- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Quantifying the Capacity of Phloem Loading in Leaf Disks with [14C]Sucrose
Published: Vol 7, Iss 24, Dec 20, 2017 DOI: 10.21769/BioProtoc.2658 Views: 8054
Reviewed by: John F. C. SteeleAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
A Quick Method to Quantify Iron in Arabidopsis Seedlings
Chandan Kumar Gautam [...] Wolfgang Schmidt
Mar 5, 2022 3960 Views

13CO2-labelling and Sampling in Algae for Flux Analysis of Photosynthetic and Central Carbon Metabolism
Or Geffen [...] Haim Treves
Sep 5, 2023 1985 Views
CAPS-Based SNP Genotyping for Nitrogen-Response Phenotypes in Maize Hybrids
Jannis Jacobs [...] Peter K. Lundquist
Dec 20, 2025 586 Views
Abstract
Phloem loading and transport of photoassimilate from photoautotrophic source leaves to heterotrophic sink organs are essential physiological processes that help the disparate organs of a plant function as a single, unified organism. We present three protocols we routinely use in combination with each other to assess (1) the relative rates of sucrose (Suc) loading into the phloem vascular system of mature leaves (this protocol), (2) the relative rates of carbon loading and transport through the phloem (Yadav et al., 2017a), and (3) the relative rates of carbon unloading into heterotrophic sink organs, specifically roots, after long-distance transport (Yadav et al., 2017b). We propose that conducting all three protocols on experimental and control plants provides a reliable comparison of whole-plant carbon partitioning, and minimizes ambiguities associated with a single protocol conducted in isolation (Dasgupta et al., 2014; Khadilkar et al., 2016). In this protocol, Arabidopsis leaf disks isolated from mature rosette leaves are infiltrated with a buffered solution containing [14C]Suc. Suc transporters (SUCs or SUTs) load Suc into the phloem and excess, unloaded Suc in the leaf disk is then washed away. Loading of labeled Suc into the veins is visualized by autoradiography of lyophilized leaf disks and quantified by scintillation counting. Results are expressed as disintegration per minute per unit of leaf disk fresh weight or area.
Keywords: ArabidopsisBackground
Transport of photoassimilates from source to sink organs is essential for normal growth and maintenance of whole plants. Phloem loading in leaves is the delivery of photoassimilate synthesized in mesophyll cells to the companion cells (CC) and sieve elements (SE) of the phloem vasculature system. Three distinct loading mechanisms are recognized. Two of these expend energy to accumulate high concentrations of sugar in the CC and SEs and generate a high hydrostatic pressure in source-leaf phloem. The first is apoplastic phloem loading, in which Suc (and/or sugar alcohols in some species) is loaded across the plasma membrane from the cell wall space (i.e., the apoplast) into the CCs at the expense of the proton motive force (Giaquinta, 1983). The second is polymer trapping, in which Suc diffuses into the phloem through specialized plasmodesmata and is converted to oligosaccharides that are too large to diffuse back out (Turgeon, 1996). The third mechanism is passive loading, in which the highest solute concentrations are in the mesophyll cells and plasmodesmata provide an open path for passive movement into the CCs and SEs (Rennie and Turgeon, 2009).
Due to the central role of phloem loading and transport to plant physiology and productivity, it is desirable to have reliable methods to identify and quantify the contents of the phloem and the rates of transport. In addition, phloem loading and transport are targets for biotechnology and metabolic engineering to enhance productivity (Ainsworth and Bush, 2011; Cao et al., 2013; Dasgupta et al., 2014; Zhang et al., 2015; Yadav et al., 2015). Quantitatively assessing the rates and capacity of phloem loading and transport in natural and engineered systems is difficult for several reasons. As examples, CCs and SEs are imbedded in surrounding tissues and are very narrow relative to surrounding non-phloem cells; the phloem is under high pressure and seals rapidly when damaged; collected phloem sap is usually contaminated with the content of other cells; because transport is a dynamic process, estimates of phloem content do not indicate rates of transport (Turgeon and Wolf, 2009; Dinant and Kehr, 2013; Tetyuk et al., 2013). C isotope 11C, 13C and 14C, in CO2 or labeled sugars, have provided, and continue to provide, critical information on phloem loading mechanisms (Sovonick et al., 1974; Turgeon and Gowan, 1990; Turgeon and Medville, 1998), as well as quantitative information on the rates of loading and transport (Thorpe and Minchin, 1988; Karve et al., 2015; Dersch et al., 2016).
Arabidopsis loads Suc from the apoplast with the Suc transporter (SUT) encoded by AtSUC2 (Truernit and Sauer, 1995; Gottwald et al., 2000; Srivastava et al., 2008). AtSUC2 and other transporters have been assessed in Saccharomyces cerevisiae and Xenopus laevis oocytes as heterologous systems to establish Michaelis Menten kinetic parameters (reviewed in [Ayre, 2011]). While valuable for comparing the activities and affinities among transporters, these do not inform on the activity of the transporters in planta, or on the overall impact on plant growth and productivity. As an example of this, SUTs from Solanaceae species with roughly the same kinetic properties as AtSUC2 did not rescue an Arabidopsis Atsuc2 mutant, while an AtSUC2 cDNA and ZmSUT1 cDNA from Zea mays did rescue the mutant (Dasgupta et al., 2014). In this protocol, and two that follow (Yadav et al., 2017a and 2017b), we detail the use of [14C]CO2 and [14C]Suc to quantitatively assess phloem loading and carbon transport in living explants and intact plants. These methods are used routinely in our laboratory, and recently contributed to our demonstration that plants over expressing certain SUTs in CCs showed increased phloem loading and transport, despite having stunted growth (Dasgupta et al., 2014) and, in a separate study, over expression of a H+-pumping pyrophosphatase enhanced Suc loading and transport without a corresponding increase in SUT expression levels (Khadilkar et al., 2016).
In the procedure described here (Figure 1), Arabidopsis leaf disks isolated from mature rosette leaves are vacuum infiltrated with a buffered solution containing [14C]Suc. SUTs load the Suc into CCs, and excess Suc is washed away. The disks are quickly frozen in powdered dry ice, lyophilized, and autoradiography is used to visualize loading into the veins. Rapid freezing and lyophilization prevent the highly soluble labeled sugar from diffusing away from the sites of loading. Scintillation counting then quantifies the loaded label. We generally pool four leaf disks into one replicate, and perform 6 biological replicates for each control and experimental system. This protocol was adapted for Arabidopsis (Dasgupta et al., 2014; Khadilkar et al., 2016) from procedures described by Turgeon and colleagues (Turgeon and Wimmers, 1988; Turgeon and Gowan, 1992; Turgeon and Medville, 1998; Haritatos et al., 2000; Goggin et al., 2001), and can be further adapted for other plants. Typical autoradiography results for Arabidopsis, tobacco, and wheat are shown in Figure 2; representative quantitative results of loading in different transgenic Arabidopsis plants are published (Dasgupta et al., 2014; Khadilkar et al., 2016).
Materials and Reagents
- Potting containers (The HC Companies, catalog number: IJT06060 ;Jumbo Insert)
- Potting mixture (Sun Gro Horticulture, catalog number: Fafard 3B Mix ; or similar)
- 24-well culture plates (Greiner Bio-One International, catalog number: 662160 )
- 5 ml transport vials (Stockwell Scientific, catalog number: 3205 )
- Nylon window screen (available at most hardware stores)
- Cyanoacrylate (Super Glue, or equivalent)
- Razor blades (Double-edged PERSONNA) (Electron Microscopy Sciences, catalog number: 72000 )
- Petri dishes (100 x 25 mm) (Fisher Scientific, catalog number: FB0875711 )
- Filter paper (Fisher Scientific, catalog number: 09-795C )
- Glass beads 4 mm (Water Stern, catalog number: 100E )
- Dow Corning high vacuum grease
- Thick, smooth paper (e.g., Bristol board or manila folder)
- Scintillation vials (Fisher Scientific, catalog number: 12383317 )
- Wax paper (Reynolds CUT-RITE)
- Aluminum foil envelopes (70 x 70 mm–homemade from standard aluminum foil)
- Autoradiography film (Kodak BioMax MR Film, Eastman Kodak, catalog number: 870 1302 )
- Plant material (e.g., Arabidopsis thaliana Col-0, control and experimental material), 10 to 12 healthy plants for each treatment
- Developer and fixer solutions (Eastman Kodak, catalog numbers: 190 0984 and 190 2485 )
- Sodium hypochlorite (NaClO) (Commercial bleach)
- Ecolume scintillation fluid (MP Biomedicals, catalog number: 0188247004 )
- 95% ethanol (Pharmco-AAPER, catalog number: 111000200 )
- 2(N-morpholino) ethane-sulfonic acid [MES] (Fisher Scientific, catalog number: BP300-100 )
- Calcium chloride dihydrate (CaCl2·2H2O) (Sigma-Aldrich, catalog number: 223506 )
- Potassium hydroxide (KOH) (Merck, catalog number: PX1480-1 )
- Sucrose (Fisher Scientific, catalog number: BP220-212 )
- [14C]Sucrose (100 µCi ml-1, > 350 mCi mmol-1) (MP Biomedicals, catalog number: 011113791 )
- Powdered dry ice (preferred) or liquid nitrogen
- Drierite desiccant (W A Hammond Drierite, catalog number: 13001 )
- MES/CaCl2 buffer (see Recipes)
- 1 mM Suc in MES/CaCl2 buffer (see Recipes)
- [14C]Suc in 1 mM Suc and MES/CaCl2 buffer (see Recipes)
Equipment
- Personal safety equipment: lab coat, nitrile gloves (or similar), and eye protection
- Growth chamber for Arabidopsis, 12 h light/12 h dark at ~100 µmol photons m-2 sec-1
- Cork borer (Carolina Biology, catalog number: 712202 )
- Cover glass forceps or similar (Fine Science Tools, catalog number: 11073-10 )
- Fine scissors (Fisher Scientific, catalog number: 08-951-20 )
- Vacuum bell jar (Kimble, catalog number: 31200-150 )
- Vacuum pump (Franklin Electric, catalog number: 1112585400 )
- Rotary shaker (Orbital Shaker Variable) (BioExpress, GeneMate, catalog number: S-3200-LS )
- Balance (METTLER TOLEDO, model: AE100 )
- Geiger counter (Ludlum Measurements, model: Model 3 )
- Lyophilizer (The VIRTIS company, catalog number: 10-030 )
- Mesh basket for lyophilizer chamber (homemade from metal window screen)
- Heavy duty bench vise (Kobalt 4” Vise)
- Two flat-surface, metal plates (~15 x 3 cm)
- Autoradiography cassettes (Exposure Cassette Kodak®) (Sigma-Aldrich, catalog number: E9010 )
- Autoradiography developing trays (Commercial plastic trays ~35 x 25 x 8 cm)
- Scintillation counter (Beckman Counter, model: LS 6000IC )
Procedure
- Working with any radionuclide requires special consideration and approval from the appropriate institutional office. Clearance can take a long time (months or more) and the application process should be started early, or collaborations should be established with groups that already have approvals in place.
- Grow Arabidopsis plants under short-day conditions with potting mix (Fafard 3B mix, or similar) to achieve well separated, large, and healthy rosettes; larger shade leaves are easier to work with than small sun leaves: 12 h light/12 h dark at ~100 µmol photons m-2 sec-1 works well. Select source leaves for labeling from each rosette that are fully expanded but not shaded or beginning to senesce (e.g., leaves 8 through 12 on ~25-d old rosettes) (Figure 1A). We generally use 32 leaf disks per treatment (8 pools of 4 disks, see below), and prepare 10 to 12 plants for each treatment to ensure sufficient healthy material.
- This procedure requires numerous (3 or 4) washing steps, which are made easier with small baskets constructed to fit inside the wells of a 24-well culture plate (shown in Figure 1C). We constructed baskets from polypropylene vials (5 ml transport vials, Stockwell Scientific). The top and bottom of each vial was cut off, leaving ¾ inches of open cylinder. Nylon window screen was attached to the bottom opening with cyanoacrylate (Super Glue, or equivalent) and excess screen trimmed away with a razor blade. These baskets maximize use of the space in the culture plate wells while still moving freely in and out of the wells during washing. Other forms of tubing will undoubtedly work, but investigators are cautioned that standard commercial tubing with 5/8 inch outer diameter and 1/2 inch inner diameter will likely not move freely in the wells.
- Cut the petioles near the stem and transfer the leaves to a Petri dish lined with three layers of filter paper and MES/CaCl2 buffer (20 mM MES, 2 mM CaCl2, pH 5.5 with KOH) to a depth of a few mm to ensure leaf disks are completely submerged. Under buffer, cut out a leaf disk with a #5 cork bore (~8 mm diameter, or smaller, depending on the size of the source leaves) (Figure 1B). Transfer the leaf disks abaxial side down into a basket in the well of a 24-well microtiter plate containing 1 ml MES/CaCl2 buffer (Figure 1D); pool 4 disks in each basket in each well and gently cover with 4 mm glass beads to keep them submerged (Figure 1E). We generally use 6 (minimum) to 8 (preferred) replicates of 4 pooled disks, so 24 to 32 disks are required for each treatment. The disks are randomized among the pools.
Note: All solutions should be at temperatures approximating leaf temperature. Cold solutions, such as from a refrigerator, will inhibit phloem loading. - Once all the baskets in the wells are prepared, lift the baskets (with leaf disks and glass beads) out of the culture plates and transfer to the wells of fresh plate pre-filled with 0.5 ml MES/CaCl2 buffer supplemented with 1 mM Suc solution (1 mM unlabeled Suc supplemented with 0.81 µCi (3 x 104 Bq) ml-1 [14C]Suc) (Figure 1F). The leaf disks should be submerged owing to the glass beads; more solution can be added to each well if necessary.
- Vacuum infiltrate the MES/CaCl2 with [14C]Suc solution into the leaf disks. In a bell jar and with a vacuum pump, pull a strong vacuum on the microtiter plate until bubbles no longer emerge from the leaf disks and then release the vacuum (Figure 1G). This pulls air out of the leaf air space and replaces it with labeling [14C]Suc solution. Imbibed sections of the disks leaves will look darker than non-imbibed sections; repeat the vacuum infiltration until the disks are completely imbibed and saturated.
Note: It is important to keep the disks submerged in MES/CaCl2 with [14C]Suc solution for effective infiltration, otherwise, air will be pulled back into the disks when the vacuum is released, rather than solution. - Cover the culture plate and incubate the vacuum-infiltrated discs for 20 min with gentle shaking on rotating platform (shaker) at room temperature.
- Lift the baskets (with labeled disks and glass beads), letting the MES/CaCl2 with [14C]Suc solution drain, and transfer the baskets to fresh culture plates pre-filled with 1 ml MES/CaCl2 buffer (Figure 1H). Incubate with gentle rotation on a rotary platform for 10 min.
- Repeat the above wash step another two times. Treat all wash solutions as a radioactive, [14C] containing liquid waste as it contains a significant amount of [14C]Suc. Process wells as quickly as practical to minimize deviations in incubation times between samples.
Note: The MES/CaCl2 with [14C]Suc solution can be reused, if desired, since only a small portion of the [14C]Suc is absorbed into the leaf disks. However, this increases the risk of contamination by enzymes such as invertase and microorganisms. We recommend filter sterilizing the solution and freezing promptly, with appropriate labels and storage for isotopes, if it is to be reused. - Blot the leaf disks on absorbent filter paper (Figure 1I) and place them in perforated aluminum foil envelopes (70 x 70 mm), prepared and labeled in advance. To make envelopes, fold a piece of aluminum foil in half and then fold in two sides to leave one side open; perforate the foil with a tack. We put the four disks representing one repetition into a single envelope. Place each envelope immediately on powdered dry ice and cover the envelope with more powered dry ice, repeat for each envelope, keep them frozen until all samples are processed.
Note: Liquid nitrogen can also be used, but layering the envelopes in powdered dry ice is preferred since it causes less cracking than submerging in liquid nitrogen: intact leaf disks are much easier to work with during autoradiography. - Lyophilize the frozen disks in a lyophilizer for 48 h. It is imperative that they remain frozen to prevent movement of the highly soluble [14C]Suc (Figure 1J).
Note: A lyophilizer with a sample chamber cooled to well below the freezing point (~-30 °C) of the tissue works best; lyophilizers with external sample vessels attached via a manifold work poorly because the disks thaw sufficiently to allow diffusion of the label. In Figure 1J, the aluminum foil envelopes are placed inside a homemade metal mesh basket (made from metal window screen) that fits snuggly within the coiling coils of our lyophilizer. The coiling coils themselves are the coldest part of the system, so water sublimates from the leaf disks and condenses on the coils, while the entire chamber stays well below the freezing point of the samples. - After lyophilizing, transfer the envelopes with leaf disks to a bell jar with Drierite desiccant: it is important to prevent condensation and exposure to humid conditions to avert diffusion of the [14C]Suc.
- Autoradiography and scintillation counting
- Visualizing phloem loading into leaf veins by autoradiography
- Visualizing phloem loading by autoradiography is based on the veins accumulating [14C]Suc from the apoplast while it is washed out of the areoles. Because 14C emits weak β particles and the signal disperses with distance from the source, the best autoradiograms are obtained with flat, thin leaf disks placed directly on single emulsion, high-resolution film, such as Kodak BioMax MR Film.
- Press the leaf disks flat between two metal plates (15 x 3 cm) in a heavy duty bench vise. On top of one metal plate (Figure 1K), place a piece of paperboard (Bristol paper or manila folder paper) and a piece of wax paper (~12.5 x 2.5 cm). Arrange the lyophilized leaf disks abaxial side up on the wax paper, keeping track of which disks belong to each repetition by labeling either the wax paper or the underlying paperboard. Cover the disks with another piece of paperboard and then the second metal plate. Insert the ‘sandwich’ into a heavy duty bench vise and compress as flat as possible (Figure 1L). Disassemble the sandwich but do not separate the disks from the wax paper; after compression, the wax paper provides a slightly tacky surface to help keep the disks in place during autoradiography. Repeat until all the disks are pressed flat.
- Arrange the bottom pieces of paperboard, wax paper and loosely adhered leaf disks (abaxial side up) in an autoradiography cassette (Figure 1M). In a darkroom licensed for work with isotopes, and under safe lights, place a piece of Kodak BioMax MR Film (or similar) with the emulsion side down so that it contacts the leaf disks.
- Depending on the strength of the [14C]Suc labeling, expose the autoradiography film to the leaf disks for 24 to 48 h. We generally start with a 48 h exposure, and after developing we make a decision if a second exposure, for a shorter or longer period of time, is warranted.
- In a dark room, remove the autoradiography film from the cassette taking care that the leaf disks stay adhered to the wax paper. Keeping the disks organized is important if a second exposure will be done. Develop the autography film by hand with freshly prepared developer and fixer (Kodak GBX) according to the manufacturer’s instructions. In brief, place the film in a tray with developer solution and incubate at room temperature with gentle shaking for 5 min. Transfer the film to a tray with water for 30 sec and then transfer to a tray with fixer solution for 5 min. Rinse with running water for 15 min and hang the film to drip and air dry.
- A stereo microscope equipped with a digital camera works well for photographing the autoradiographs.
- Visualizing phloem loading by autoradiography is based on the veins accumulating [14C]Suc from the apoplast while it is washed out of the areoles. Because 14C emits weak β particles and the signal disperses with distance from the source, the best autoradiograms are obtained with flat, thin leaf disks placed directly on single emulsion, high-resolution film, such as Kodak BioMax MR Film.
- Quantifying phloem loading into leaf veins by scintillation counting
- After autoradiography, remove the leaf disks from the wax paper and place the disks from each replicate in a scintillation vial (Figure 1N).
- To extract the pigments and solutes, add 500 µl 80% ethanol and shake gently for 20 min. Add 500 µl commercial bleach and shake gently for 20 min to destroy the pigments.
- Add 5 ml of biodegradable scintillation cocktail suitable for aqueous solutions. Mix thoroughly by shaking the vial to ensure the scintillation cocktail permeates the leaf disk and creates a monophasic solution. Include a negative control vial containing ethanol, bleach and scintillation cocktail, but without leaf disks.
- Perform scintillation counting with a program suitable for 14C with a scintillation counter.
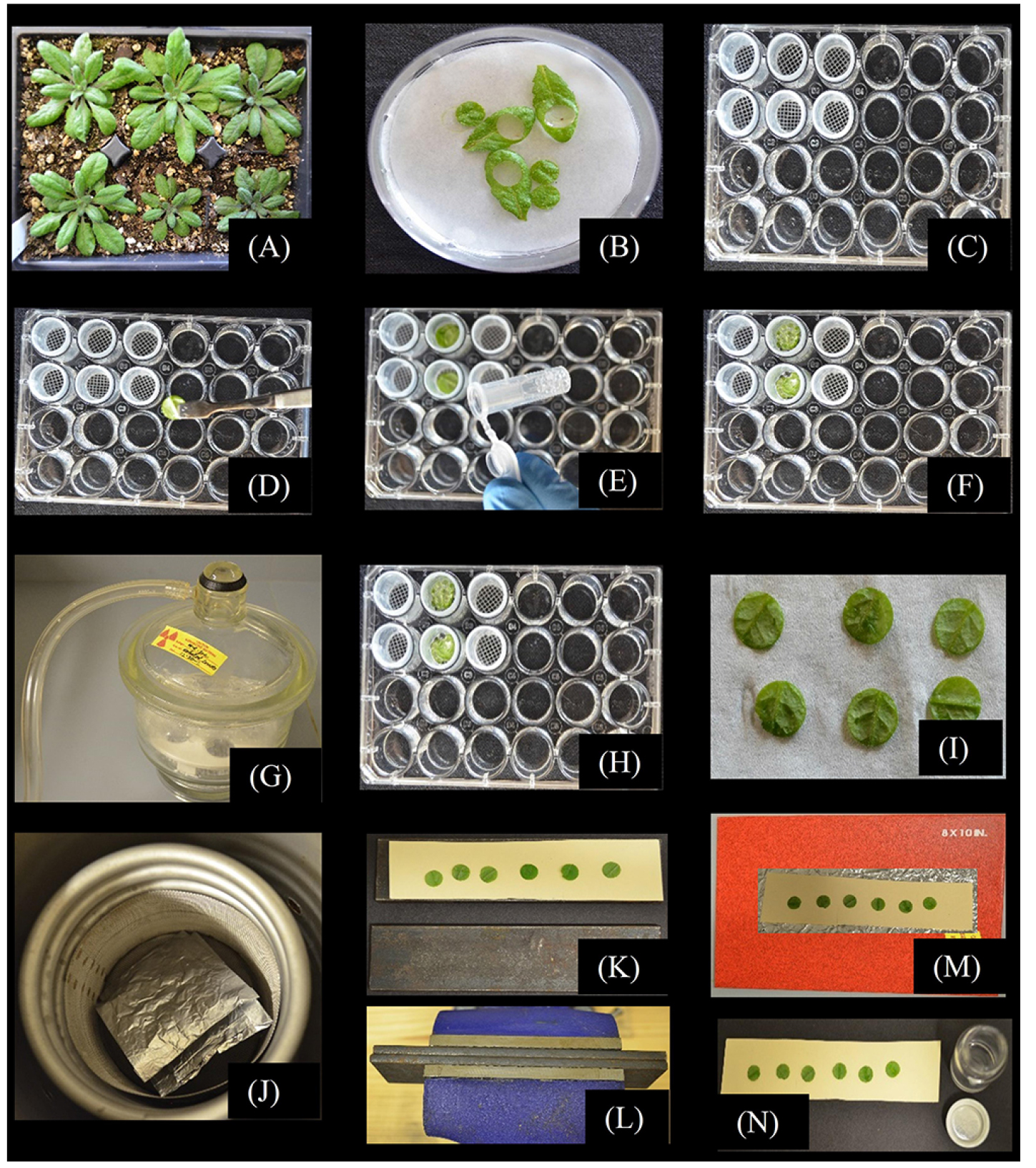

Figure 1. Experimental procedure to study phloem loading using [14C]Suc in Arabidopsis leaf disks. A. Grow Arabidopsis thaliana Col-0 control and experimental plants on potting mixture under conditions suitable to the experiment. B. Cut healthy mature leaves at the petiole base, transfer leaves to a Petri dish lined with filter paper and while the leaves are submerged in MES/CaCl2 buffer, use a #5 mm cork borer to isolate leaf disks. C. Have prepared 24-well microtiter plates filled with 1 ml MES/CaCl2 buffer in each well, and mesh-bottom baskets for convenient washing in later steps. D. Pool 4 disks in each well and (E) gently cover with glass beads to submerge the leaf disks. F. Once all the baskets with leaf disks and glass beads are prepared, transfer the baskets to the wells of a fresh plate pre-filled with MES/CaCl2 buffer supplemented with 1 mM Suc solution (1 mM unlabeled Suc supplemented with 0.81 µCi (3 x 104 Bq) ml-1 [14C]Suc). G. In a bell jar, vacuum infiltrate the MES/CaCl2/[14C]Suc solution into the leaf disks making sure to imbibe the air-space with solution. H. Incubate the disks for 20 min with gentle shaking on a rotary platform, and wash disks three times with MES/CaCl2 solution without Suc. I. Blot the leaf disks on absorbent paper (I) and place them into premade and prelabeled, perforated aluminum foil envelopes. Layer the envelopes in powdered dry ice (preferred method) or submerge in liquid nitrogen; the perforations in the aluminum foil envelopes allow liquid N2 to enter and permit N2 gas to escape. J. Lyophilize the discs for 48 h such that they remain frozen while drying (shown are aluminum foil envelopes inside a metal mesh basket made to fit inside the coiling coils of the lyophilizer we use; temperature is maintained at ~-30 °C). After lyophilizing, move the aluminum foil envelopes to a desiccator to warm to room temperature without condensation forming. K. Carefully remove the disk from the envelopes and arrange, abaxial side up, on wax paper on Bristol paper on a flat-surface metal plate. Put a second sheet of Bristol paper (without wax paper) over the leaf disk and a second, flat surface metal plate. L. Transfer the sandwich to a heavy duty bench vise and press the leaf disks as flat as possible. M. The leaf disks will loosely adhere to the wax paper, and can be transferred to an autoradiography cassette for exposure to single emulsion, high-resolution autoradiography film. N. After satisfactory autoradiography, remove the leaf disks from the wax paper and quantify 14C label by scintillation counting.
Data analysis
Autoradiography is a qualitative visual assessment of loading whereas scintillation counting provides quantitative data. Representative autoradiograms of Arabidopsis, tobacco, and wheat are shown in Figure 2. The results of scintillation counting are expressed as disintegrations per minute (dpm) per unit of leaf area (mm2) or as a percent relative to controls: (experimental dpm/control dpm) x 100 (Khadilkar et al., 2016). For quantitative analysis, the mean is calculated for each of the 8 replicates of 4 pooled disks. Statistically significant differences in loading among treatments are determined by standard calculations such as Student’s t-test or analysis of variance (ANOVA) with post-hoc analysis.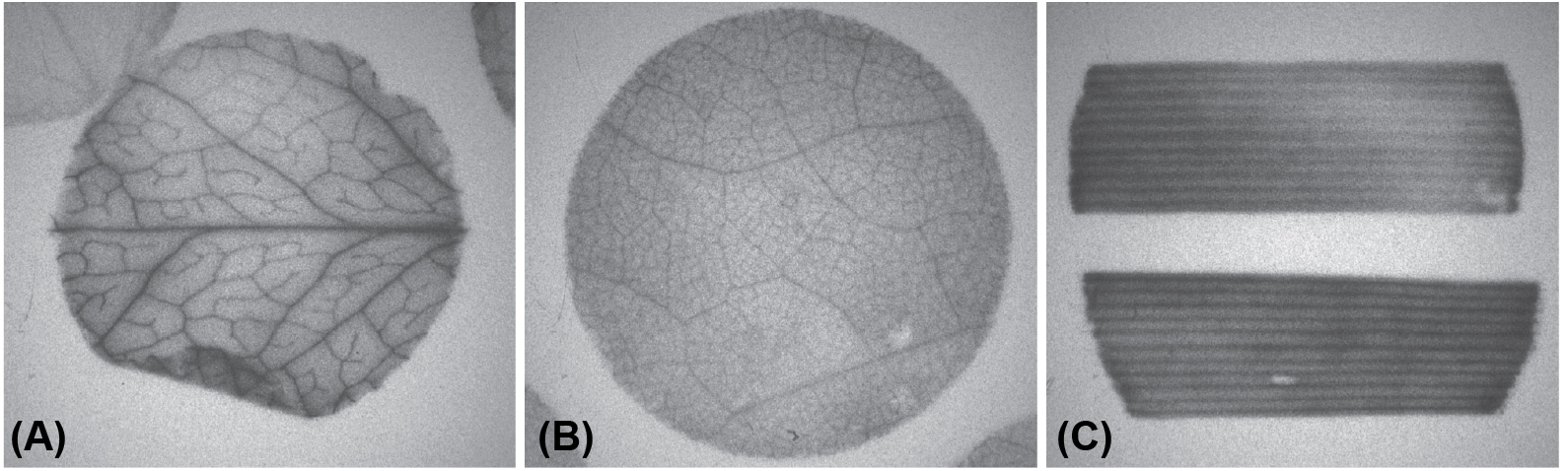
Figure 2. Representative autoradiography of (A) Arabidopsis, (B) tobacco, and (C) wheat leaf segments prepared with this protocol
Recipes
- MES/CaCl2 buffer (500 ml)
MES anhydrous 1.9524 g (FW 195.24) (20 mM)
CaCl2·2H2O 0.147 g (FW 147.01) (2 mM)
Adjust pH 5.5 with KOH (5 N) - 1 mM Suc in MES/CaCl2 buffer (100 ml)
Sucrose 0.034 g (FW 342.30) (1 mM) in 100 ml MES/CaCl2 buffer, or dilute appropriately from a more concentrated Suc stock - [14C]Suc in 1 mM Suc and MES/CaCl2 buffer (10 ml, make fresh, calculate how much will be needed for the specific experiment and prepare ~10% extra)
Commercial [14C]Suc stock, 100 µCi ml-1, > 350 mCi mmol-1
Add 0.081 ml (3 x 104 Bq) of commercial stock to 9.92 ml of 1 mM Suc in MES/CaCl2 buffer
Note: [14C]Suc specific activity may vary with different batches and suppliers. 0.081 ml of the stock described contributes 0.023 µmoles of Suc per 10 ml of 1 mM Suc and MES/CaCl2 buffer and increases the final Suc concentration to 1.002 mM, which we consider negligible.
(0.081 ml) x (100 µCi ml-1)/(350 mCi mmol-1) = 0.023 µmoles
0.023 µmoles/10 ml = 0.0023 mM
Acknowledgments
This protocol is based on methods published in Dasgupta et al. (2014) and Khadilkar et al. (2016). We thank John Evers for providing the plants contributing to Figure 2. Work on phloem loading and long distance transport in B.G. Ayre’s laboratory is/was supported by the National Science Foundation 0344088, 0922546, 1121819, and 1558012. The authors report no conflicts of interest or competing interests.
References
- Ainsworth,E. A. andBush, D. R. (2011). Carbohydrate export from theleaf: a highly regulated process and target to enhance photosynthesis andproductivity. Plant Physiol 155(1):64-69.
- Ayre,B. G. (2011). Membrane-transport systems forsucrose in relation to whole-plant carbonpartitioning. Mol Plant 4(3):377-394.
- Cao,T., Lahiri, I., Singh, V., Louis, J., Shah, J. and Ayre, B. G. (2013). Metabolicengineering of raffinose-family oligosaccharides in the phloem revealsalterations in carbon partitioning and enhances resistance to green peach aphid. FrontPlant Sci 4: 263.
- Dasgupta,K., Khadilkar, A. S., Sulpice, R., Pant, B., Scheible, W. R., Fisahn, J.,Stitt, M. and Ayre, B. G. (2014). Expression of sucrosetransporter cDNAs specifically in companion cells enhances phloem loading andlong-distance transport of sucrose but leads to an inhibition of growth and theperception of a phosphate limitation. PlantPhysiol 165(2): 715-731.
- Dersch,L. M., Beckers, V., Rasch, D., Melzer, G., Bolten, C., Kiep, K., Becker, H.,Blasing, O. E., Fuchs, R., Ehrhardt, T. and Wittmann, C. (2016). Novel approach for high-throughput metabolicscreening of whole plants by stable isotopes. PlantPhysiol 171(1): 25-41.
- Dinant,S. and Kehr, J. (2013). Sampling and analysis ofphloem sap. Methods Mol Biol 953:185-194.
- Giaquinta,R. T. (1983). Phloem loading of sucrose. AnnuRev Plant Physiol Plant Mol Biol 34: 347-387.
- Goggin,F. L., Medville, R. and Turgeon, R.(2001). Phloem loading in the tuliptree. Mechanisms and evolutionary implications. PlantPhysiol 125(2): 891-899.
- Gottwald,J. R., Krysan, P. J., Young, J. C., Evert, R. F. and Sussman, M. R. (2000). Geneticevidence for the in planta role of phloem-specific plasma membrane sucrosetransporters. Proc Natl Acad Sci U S A 97(25): 13979-13984.
- Haritatos,E., Ayre, B. G. and Turgeon, R. (2000). Identification of phloeminvolved in assimilate loading in leavesby the activity of the galactinol synthase promoter. PlantPhysiol 123(3): 929-937.
- Karve,A. A., Alexoff, D., Kim, D., Schueller, M. J., Ferrieri, R. A. and Babst, B. A.(2015). In vivo quantitative imagingof photoassimilate transport dynamics and allocation in large plants using acommercial positron emission tomography (PET) scanner. BMCPlant Biol 15: 273.
- Khadilkar,A. S., Yadav, U. P., Salazar, C., Shulaev, V., Paez-Valencia, J., Pizzio, G.A., Gaxiola, R. A. and Ayre, B. G. (2016). Constitutive and companioncell-specific overexpression of AVP1, encoding a proton-pumpingpyrophosphatase, enhances biomass accumulation, phloem loading, andlong-distance transport. Plant Physiol 170(1): 401-414.
- Rennie,E. A. and Turgeon, R. (2009). Acomprehensive picture of phloem loading strategies. ProcNatl Acad Sci U S A 106(33): 14162-14167.
- Sovonick,S. A., Geiger, D. R. and Fellows, R. J. (1974). Evidence for active Phloemloading in the minor veins of sugar beet. Plant Physiol 54(6): 886-891.
- Srivastava,A. C., Ganesan, S., Ismail, I. O. and Ayre, B. G. (2008). Functionalcharacterization of the Arabidopsis AtSUC2 Sucrose/H+ symporter by tissue-specific complementationreveals an essential role in phloem loading but not in long-distance transport. PlantPhysiol 148(1): 200-211.
- Tetyuk,O., Benning, U. F. and Hoffmann-Benning, S. (2013). Collection and analysis ofArabidopsis phloem exudates using the EDTA-facilitated method. JVis Exp(80): e51111.
- Thorpe,M. R. and Minchin, P. E. H. (1988). Phloemloading and transport of endogenously or exogenously labeled photoassimilate inbean, beet, maize and cucurbit. J Exp Bot 39: 1709-1721.
- Truernit,E. and Sauer, N. (1995). The promoter of the Arabidopsis thaliana SUC2 sucrose-H+ symporter gene directs expression of beta-glucuronidase to the phloem: evidencefor phloem loading and unloading by SUC2. Planta 196(3): 564-570.
- Turgeon,R. (1996). Phloemloading and plasmodesmata. Trends Plant Sci 1: 418-423.
- Turgeon,R. and Gowan, E. (1990). Phloem loading in coleus blumei in the absence of carrier-mediateduptake of export sugar from the apoplast. PlantPhysiol 94(3): 1244-1249.
- Turgeon,R. and Gowan, E. (1992). Sugar synthesis and phloemloading in Coleus blumei leaves. Planta 187(3): 388-394.
- Turgeon,R. and Medville, R. (1998). The absence ofphloem loading in willow leaves. ProcNatl Acad Sci U S A 95(20): 12055-12060.
- Turgeon,R. and Wimmers, L. E. (1988). Differentpatternsof vein loading of exogenous C-14 cucrosein leaves of Pisum sativum and Coleus blumei. Plant Physiol 87: 179-182.
- Turgeon,R. and Wolf, S. (2009). Phloem transport: cellularpathways and molecular trafficking. AnnuRev Plant Biol 60: 207-221.
- Yadav,U. P., Ayre, B. G. and Bush, D. R. (2015). Transgenic approaches toaltering carbon and nitrogen partitioning in whole plants: assessing thepotential to improve crop yields and nutritional quality. FrontPlant Sci 6: 275.
- Yadav,U. P., Khadilkar, A. S., Shaikh, M. A., Turgeon, R. and Ayre, B. G. (2017a). Assessing rates of long-distancecarbon transport in Arabidopsis by collectingphloem exudations into EDTA solutions after photosynthetic labeling with [14C]CO2. Bio Protoc 7(24): e2656.
- Yadav,U. P., Khadilkar, A. S., Shaikh, M. A., Turgeon, R. and Ayre, B. G. (2017b). Assessing long-distancetransport from photosynthetic source leaves to heterotrophic sink organs with [14C]CO2. Bio Protoc 7(24): e2657.
- Zhang,L., Garneau, M. G., Majumdar, R., Grant, J. and Tegeder, M. (2015). Improvement of pea biomass and seed productivity by simultaneous increase of phloem and embryoloading with amino acids. Plant J 81(1):134-146.
Article Information
Copyright
© 2017 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Yadav, U. P., Khadilkar, A. S., Shaikh, M. A., Turgeon, R. and Ayre, B. G. (2017). Quantifying the Capacity of Phloem Loading in Leaf Disks with [14C]Sucrose. Bio-protocol 7(24): e2658. DOI: 10.21769/BioProtoc.2658.
- Khadilkar, A. S., Yadav, U. P., Salazar, C., Shulaev, V., Paez-Valencia, J., Pizzio, G. A., Gaxiola, R. A. and Ayre, B. G. (2016). Constitutive and companion cell-specific overexpression of AVP1, encoding a proton-pumping pyrophosphatase, enhances biomass accumulation, phloem loading, and long-distance transport. Plant Physiol 170(1): 401-414.
Category
Plant Science > Plant physiology > Nutrition
Plant Science > Plant physiology > Photosynthesis
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










