- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Glycolate Oxidase Activity Assay in Plants
Published: Vol 2, Iss 20, Oct 20, 2012 DOI: 10.21769/BioProtoc.277 Views: 12707

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
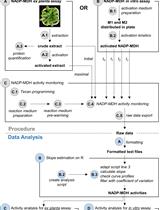
A Semi-throughput Procedure for Assaying Plant NADP-malate Dehydrogenase Activity Using a Plate Reader
Kevin Baudry and Emmanuelle Issakidis-Bourguet
Aug 20, 2023 1508 Views
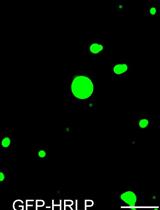
An in vitro Assay to Probe the Formation of Biomolecular Condensates
Yu Zhang and Shen Lisha
Sep 5, 2023 3261 Views
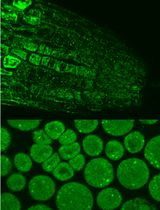
Immunofluorescence for Detection of TOR Kinase Activity In Situ in Photosynthetic Organisms
Ana P. Lando [...] Giselle M. A. Martínez-Noël
Dec 20, 2024 1859 Views
Abstract
Glycolate oxidase is located in the peroxisome and is involved in the photorespiratory cycle which recovers some of the carbon loss during photosynthesis. Glycolate oxidase converts glycolate to glyxoylate with the concomitant production of H2O2.In this assay, the H2O2 generated, in the presence of HRP, oxidizes O-dianisidine into a colored O-dianisidine radical cation that can be quantified spectrophotometrically. The amount of color produces is directly proportional to the glycolate oxidase activity.
Materials and Reagents
- Sucrose
- HEPES
- EDTA
- DTT
- L-cysteine
- MgCl2
- PVP
- BSA
- Complete, Mini, EDTA-free Protease Inhibitor Cocktail Tablets (F. Hoffmann-La Roche, catalog number: 04693159001 ).
- Bio-Rad Protein Assay (Bio-Rad Laboratories, catalog number: 500-0006 )
- Horseradish peroxidase(HRP) (Sigma-Aldrich, catalog number: P8375 )
- O-Dianisidine (Sigma-Aldrich, catalog number: D9143 )
- Sodium glycolate (Thermo Fisher Scientific, Acros Organics,catalog number: 351570250 )
- Potassium phosphate
- Triton X-100
- Protein extraction buffer (see Recipes)
- Glycolate oxidase assay buffer (see Recipes)
Equipment
- Microtiter plate reader (Infinite M200 Pro, Tecan)
- Microcentrifuge (AqquSpin Micro R) (Thermo Fisher Scientific)
- 96-well microtiter plate flat bottom(BD Biosciences, catalog number: 353075 )
- 2 ml-microcentrifuge tubes
Procedure
- Total Protein extraction from plant tissues
- Harvest tissue in liquid nitrogen. If not used immediately, keep at -80 °C until processing.
- Grind tissue in liquid nitrogen and weigh out 0.5 g of the ground tissues in empty falcon tube that has been pre-chilled in liquid nitrogen and used to tare the scale.
- Add 6 ml of ice cold protein extraction buffer to ground tissues on ice.
- Vortex at room temperature to mix thoroughly.
- Filter homogenized tissue through four layers of cheese cloth andtransfer filtrate (flow through) to 2 ml-microcentrifuge tubes, on ice.
- Centrifuge at 10,000 x g for 45 min at 4 °C and transfer supernatant to new tubes on ice.
- Use supernatant to estimate protein concentration and to measure glycolate oxidase activity.
- Harvest tissue in liquid nitrogen. If not used immediately, keep at -80 °C until processing.
- Protein estimation using Bradford microassay (160 μl)
- Prepare BSA standards ranging from 5 μg-25 μg/ml as follows.
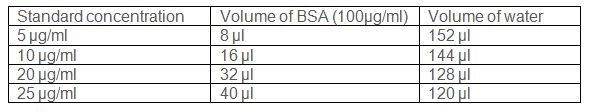

- Use 96 well microtiter plate to estimate protein content.
- Prepare blank by160 μl of water to one well in triplicates.
- Prepare test samplesby adding 2 μl of supernatant (from section 1) to 158 μl of water.
- Add 40 μl of Bradford Assay reagent to BSA standards, blank and test samples.
- Mix and incubate at room temperature for 5 min and read absorbance at 595 nm (A595) on plate reader spectrophotometer.
Note: If spectrophotometer does not include a software to generate standard curve to automatically estimate protein content, generate a BSA standard curve by plotting known protein concentration (X-axis) vs. Absorbance (in Y-axis). Protein concentration for a given unknown sample is estimated by plotting the A595 absorbance of the unknown (in the y-axis)and determining the intersection point with the BSA standard curve and then find the concentration associated with that particular point (in the x-axis). If using excel, after plotting concentration vsA595, obtain the trendline and use the equation for the line and the A595 of the unknown to resolve the unknown concentration.
- Prepare BSA standards ranging from 5 μg-25 μg/ml as follows.
- Glycolate oxidase activity assay
- Prepare blank by adding 10 μl of 0.1 M sodium phosphate buffer (pH 8.3), to 250 μl of glycolate oxidase assay bufferin 96 well microtiter plate.
- Prepare test samples by adding 10 μl of supernatant (from section 1) to 250 μl of glycolate oxidase assay buffer.
- Read blank and samples at 440 nm for 0 min and then at 20 min intervals for one hour or until saturation point reached, on plate reader spectrophotometer.
- Calculate the generation of O-dianisidine radical using the following formula:
(ΔA440nm/min test-ΔA440nm/min blank))/ (11.60) (0.04)
ΔA440nm/mintest =A440 (sample X) at saturation point - A440 (sample X) at 0 min
ΔA440nm/min blank = A440nm (blank) at saturation point - A440nm (blank) at 0 min
Where
11.60 =extinction coefficient for O-dianisidine. (Macheroux et al, 1991)
0.04= dilution factor (10 μl/250 μl)
To calculate specific activity, divide the value obtained in equation by the amount of protein present in the sample (converted to mg/ml).
- Prepare blank by adding 10 μl of 0.1 M sodium phosphate buffer (pH 8.3), to 250 μl of glycolate oxidase assay bufferin 96 well microtiter plate.
Recipes
- Protein extraction buffer
Working solution:
0.25 M sucrose
50 mM HEPES
3 mM EDTA
1 mM DTT
3.6 mM L-cysteine
0.1 mM MgCl2
0.6% PVP
10 tablets of complete, Mini, EDTA-free Protease Inhibitor Cocktail Tablets.
Protein extraction buffer (100 ml)
Use following stock solutions to make working solution:
In 80 ml of water add the following reagents: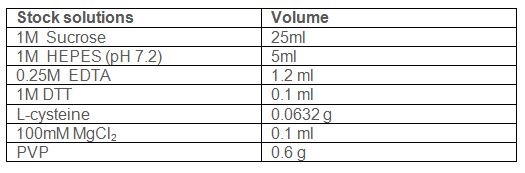
Add 10 tablets of complete, Mini, EDTA-free Protease Inhibitor Cocktail Tablets.
Mix well and adjust volume to 100 ml. - Glycolate oxidase assay buffer
Working solution:
10 μg ml-1 HRP
0.4 mM O-dianisidine
10 mM sodium glycolate
in 0.1 M potassium phosphate (pH 8.3)
Glycolate oxidase assay buffer (1 ml)
PrepareGlycolate oxidase assay buffer using following stock solutions:
Acknowledgments
This protocol has been adapted and modified to use in Arabidopsis from Macheroux et al. (1991). This work was supported by the Samuel Roberts Noble Foundation.
References
- Macheroux, P., Massey, V., Thiele, D. J. and Volokita, M. (1991). Expression of spinach glycolate oxidase in Saccharomyces cerevisiae: purification and characterization. Biochemistry 30(18): 4612-4619.
- Rojas, C. M., Senthil-Kumar, M., Wang, K., Ryu, C. M., Kaundal, A. and Mysore, K. S. (2012). Glycolate oxidase modulates reactive oxygen species-mediated signal transduction during nonhost resistance in Nicotiana benthamiana and Arabidopsis. Plant Cell 24(1): 336-352.
Article Information
Copyright
© 2012 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Kaundal, A., Rojas, C. M. and Mysore, K. S. (2012). Glycolate Oxidase Activity Assay in Plants. Bio-protocol 2(20): e277. DOI: 10.21769/BioProtoc.277.
Category
Plant Science > Plant biochemistry > Protein > Activity
Biochemistry > Protein > Activity
Biochemistry > Other compound > Reactive oxygen species
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










