sucrose, glucosylglycerol, in their cells in response to salt stress. Here we describe a protocol of our laboratory for extraction and quantification of cyanobacterial intracellular sucrose and glucosylglycerol. We have confirmed this protocol was applicable to at least four kinds of cyanobacteria, filamentous cyanobacterium Anabaena sp. PCC 7120, unicellular cyanobacterium Synechocystis sp. PCC 6803, Synechococcus elongatus PCC 7942 and halotolerant unicellular cyanobacterium Synechococcus sp. PCC 7002.
Keywords: OsmolyteBackground
Osmolytes (or compatible solutes) are a group of low-molecular-weight organic solutes, and play important physiological roles on abiotic stress acclimation in microbes including cyanobacteria (Reed and Stewart, 1985; Klähn and Hagemann, 2011; Slama et al., 2015). For determination of intracellular osmolytes from cyanobacterial cells, several protocols have been established (Reed and Stewart, 1985; Hagemann et al., 1997; Motta et al., 2004; Du et al., 2013; Fa et al., 2015).
Among these methods, nuclear magnetic resonance (NMR) spectroscopy based method was the only one which could be directly applied on microbe cultures without any extraction procedure (Motta et al., 2004). However, this protocol has just been tested in cultures of Halomonas pantelleriensis and Sulfolobus solfataricus rather than in cyanobacterial cultures. For all other methods, 80% ethanol was used for extraction of osmolytes from cyanobacterial cells. After derivatization by some trimethyl-silyl reagents, the derivatives of osmolytes could be analyzed by gas chromatography (Reed and Stewart, 1985). Alternatively, the extracted osmolytes could be directly analyzed by high-performance liquid chromatography (HPLC) (Hagemann et al., 1997). Our lab has firstly reported our protocol for osmolyte determination by ion chromatography (IC) equipped with a carb-Pac®MA1 analytical column (Du et al., 2013). In this protocol (Du et al., 2013), the column was equilibrated with 600 mM NaOH with a flow rate of 0.4 ml/min, and the running time for one sample was 45 min. Later, the concentration of NaOH was increased to 800 mM, and the running time for each sample was shortened to 32 min (Song et al., 2016 and 2017). Recently, the PA10 analytical column was successfully used for osmolyte analysis by ion chromatography (Qiao et al., 2017), and the running time for one sample was further shortened to 10 min. It is worthy to note that our collaborator has established a novel method for osmolyte determination by combination of separation by capillary ion chromatography and detection by mass spectrometry (CIC-MS) (Fa et al., 2015). The NMR and CIC-MS based methods are suitable for determination of unknown osmolytes from cyanobacteria. Compared with these two methods as well as the GC and HPLC based methods, our IC based method has advantages on running time and accuracy which would be helpful for the high throughput osmolyte detection used in some cyanobacterial metabolic engineering reseaches.
Therefore, we detailedly present our recent IC-based protocol here for osmolyte determination in cyanobacterial cells.
Materials and Reagents
- 2 ml microcentrifuge tubes (Cypress
- 10 ml Centrifuge tube, Snap-Cap (Kangjian)
- 1 ml syringe (Jianshi)
- Syringe membrane filters, 0.22 μm (Jinteng)
- Synechocystis sp. PCC 6803
- Nitrogen (Dehai)
- CO2 (Dehai)
- MilliQ water (Millipore, Germany)
- Potassium phosphate dibasic trihydrate (K2HPO4∙3H2O) (Sinopharm Chemical Reagent, catalog number: 10017518 )
- Magnesium sulfate heptahydrate (MgSO4·7H2O) (Sinopharm Chemical Reagent, catalog number: 10013018 )
- Calcium chloride dihydrate (CaCl2·2H2O) (Sinopharm Chemical Reagent, catalog number: 20011160 )
- Critic acid (Sinopharm Chemical Reagent, catalog number: 10007118 )
- Ferric ammonium citrate (Sinopharm Chemical Reagent, catalog number: 30011428 )
- EDTA·2Na (Sinopharm Chemical Reagent, catalog number: 10009717 )
- Sodium carbonate (Na2CO3) (Sinopharm Chemical Reagent, catalog number: 10019260 )
- Boric acid (H3BO3) (Sinopharm Chemical Reagent, catalog number: 10004818 )
- Manganese(II) chloride tetrahydrate (MnCl2·4H2O) (Sinopharm Chemical Reagent, catalog number: 20026118 )
- Zinc sulfate heptahydrate (ZnSO4·7H2O) (Sinopharm Chemical Reagent, catalog number: 10024018 )
- Sodium molybdate dihydrate (Na2MoO4·2H2O) (Sinopharm Chemical Reagent, catalog number: 10019818 )
- Copper(II) sulfate pentahydrate (CuSO4·5H2O) (Sinopharm Chemical Reagent, catalog number: 10008218 )
- Cobalt(II) chloride hexahydrate (CoCl2·6H2O) (Sinopharm Chemical Reagent, catalog number: 10007216 )
- Sodium nitrate (NaNO3) (Sinopharm Chemical Reagent, catalog number: 10019918 )
- Sodium chloride (NaCl) (Sinopharm Chemical Reagent, catalog number: 10019318 )
- Ethanol (Sinopharm Chemical Reagent, catalog number: 10009218 )
- Glycerol standard (99%, Sinopharm Chemical Reagent, catalog number: 10010618 )
- Glucosylglycerol standard (50%, Bitop, http://www.bitop.de/en/products/cosmetic-active-ingredients/glycoin)
- Glucose standard (Sinopharm Chemical Reagent, catalog number: 10010518 )
- Sucrose (Sinopharm Chemical Reagent, catalog number: 10021418 )
- 50% Sodium chloride (NaOH) solution (Thermo Fisher Scientific)
- BG11 medium (see Recipes)
- Saturated NaCl solution prepared in BG11 media (see Recipes)
- 80% ethanol (see Recipes)
- Osmolytes standards (see Recipes)
- 200 mM NaOH (see Recipes)
Equipment
- 200, 1,000 ml Pipettes (Eppendorf, Germany)
- Glass column photobioreactors (Sanhehuaxing, China) (Tan et al., 2011)
- Centrifuge (Sigma-Zentrifugen, model: Sigma 1-14 )
- Water bath (Yarong, model: B260 )
- Organomation (Hengao, model: HGC-24A )
- Ion chromatography (Thermo Fisher Scientific, Thermo ScientificTM, model: DionexTM ICS-5000+ )
- DinexTM CarboPacTM analytical column (4 x 250 mm, Thermo Fisher Scientific, model: DinexTM CarboPacTM PA10 )
- -80 °C freezer (Thermo Fisher Scientific, Thermo ScientificTM, model: Forma 705 )
- Vortex-Genie 2 (Scientific Industries, model: Vortex-Genie 2 )
- Autoclave (Hirayama, model: HV-50 )
Software
- Chromeleon software (version 6.80, Thermo Fisher Scientific)
- IBM SPSS Statistics (IBM, version 19)
Procedure
- Cultivation of Synechocystis sp. PCC 6803
Note: Monitor the growth of cyanobacterial cells by measuring the optical density at 730 nm (OD730) with a spectrophotometer. Culture volume should be less than 150 ml in 200 ml columns.- Inoculate Synechocystis cultures into liquid BG11 media in glass column photobioreactors at 30 °C under constant white fluorescent light with a light intensity of 100 μE/m2/sec. Adjust the initial OD730 to 0.5.
- Bubble the cultures with CO2-enriched air flow (3%).
- Add the saturated NaCl solution prepared in BG11 media into the late exponential phase culture (OD730 ≈ 8-10) to reach a final NaCl concentration of 600 mM.
- Inoculate Synechocystis cultures into liquid BG11 media in glass column photobioreactors at 30 °C under constant white fluorescent light with a light intensity of 100 μE/m2/sec. Adjust the initial OD730 to 0.5.
- Harvesting cells: Centrifuge 2 ml aliquots of the above Synechocystis cultures at 12,000 x g for 5 min at room temperature. Cells (Figure 1A) can be stored at -80 °C if not proceed to the next step immediately.
Note: Regularly, the highest intracellular glucosylglycerol concentration of Synechocystis would be reached within two days after salt shock. For sucrose, the highest concentration will be reached around 12 h after salt shock. If cells could not be extracted immediately, Synechocystis cells should be spun down by centrifugation and stored without supernatants at a -80 °C freezer. - Extraction of intracellular osmolytes from Synechocystis cells
- Re-suspend Synechocystis cells in 1 ml of 80% ethanol (v/v) (Figure 1B) and then incubate at 65 °C for 4 h.
Note: When incubating at 65 °C, it is better to mix the samples gently by inverting tubes once for each hour. - After centrifugation at 12,000 x g for 5 min at room temperature, transfer the supernatant to a clean 10 ml tube, and then dry at 55 °C under a stream of nitrogen.
- Dissolve the dry residues (Figure 1C) in 1 ml of Milli-Q water, and filter through membranes (0.22 μm).
Note: For drying, it takes about 30 min. If the extracted samples (Figure 1D) could not be analyzed immediately, they should be stored in 4 °C for less than seven days (-20 °C for less than 30 days), and filtered again before analyzing by ion chromatography.
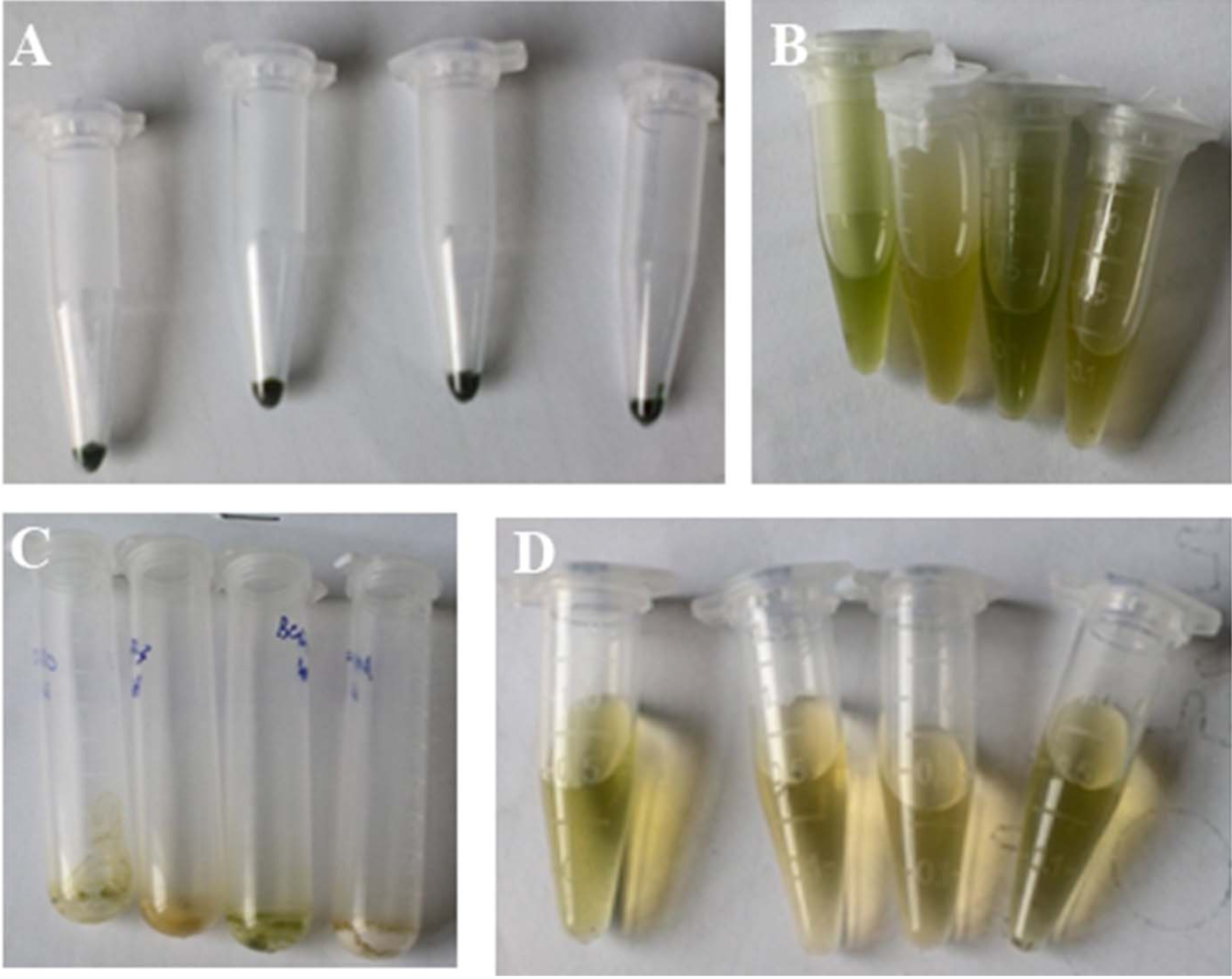

Figure 1. Extraction of osmolytes from Synechocystis cells. A. Cells of Synechocystis for an osmolyte extraction experiment; B. Synechocystis cells re-suspended in 80% ethanol; C. The dry residues dried at 55 °C under a stream of nitrogen; D. The extracted osmolyte samples from Synechocystis cells. - Re-suspend Synechocystis cells in 1 ml of 80% ethanol (v/v) (Figure 1B) and then incubate at 65 °C for 4 h.
- Detection of intracellular osmolytes by ion chromatography
- Dilute the extracted samples to the suitable concentrations ranging from 1~20 mg/L.
Note: Normally, the intracellular sucrose and glucosylglycerol concentrations in the wild type cells of Synechocystis sp. PCC 6803 are around 100 mg/L. Therefore, the samples should be diluted 5-10 fold. - Subject 25 μl of samples to ICS-5000+ ion-exchange chromatography system equipped with an electrochemical detector and a DinexTM CarboPacTM PA10 analytical column (4 x 250 mm, Thermo Fisher Scientific, Waltham, MA, USA). Elute the column with 200 mM NaOH at a flow rate of 1.0 ml/min.
- Mix the standards of glucosylglycerol, sucrose, and glycerol, dilute into 1, 5, 10, 15, 20 mg/L, and analyze by the same methods with the extracted samples.
- Dilute the extracted samples to the suitable concentrations ranging from 1~20 mg/L.
- Quantification of intracellular osmolytes
- For the standard mixtures with 15 mg/L of each osmolyte, retention times of glycerol, glucosylglycerol, glucose and sucrose using the method described above are 1.7, 2.1, 3.3 and 5.5 min, respectively (Figure 2A).
- Based on the mixed osmolyte standards, standard curves are made by using the peak area calculated by the Chromeleon software (Figure 2B).
- For quantification of intracellular osmolytes in cyanobacteria, areas of the target peak obtained by ion chromatography are calculated by the Chromeleon software, and then the osmolyte concentration is determined using the standard curve of osmolyte standards.
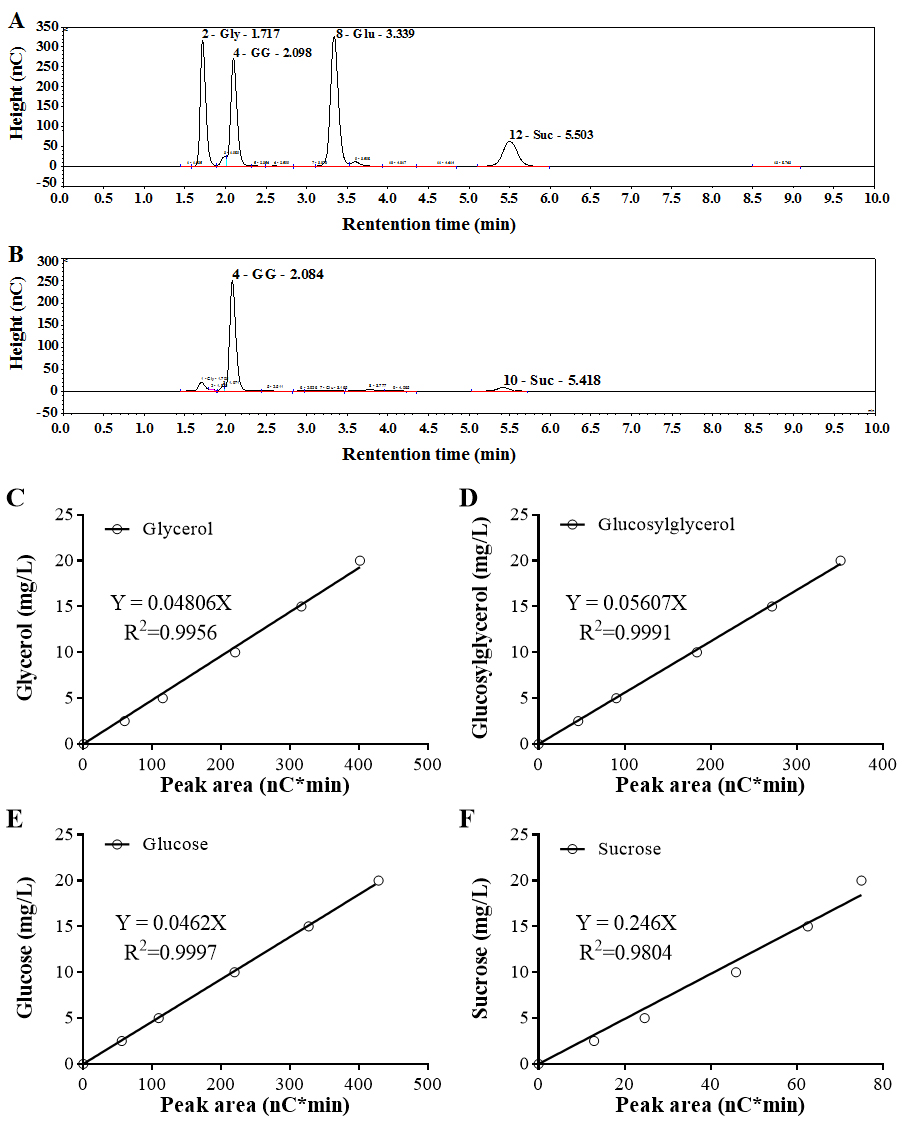
Figure 2. Ion chromatography profile and standard curves of osmolyte standards. A. Chromatogram of the standard mixture. 15 mg/L of each kind of osmolyte standard were mixed together and analyzed by ion chromatography. Gly, Glycerol; GG, Glucosylglycerol; Glu, Glucose; Suc, Sucrose. B. Chromatogram of the sample isolated from Synechocystis cells. Standard curves of glycerol (C), glucosylglycerol (D), glucose (E) and sucrose (F) were established by plotting peak areas and concentrations of each kind of osmolyte standard.
- For the standard mixtures with 15 mg/L of each osmolyte, retention times of glycerol, glucosylglycerol, glucose and sucrose using the method described above are 1.7, 2.1, 3.3 and 5.5 min, respectively (Figure 2A).
Data analysis
- For statistical analysis, IBM SPSS Statistics version 19 was used.
- For comparing differences between two data sets, an independent samples t-test was performed.
Recipes
- BG11 medium (Rippka et al., 1979)
- Prepare 8 kinds of 10x stocks as follows:
Stock 1 (40 g/L K2HPO4·3H2O)
Stock 2 (75 g/L MgSO4·7H2O)
Stock 3 (36 g/L CaCl2·2H2O)
Stock 4 (6 g/L critic acid)
Stock 5 (6 g/L ferric ammonium citrate)
Stock 6 (1 g/L EDTA·2Na)
Stock 7 (20 g/L Na2CO3)
Stock A5 (2.86 g/L H3BO3, 1.81 g/L MnCl2·4H2O, 0.22 g/L ZnSO4·7H2O, 0.39 g/L NaMoO4·2H2O, 0.08 mg/L CuSO4·5H2O, 0.01 g/L CoCl2·6H2O)
Autoclave Stocks 1 and 5 at 121 °C for 20 min. Store the autoclaved stocks and other stocks at 4 °C before use - Dissolve 1.5 g NaNO3 in 992 ml of ddH2O, add 1 ml of Stocks 2, 3, 4, 6, 7 and A5 into the medium respectively
- Autoclave the medium at 121 °C for 20 min, and supplement with 1 ml of both Stocks 1 and 5 before inoculations
- Prepare 8 kinds of 10x stocks as follows:
- Saturated NaCl solution prepared in BG11 media
Dissolve 292.5 g NaCl in BG11 media
Adjust the final volume with BG11 media to 1 L
Autoclave the NaCl solution at 121 °C for 20 min - 80% (v/v) ethanol
For 100 ml of ethanol solution, supplement 80 ml of ethanol with Milli-Q water to reach the final volume of 100 ml - Osmolytes standards
- First, dissolve 6 mg of glycerol, sucrose, glucose and glucosylglycero in 100 ml of Milli-Q water
- Then, prepare different concentrations of osmolytes standards according to the following table
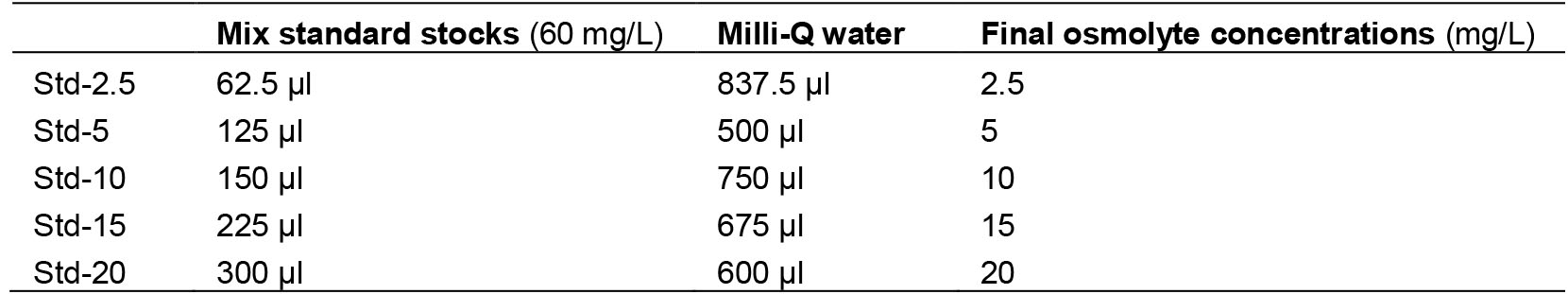

- 200 mM NaOH
For 1 L of 200 mM NaOH solution, dilute 16 ml of 50% NaOH solution to the final volume of 1 L with Milli-Q water
Acknowledgments
The protocol was adopted from the publication ‘Effects of lowered and enhanced glycogen pools on salt-induced sucrose production in a sucrose-secreting strain of Synechococcus elongatus PCC 7942’ (Qiao et al., 2017). This work was supported by the National Science Fund for Distinguished Young Scholars of China (31525002 to X. Lu), Shandong Key basic Research project (ZR2017ZB0211), the Joint Sino-German Research Project (grant GZ 984 to X. Lu), the Shandong Taishan Scholarship (X. Lu), the National Science Foundation of China (31301018 to X. Tan), the Key Research Program of the Chinese Academy of Sciences (ZDRW-ZS-2016-3 to X. Tan) and Qingdao Innovative Leading Talent (15-10-3-15-(31)-zch). The authors have no conflicts of interest or competing interests to declare.
References
- Du, W., Liang, F., Duan, Y., Tan, X. and Lu, X. (2013). Exploring the photosynthetic production capacity of sucrose by cyanobacteria. Metab Eng 19: 17-25.
- Fa, Y., Liang, W., Cui, H., Duan, Y., Yang, M., Gao, J. and Liu, H. (2015). Capillary ion chromatography-mass spectrometry for simultaneous determination of glucosylglycerol and sucrose in intracellular extracts of cyanobacteria. J Chromatogr B Analyt Technol Biomed Life Sci 1001: 169-173.
- Hagemann, M., Richter, S. and Mikkat, S. (1997). The ggtA gene encodes a subunit of the transport system for the osmoprotective compound glucosylglycerol in Synechocystis sp. strain PCC 6803. J Bacteriol 179(3): 714-720.
- Klähn, S. and Hagemann, M. (2011). Compatible solute biosynthesis in cyanobacteria. Environ Microbiol 13(3): 551-562.
- Motta, A., Romano, I. and Gambacorta, A. (2004). Rapid and sensitive NMR method for osmolyte determination. J Microbiol Methods 58(2): 289-294.
- Qiao, C., Duan, Y., Zhang, M., Hagemann, M., Luo, Q. and Lu, X. (2017). Effects of lowered and enhanced glycogen pools on salt-induced sucrose production in a sucrose-secreting strain of Synechococcus elongatus PCC 7942. Appl Environ Microbiol.
- Reed, R. H. and Stewart, W. D. P. (1985). Osmotic adjustment and organic solute accumulation in unicellular cyanobacteria from freshwater and marine habitats. Mar Biol 88(1):1-9.
- Rippka, R., Deruelles, J., Waterbury, J. B., Herdman, M. and Stanier, R. Y. (1979). Generic assignments, strain histories and properties of pure cultures of cyanobacteria. J Gen Microbiol 111(1):1-61.
- Slama, I., Abdelly, C., Bouchereau, A., Flowers, T. and Savoure, A. (2015). Diversity, distribution and roles of osmoprotective compounds accumulated in halophytes under abiotic stress. Ann Bot 115(3): 433-447.
- Song, K., Hagemann, M., Tan, X. and Lu, X. (2017). The response regulator Slr1588 regulates spsA but is not crucial for salt acclimation of Synechocystis sp. PCC 6803. Front Microbiol 8: 1176.
- Song, K., Tan, X., Liang, Y. and Lu, X. (2016). The potential of Synechococcus elongatus UTEX 2973 for sugar feedstock production. Appl Microbiol Biotechnol 100(18): 7865-7875.
- Tan, X., Yao, L., Gao, Q., Wang, W., Qi, F. and Lu, X. (2011). Photosynthesis driven conversion of carbon dioxide to fatty alcohols and hydrocarbons in cyanobacteria. Metab Eng 13(2): 169-176.
- First, dissolve 6 mg of glycerol, sucrose, glucose and glucosylglycero in 100 ml of Milli-Q water
Article Information
Copyright
© 2018 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Tan, X., Song, K., Qiao, C. and Lu, X. (2018). Determination of Intracellular Osmolytes in Cyanobacterial Cells. Bio-protocol 8(8): e2812. DOI: 10.21769/BioProtoc.2812.
Category
Microbiology > Microbial metabolism > Carbohydrate
Microbiology > Microbial physiology > Adaptation
Biochemistry > Carbohydrate > Polysaccharide
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









