- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
In vivo Leukemogenesis Model Using Retrovirus Transduction
Published: Vol 7, Iss 23, Dec 5, 2017 DOI: 10.21769/BioProtoc.2627 Views: 9636
Reviewed by: Jia LiXi FengXiaoyi Zheng

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Measurement of Bone Metastatic Tumor Growth by a Tibial Tumorigenesis Assay
Baotong Zhang [...] Jin-Tang Dong
Nov 20, 2021 4547 Views
Generation and Maintenance of Patient-Derived Endometrial Cancer Organoids
Mali Barbi [...] Semir Beyaz
Oct 20, 2024 2107 Views
Temporally and Spatially Controlled Age-Related Prostate Cancer Model in Mice
Sen Liu [...] Qiuyang Zhang
Jan 5, 2025 2744 Views
Abstract
Various genetic alterations such as chromosomal translocation cause leukemia. For examples, gene rearrangements of the mixed-lineage leukemia (MLL) gene generate MLL fusion genes, whose products are potent oncogenic drivers in acute leukemia. To better understand the mechanism of disease onset, several murine leukemia models using retroviral gene transduction, xenograft, or Cre-mediated chromosomal translocation have been developed over the past twenty years. Particularly, a retroviral gene transduction-mediated murine leukemia model has been frequently used in the leukemia research field. Here, we describe the detailed protocol for this model.
Keywords: Mixed lineage leukemiaBackground
Gene rearrangements generate mixed-lineage leukemia (MLL) fusion genes, which cause highly aggressive acute leukemia. MLL-rearrangements are often associated with few additional genetic alterations and poor clinical outcomes (Andersson et al., 2015). Wild-type MLL enhances and maintains the expression of a subset of genes, including homeobox (Hox) genes, to stimulate the expansion of immature progenitors (Jude et al., 2007). The expression of Hoxa9 and Meis1 is highest in the immature progenitor/stem cell fraction, but gradually declines as cells differentiate, and eventually diminishes in terminally-differentiated cell fractions (Somervaille and Cleary, 2006; Yokoyama et al., 2013). The MLL fusion protein constitutively up-regulates the expression of target genes, including Hoxa9 and Meis1, to immortalize immature progenitor cells and cause leukemia in vivo (Ayton and Cleary, 2003; Lavau et al., 1997). To date, more than 130 different MLL-rearrangements have been identified (Meyer et al., 2017). Two-thirds of MLL-rearranged leukemia cases are caused by fusion with a gene that is part of the AF4 family-ENL family-P-TEFb (AEP) complex (Yokoyama et al., 2010). The MLL fusion proteins constitutively form an MLL/AEP hybrid complex on the target chromatin (Okuda et al., 2014; Yokoyama et al., 2010), which further associates with the SL1 complex to activate RNA polymerase II-dependent transcription (Okuda et al., 2015 and 2016). AEP-mediated transactivation of MLL target genes transformed myeloid progenitors ex vivo, but did not cause leukemia in vivo, which suggested that other function is additionally required for in vivo leukemogenesis (Okuda et al., 2017). Recently, we showed that the ability to recruit the DOT1L complex is necessary to cause leukemia in vivo in addition to the ability to recruit AEP using in vivo leukemogenesis model. Thus, the combinatorial use of the in vivo leukemogenesis model and myeloid progenitor transformation assay is necessary to dissect the functional properties of oncogenes. In this protocol, we describe the in vivo leukemogenesis model using retroviral transduction in detail.
Materials and Reagents
- Pipette tips
- Tissue culture 10-cm dish (Greiner Bio One International, catalog number: 664160 )
- Collagen-coated tissue culture 10-cm dishes (Corning, catalog number: 354450 )
- 15-ml conical centrifuge tubes (Thermo Fisher Scientific, Thermo ScientificTM, catalog number: 339650 )
- Terumo syringe® 10-ml (Terumo, catalog number: SS-10ESZ )
- Terumo needle 21 G x1 ½” (Terumo, catalog number: NN-2138S )
- MS column (Miltenyi Biotec, catalog number: 130-042-201 )
- Pre-separation filter (Miltenyi Biotec, catalog number: 130-041-407 )
- Tissue culture 48-well plate (Thermo Fisher Scientific, Thermo ScientificTM, catalog number: 150687 )
- Millex-GV 0.45-µm PVDF 33 mm sterile syringe filter (Merck, catalog number: SLHV033RB )
- Terumo® tuberculin syringe 26 G x ½” (Terumo, catalog number: SS-01T2613S )
- Terumo Myjector® insulin syringe 29 G x ½” (Terumo, catalog number: SS-05M2913 )
- Terumo syringe® 1-ml (Terumo, catalog number: SS-01T )
- Terumo needle 18 G x1 ½” (Terumo, catalog number: NN-1838S )
- Tissue culture 12-well plates (Thermo Fisher Scientific, Thermo ScientificTM, catalog number: 150628 )
- Bottle-top filter 0.2-µm PVDF (Corning, catalog number: 431098 )
- Five-week-old female C57BL/6JJcl mice (donor mice) (CLEA Japan, Tokyo, Japan)
- Seven-week-old female C57BL/6JJcl mice (recipient mice) (CLEA Japan)
- Platinum-E packaging (PLAT-E) cell line (Morita et al., 2000) (gifted from Dr. Toshio Kitamura, or Cell Biolabs, catalog number: RV-101 )
- WEHI-3 (ATCC, catalog number: TIB-68 )
- pMSCV-neo vector (Takara Bio, ClontechTM, catalog number: 634401 )
- pMSCV-neo-MLL-ENL (request to Akihiko Yokoyama: ayokoyam@ncc-tmc.jp)
- Trypsin-EDTA solution (NACALAI TESQUE, catalog number: 32778-34 )
- LipofectamineTM 2000 transfection reagent (Thermo Fisher Scientific, InvitrogenTM, catalog number: 11668019 )
- Opti-MEMTM I medium (Thermo Fisher Scientific, InvitrogenTM, catalog number: 31985070 )
- 70 % ethanol
- CD117 microbeads, mouse (Miltenyi Biotec, catalog number: 130-091-224 )
- 2.5% Baytril solution (Bayer, BaytrilTM, https://www.baytril.com/en/farm-animals/product/oral/)
- Polybrene infection/transfection Reagent (10 mg/ml) (Merck, catalog number: TR-1003-G )
- Bambanker serum-free freezing media (NIPPON Genetics, catalog number: BB01 )
- G418 solution (NACALAI TESQUE, catalog number: 16513-84 )
- Sodium chloride (NaCl) (NACALAI TESQUE, catalog number: 31320-05 )
- Na2HPO4·12H2O (Wako Pure Chemical Industries, catalog number: 196-02835 )
- Potassium dihydrogenphosphate (KH2PO4) (Wako Pure Chemical Industries, catalog number: 164-22635 )
- Potassium chloride (KCl) (Wako Pure Chemical Industries, catalog number: 160-22115 )
- Fetal bovine serum (FBS) (NICHIREI, Sigma-Aldrich, catalog number: 172012-500ml )
- Penicillin-streptomycin-glutamine (P/S) solution (NACALAI TESQUE, catalog number: 06168-34 )
- Dulbecco’s modified Eagle medium (DMEM) (NACALAI TESQUE, catalog number: 08459-64 )
- RPMI 1640 (NACALAI TESQUE, catalog number: 30264-56 )
- Ethylenediaminetetraacetic acid (EDTA) (NACALAI TESQUE, catalog number: 15130-95 )
- Sodium hydroxide (NaOH) (Wako Pure Chemical Industries, catalog number: 198-13765 )
- Ammonium chloride (NH4Cl) (Wako Pure Chemical Industries, catalog number: 017-02995 )
- Potassium hydrogen carbonate (KHCO3) (Wako Pure Chemical Industries, catalog number: 166-03275 )
- Bovine serum albumin (BSA) (Wako Pure Chemical Industries, catalog number: 019-23293 )
- Murine stem cell factor (SCF) (PeproTech, catalog number: 250-03 )
- Murine interleukin-3 (IL-3) (PeproTech, catalog number: 213-13 )
- Murine interleukin-6 (IL-6) (PeproTech, catalog number: 216-16 )
- Murine granulocyte-macrophage colony-stimulating factor (GM-CSF) (PeproTech, catalog number: 315-03 )
- IMDM powder (Thermo Fisher Scientific, GibcoTM, catalog number: 12200036 )
- Sodium hydrogen carbonate (NaHCO3) (Wako Pure Chemical Industries, catalog number: 195-14515 )
- Methyl cellulose (viscosity: 4,000 cP) (Sigma-Aldrich, catalog number: M0512 )
- Beta-mercaptoethanol (NACALAI TESQUE, catalog number: 21418-84 )
- Sodium pentobarbital (NACALAI TESQUE, catalog number: 02095-04 )
- 25x phosphate-buffered saline Ca2+/Mg2+-free (PBS) (see Recipes)
- D10 media (see Recipes)
- R10 media (see Recipes)
- R10W10 media (see Recipes)
- 0.5 M EDTA solution (see Recipes)
- ACK lysis buffer (see Recipes)
- SM buffer (see Recipes)
- Cytokine stocks (see Recipes)
- AC media (see Recipes)
- Anesthetic solution (see Recipes)
Equipment
- Pipetman P2, P20, P200, P1000
- 5% CO2 incubator 37 °C
- 5% CO2 incubator 32 °C
- Centrifuge for 15-ml conical tubes
- Cell counter (Hemacytometer)
- Laminar flow cabinet
- Surgical scissors and forceps
- MACS multistand (Miltenyi Biotec, catalog number: 130-042-303 )
- MiniMACS separator (Miltenyi Biotec, catalog number: 130-042-102 )
- Portable Pipet-Aid® XP pipette controller (Drummond Scientific, catalog number: 4-000-101 )
- -80 °C freezer
- 1-L glass bottle
- Autoclave
- Shaker
- Gammacell® 40 exactor with cesium-137 (Best Theratronics, model: Gammacell® 40 )
- Inverted microscope for general cell culture
Software
- GraphPad Prism (GraphPad Software, La Jolla, CA, USA)
Procedure
Schedule:
Day 1. Start culturing PLAT-E cells from freeze stock
Day 4. Replate PLAT-E cells for transfection
Day 5. Transfect PLAT-E cells
Day 6. Preparation of c-kit positive cells and irradiation of mice
Day 7. Transduction of retrovirus to c-kit positive cells and injection
Notes:
- Prepare a donor mouse (five-week-old female C57BL/6J mouse) and five recipient mice (seven-week-old female C57BL/6J mice) per sample.
- One positive control (MLL-ENL) and one negative control (Empty vector; pMSCV-neo) are required in this experiment.
- Virus preparation
- On day 1, thaw 4 x 106 PLAT-E cells from freeze stock and culture in 10-cm dishes with D10 media (see Recipes). The freezing media need to be removed for the initial culture.
Note: Culture size depends on the sample size. - On day 4, replate 4 x 106 PLAT-E cells in two 10-cm collagen-coated dishes with 10 ml of D10 media for a sample. In detail, remove the culture media, add 10 ml of PBS, remove PBS, add 1 ml of trypsin-EDTA and incubate for 5 min in a 37 °C 5% CO2 incubator. Add 9 ml of D10 media, transfer into a 15 ml tube, spin down cells at 400 x g for 5 min at room temperature, and then remove the supernatant. Add 10 ml of D10 media, count the cell number, and replate 4 x 106 PLAT-E cells in two 10-cm collagen-coated dishes with 10 ml of D10 media for a sample.
- On day 5, transfect PLAT-E cells with 16 µg of DNA and 40 µl of LipofectamineTM 2000 transfection reagent. At the time of transfection, the cell confluency should be 60-70%. In detail, dilute 16 µg of DNA in 1 ml of Opti-MEMTM media and dilute 40 µl of LipofectamineTM 2000 transfection reagent in 1 ml of Opti-MEMTM media, separately. After 5 min, combine the DNA solution with the LipofectamineTM 2000 solution. Mix gently and incubate for 20 min at room temperature. Add the DNA-Lipofectamine mixture to each dish and incubate in a 37 °C 5% CO2 incubator. After 6 h, replace the media with fresh D10 media and incubate in a 32 °C 5% CO2 incubator for 2 days.
- On day 1, thaw 4 x 106 PLAT-E cells from freeze stock and culture in 10-cm dishes with D10 media (see Recipes). The freezing media need to be removed for the initial culture.
- Preparation of c-kit (CD117) positive progenitor cells
- On day 6, euthanize the five-week-old female C57BL/6J mice by cervical dislocation, spray 70 % ethanol to the whole body in open air, and dissociate the femurs and tibiae from the donor mice in a laminar flow cabinet (Figure1). Peripheral muscle should be removed by surgical scissors as thoroughly as possible.
Note: The volume of solution described in the following procedures is for the cells from one donor mouse. If the temperature is not specifically noted, all the procedures should be conducted at room temperature.
Figure 1. Illustrative image of mouse tibia and femur - Cut the end of the femurs and tibiae and flush the bone marrow with 10 ml of PBS (see Recipes) using a 10-ml syringe attached with a 21 G needle (Figure 2).
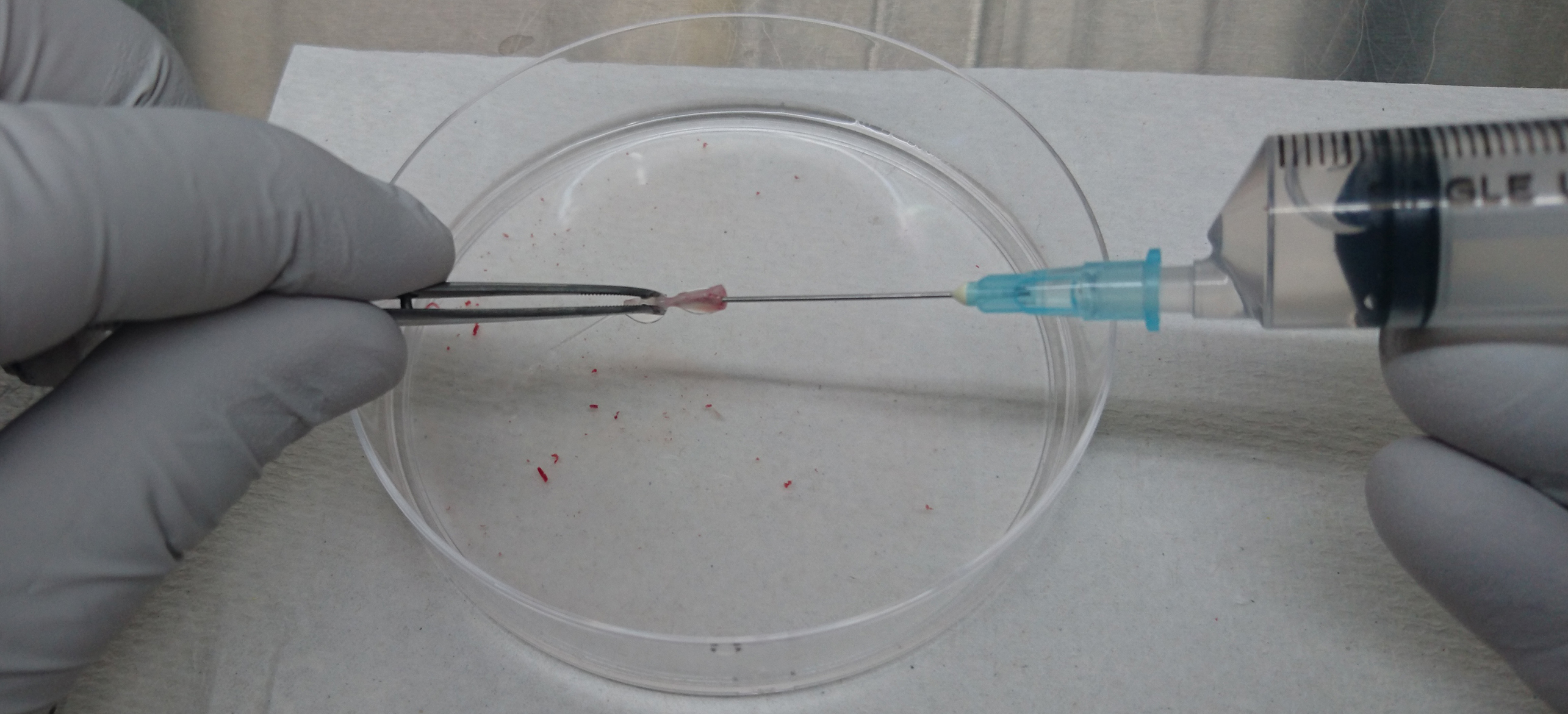

Figure 2. Isolation of bone marrow cells from tibia and femur - Homogenize the bone marrow cells gently by passing them through the 21 G needle several times.
- Spin down cells at 400 x g for 5 min, and then remove as much of the supernatant as possible.
- Resuspend the cells in 1 ml of ACK lysis buffer (see Recipes) and incubate on ice for 1 min.
- Add 10 ml of R10W10 media (see Recipes) and spin down cells at 400 x g for 5 min, and then remove the supernatant.
- Resuspend the cells in 10 ml of SM buffer (see Recipes) and spin down at 400 x g for 5 min, and then remove the supernatant.
- Resuspend the cells in 0.5 ml of SM buffer, add 20 µl of CD117 microbeads, and incubate for 20 min on ice or in the refrigerator.
- Add 10 ml of SM buffer and spin down cells at 400 x g for 5 min, and then remove the supernatant to wash the cells.
- Wash the cells again as in step B9.
- Install an MS column on a MACS magnetic stand and equilibrate with 1 ml of SM buffer.
- Place a pre-separation filter on top of the column and load the cells onto MS column through the pre-separation filter.
- Wash the column with 1 ml of SM buffer twice.
- Remove the column from the magnetic stand and place it on a new 15-ml conical tube.
- Add 1 ml of SM buffer to the column to elute c-kit-positive cells by gravity flow (optionally using a plunger).
- Add 10 ml of R10W10 media and spin down cells at 400 x g for 5 min, and then remove the supernatant.
- Resuspend all of the cells in 1 ml of R10W10 and transfer the cells into one well in a 48-well plate.
- Add cytokines (10 ng/ml SCF, 10 ng/ml IL-3, 10 ng/ml IL-6 at the final concentration) and incubate the cells in a 37 °C 5% CO2 incubator overnight.
- On day 6, euthanize the five-week-old female C57BL/6J mice by cervical dislocation, spray 70 % ethanol to the whole body in open air, and dissociate the femurs and tibiae from the donor mice in a laminar flow cabinet (Figure1). Peripheral muscle should be removed by surgical scissors as thoroughly as possible.
- Irradiation of mice
- On day 7, irradiate recipient mice with a sub-lethal dose (6 Gy) in a gamma-cell. Several mice can be irradiated at the same time using an individually separatable cage.
- Add 1 ml of 2.5% Baytril solution to 200 ml of drinking water and treat for 1 week. The Baytril-containing water is not necessarily changed during the 1-week treatment.
- On day 7, irradiate recipient mice with a sub-lethal dose (6 Gy) in a gamma-cell. Several mice can be irradiated at the same time using an individually separatable cage.
- Virus transduction
- On day 7, count the cells. A total of 1-2 x 106 c-kit-positive cells are expected from a donor mouse.
- Add an appropriate amount of R10W10 media to make a 6 x 105 cell/ml suspension and add 1/250 volume of 10 mg/ml polybrene solution.
- Aliquot 1 ml of cell suspensions into 15-ml conical tubes (two tubes for each sample).
- Suck 10 ml of the virus supernatant from the PLAT-E cell culture using a 10-ml syringe, attach a filter (0.45-µm) to the 10-ml syringe, and add the virus supernatant directly to the c-kit-positive cell suspension.
- Spin the cell suspension at 1,100 x g for 2.5 h at 32 °C (Spinoculation).
- After spinoculation, remove the virus supernatant and wash the cells twice each with 10 ml of PBS.
- Resuspend the cells to 2 x 105 cells/100 µl in PBS.
Note: The number of cells injected affects the duration and incidence of leukemogenesis. Thus, we can manipulate the duration by decreasing/increasing the cell number to some extent. - Place the cell suspensions on ice until injection.
- On day 7, count the cells. A total of 1-2 x 106 c-kit-positive cells are expected from a donor mouse.
- Transplantation
- Inject 300 µl of anesthetic solution (see Recipes) intraperitoneally using a tuberculin syringe. Mice will be anesthetized completely within 5 min and stay asleep up to 1 h.
- After 5 min, inject 100 µl of the cell suspension into the retro-orbital sinus using a Myjector® insulin syringe. Pop up the eye, place the needle 1-2 mm deep between the eye and skin toward the brain to reach the vein, and gently inject the cell suspension (Figure 3) (Yardeni et al., 2011).
Notes: Alternatively, the tail vein injection is applicable instead of retro-orbital injection. - Place the mice into a cage for recovery and monitoring.
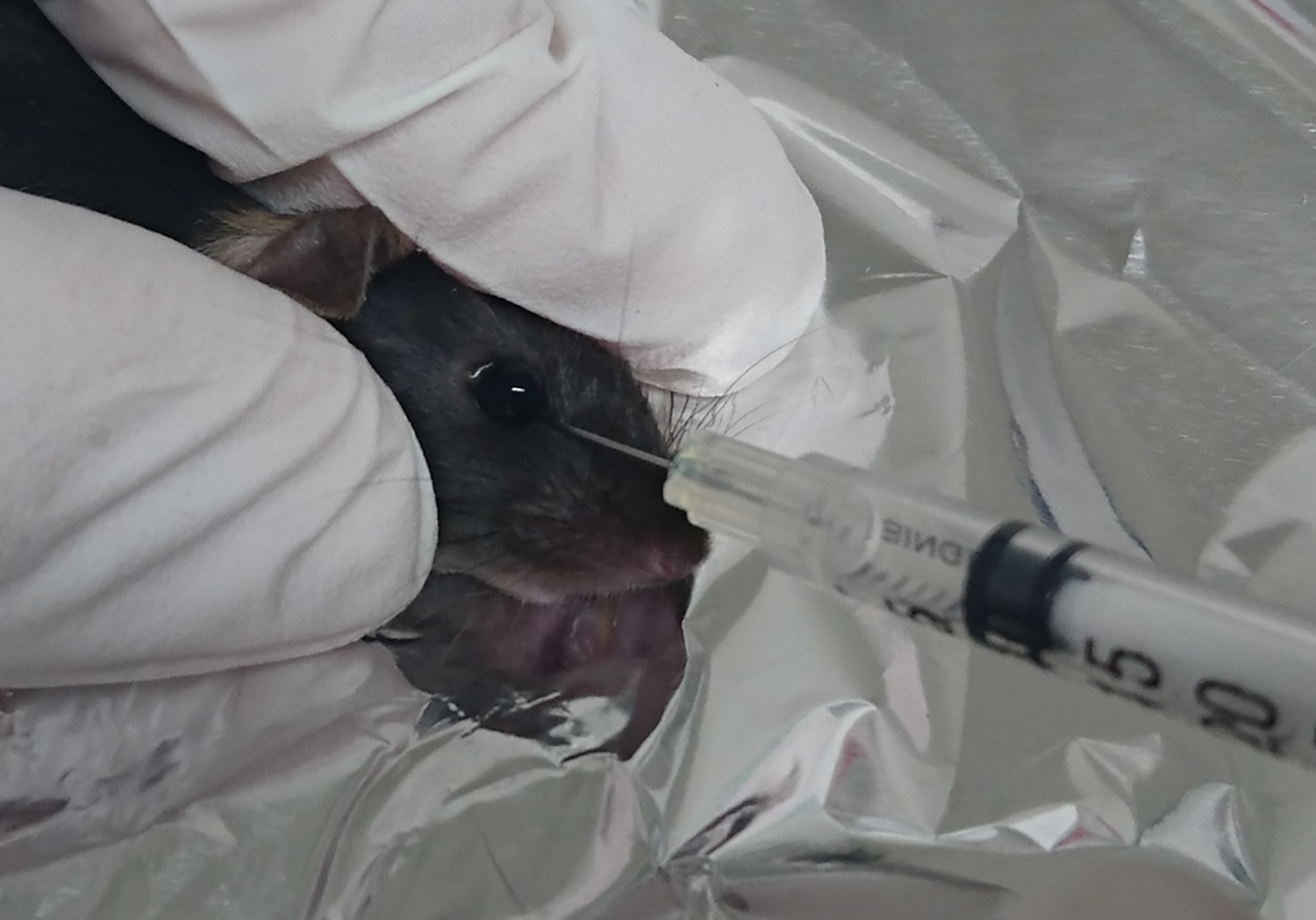
Figure 3. Retro-orbital injection in mice
- Inject 300 µl of anesthetic solution (see Recipes) intraperitoneally using a tuberculin syringe. Mice will be anesthetized completely within 5 min and stay asleep up to 1 h.
- Tumor cell isolation and culture
- When a mouse exhibits a morbid look, sacrifice the mouse, and dissociate the tumor cells from the femurs and tibiae as the same method of bone marrow preparation (steps B1-B6). Optionally, dissociate the spleen for pathological analysis.
Notes: Morbid look indicates the condition of losing weight, bad coat of fur, and minimum reaction against stimulation e.g., sound and poking with a fingertip. It can be normally observed from day 60 to day 140 in the primary transplantation, and from day 20 to day 50 in the secondary transplantation. - Prepare the bone marrow cell suspension with ACK buffer treatment described in steps B2-B6.
- Add 1 ml of R10W10 media and count the cell number.
- Add an appropriate amount of R10W10 media to make a 5 x 105 cells/ml suspension.
- Aliquot 200 µl of the cell suspension, add 1 ml of AC media (see Recipes) using a 1-ml syringe attached with an 18 G needle, and mix by vortexing. Optionally, the residual cells can be stocked. In detail, spin down cells at 400 x g for 5 min, and then remove the supernatant. Resuspend the cells in freezing media (We typically use the Bambanker freezing solution, but R10W10 plus 10% DMSO can be alternatively used.) and directly store at -80 °C. For long-term storage, storage tube should be transferred to liquid nitrogen from -80 °C freezer.
- Add 20 µl of G418 solution to eliminate normal cells and incubate in a 37 °C 5% CO2 incubator for 5 days.
- Transfer the cells into a 12-well plate using a 1-ml syringe with an 18 G needle and incubate in a 37 °C 5% CO2 incubator for several days. During the culture, prepare the recipient mice for secondary transplantation. If the media turns orange during culture because of excess cells, add an additional 1 ml of AC media with G418.
Note: It is important to maintain good culture conditions. Overgrowth drastically affects the incidence of leukemia in the second transplantation.
- When a mouse exhibits a morbid look, sacrifice the mouse, and dissociate the tumor cells from the femurs and tibiae as the same method of bone marrow preparation (steps B1-B6). Optionally, dissociate the spleen for pathological analysis.
- Secondary transplantation
- Collect cells with 10 ml of PBS, transfer to 15-ml conical tubes, and spin at 400 x g for 5 min.
- Remove the supernatant, resuspend the cells in 10 ml of PBS, and remove the debris to pass through a Pre-separation filter.
- Count the cell number and resuspend the cells to 2 x 105 cells/100 µl in PBS.
Note: The number of cells injected affects the duration and incidence of leukemogenesis. Thus, we can manipulate the duration by decreasing/increasing the cell number to some extent. - Place the cell suspension on ice.
- Inject 300 µl of anesthetic solution intraperitoneally using a tuberculin syringe.
- Inject 100 µl of the cell suspension into the retro-orbital sinus using a Myjector® insulin syringe.
- Place the mice into a cage for recovery and monitoring.
Note: The duration of survival in secondary transplantation shortens compared to that in initial transplantation (Figure 4). It could be because leukemia cells need to acquire genetic/epigenetic alterations during the first disease onset. Or, the cancer stem/initiating population may be more enriched at the time of second transplantation compared to the initial transplantation. It is unclear at this point why the duration has been shortened dramatically in the second transplantation.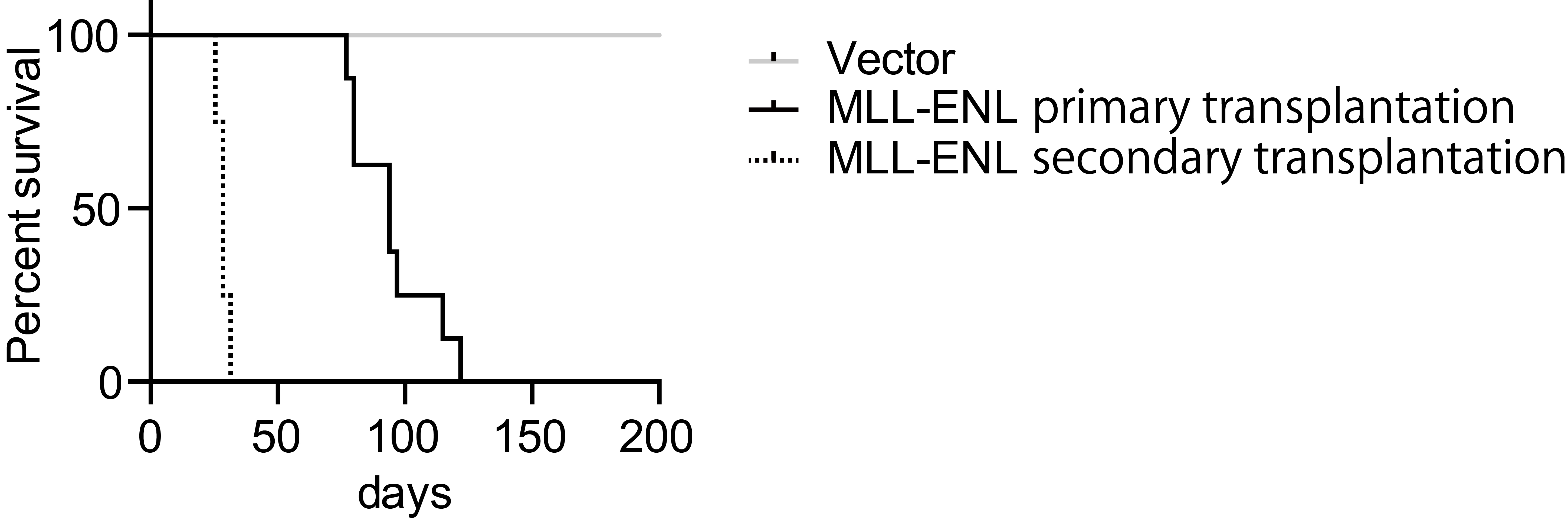
Figure 4. Leukemogenic potential of MLL-ENL in this model
- Collect cells with 10 ml of PBS, transfer to 15-ml conical tubes, and spin at 400 x g for 5 min.
Data analysis
- We typically perform this experiment a minimum of two times using five recipient mice per sample. Total ten recipient mice per sample are needed for primary transplantation in two independent experiments. To confirm the leukemogenic ability, five recipient mice are typically transplanted with primary leukemia cells.
- For survival analysis, we use GraphPad Prism software. To compare two data sets, we use a Log-rank method. To compare three or more data sets, we use the Gehan-Breslow-Wilcoxon test.
Recipes
- 25x phosphate-buffered saline Ca2+/Mg2+-free (PBS) (1 L)
- Mix 200 g NaCl, 72.4 g Na2HPO4·12H2O, 5 g KH2PO4, and 5 g KCl
- Bring to 1 L with distilled H2O
- Autoclave for 20 min at 121 °C
- Dilute to 1x with distilled H2O for working solution
- Mix 200 g NaCl, 72.4 g Na2HPO4·12H2O, 5 g KH2PO4, and 5 g KCl
- D10 media
Add 55 ml of FBS and 5.5 ml of P/S solution to 500 ml of DMEM - R10 media
Add 55 ml of FBS and 5.5 ml of P/S solution to 500 ml of RPMI 1640 - R10W10 media
- Culture WEHI-3 in R10 media to confluent
- When media turns orange, collect and filter (0.22 µm) the media
- Aliquot and store at -80 °C
- Add 55 ml of FBS, 55 ml of WEHI-3 culture media, 5.5 ml of P/S solution to 500 ml of RPMI 1640
- Culture WEHI-3 in R10 media to confluent
- 0.5 M EDTA solution
- Weigh 93.06 g EDTA
- Bring to 1 L with distilled water and adjust pH to 8.0 with NaOH
- Weigh 93.06 g EDTA
- ACK lysis buffer
- Mix 8.29 g of NH4Cl and 1 g of KHCO3
- Bring to 1 L with distilled water
- Add 200 µl of 0.5 M EDTA solution
- Filter (0.2-µm) and store at 4 °C
- Mix 8.29 g of NH4Cl and 1 g of KHCO3
- SM buffer
- Add 15 ml of FBS to 500 ml of 1x PBS
- Filter (0.2-µm) and store at 4 °C
- Add 15 ml of FBS to 500 ml of 1x PBS
- Cytokine stocks
- Dissolve 10 mg of BSA powder in 10 ml of PBS and filter (0.2-µm) (PBS + 0.1% BSA)
- Dissolve cytokines (SCF, IL-3, IL6, GM-CSF) to 50 µg/ml with PBS + 0.1% BSA
- Aliquot and store at -80 °C
- Dissolve 10 mg of BSA powder in 10 ml of PBS and filter (0.2-µm) (PBS + 0.1% BSA)
- AC media
- Dissolve IMDM powder with 500 ml of distilled water, add 3 g NaHCO3, and filter (0.2-µm)
- Weigh 16 g of methyl cellulose in a 1-L glass bottle
- Autoclave methyl cellulose powder for 20 min at 121 °C
- Dissolve sterile methyl cellulose with 300 ml of sterile water and 500 ml of IMDM in a shaker overnight
- Add 200 ml of FBS and 7 µl of beta-mercaptoethanol
- Aliquot and store at -20 °C
- Before use, add 20 µl SCF, 20 µl IL-3, 20 µl GM-CSF, and 1 ml P/S solution
- Dissolve IMDM powder with 500 ml of distilled water, add 3 g NaHCO3, and filter (0.2-µm)
- Anesthetic solution
- Dilute sodium pentobarbital solution by ten-fold with PBS before use
- The final concentration of sodium pentobarbital should be 5 mg/ml
- Dilute sodium pentobarbital solution by ten-fold with PBS before use
Acknowledgments
This study was supported by JSPS KAKENHI grants to H.O. (number 17H07379) and A.Y. (number 16H05337). This protocol is based on a previous report by Lavau et al. (1997). The authors declare no conflict of interest.
References
- Andersson, A. K., Ma, J., Wang, J., Chen, X., Gedman, A. L., Dang, J., Nakitandwe, J., Holmfeldt, L., Parker, M., Easton, J., Huether, R., Kriwacki, R., Rusch, M., Wu, G., Li, Y., Mulder, H., Raimondi, S., Pounds, S., Kang, G., Shi, L., Becksfort, J., Gupta, P., Payne-Turner, D., Vadodaria, B., Boggs, K., Yergeau, D., Manne, J., Song, G., Edmonson, M., Nagahawatte, P., Wei, L., Cheng, C., Pei, D., Sutton, R., Venn, N. C., Chetcuti, A., Rush, A., Catchpoole, D., Heldrup, J., Fioretos, T., Lu, C., Ding, L., Pui, C. H., Shurtleff, S., Mullighan, C. G., Mardis, E. R., Wilson, R. K., Gruber, T. A., Zhang, J., Downing, J. R. and St. Jude Children's Research Hospital-Washington University Pediatric Cancer Genome, P. (2015). The landscape of somatic mutations in infant MLL-rearranged acute lymphoblastic leukemias. Nat Genet 47(4): 330-337.
- Ayton, P. M., and Cleary, M. L. (2003). Transformation of myeloid progenitors by MLL oncoproteins is dependent on Hoxa7 and Hoxa9. Genes Dev 17: 2298-2307.
- Jude, C. D., Climer, L., Xu, D., Artinger, E., Fisher, J. K. and Ernst, P. (2007). Unique and independent roles for MLL in adult hematopoietic stem cells and progenitors. Cell Stem Cell 1(3): 324-337.
- Lavau, C., Szilvassy, S. J., Slany, R. and Cleary, M. L. (1997). Immortalization and leukemic transformation of a myelomonocytic precursor by retrovirally transduced HRX-ENL. EMBO J 16(14): 4226-4237.
- Meyer, C., Burmeister, T., Groger, D., Tsaur, G., Fechina, L., Renneville, A., Sutton, R., Venn, N. C., Emerenciano, M., Pombo-de-Oliveira, M. S., Barbieri Blunck, C., Almeida Lopes, B., Zuna, J., Trka, J., Ballerini, P., Lapillonne, H., De Braekeleer, M., Cazzaniga, G., Corral Abascal, L., van der Velden, V. H. J., Delabesse, E., Park, T. S., Oh, S. H., Silva, M. L. M., Lund-Aho, T., Juvonen, V., Moore, A. S., Heidenreich, O., Vormoor, J., Zerkalenkova, E., Olshanskaya, Y., Bueno, C., Menendez, P., Teigler-Schlegel, A., Zur Stadt, U., Lentes, J., Gohring, G., Kustanovich, A., Aleinikova, O., Schafer, B. W., Kubetzko, S., Madsen, H. O., Gruhn, B., Duarte, X., Gameiro, P., Lippert, E., Bidet, A., Cayuela, J. M., Clappier, E., Alonso, C. N., Zwaan, C. M., van den Heuvel-Eibrink, M. M., Izraeli, S., Trakhtenbrot, L., Archer, P., Hancock, J., Moricke, A., Alten, J., Schrappe, M., Stanulla, M., Strehl, S., Attarbaschi, A., Dworzak, M., Haas, O. A., Panzer-Grumayer, R., Sedek, L., Szczepanski, T., Caye, A., Suarez, L., Cave, H. and Marschalek, R. (2017). The MLL recombinome of acute leukemias in 2017. Leukemia.
- Morita, S., Kojima, T. and Kitamura, T. (2000). Plat-E: an efficient and stable system for transient packaging of retroviruses. Gene Ther 7(12): 1063-1066.
- Okuda, H., Kanai, A., Ito, S., Matsui, H. and Yokoyama, A. (2015). AF4 uses the SL1 components of RNAP1 machinery to initiate MLL fusion- and AEP-dependent transcription. Nat Commun 6: 8869.
- Okuda, H., Kawaguchi, M., Kanai, A., Matsui, H., Kawamura, T., Inaba, T., Kitabayashi, I. and Yokoyama, A. (2014). MLL fusion proteins link transcriptional coactivators to previously active CpG-rich promoters. Nucleic Acids Res 42(7): 4241-4256.
- Okuda, H., Stanojevic, B., Kanai, A., Kawamura, T., Takahashi, S., Matsui, H., Takaori-Kondo, A. and Yokoyama, A. (2017). Cooperative gene activation by AF4 and DOT1L drives MLL-rearranged leukemia. J Clin Invest 127(5): 1918-1931.
- Okuda, H., Takahashi, S., Takaori-Kondo, A. and Yokoyama, A. (2016). TBP loading by AF4 through SL1 is the major rate-limiting step in MLL fusion-dependent transcription. Cell Cycle 15(20): 2712-2722.
- Somervaille, T. C. and Cleary, M. L. (2006). Identification and characterization of leukemia stem cells in murine MLL-AF9 acute myeloid leukemia. Cancer Cell 10(4): 257-268.
- Yardeni, T., Eckhaus, M., Morris, H. D., Huizing, M. and Hoogstraten-Miller, S. (2011). Retro-orbital injections in mice. Lab Anim (NY) 40(5): 155-160.
- Yokoyama, A., Ficara, F., Murphy, M. J., Meisel, C., Hatanaka, C., Kitabayashi, I. and Cleary, M. L. (2013). MLL becomes functional through intra-molecular interaction not by proteolytic processing. PLoS One 8(9): e73649.
- Yokoyama, A., Lin, M., Naresh, A., Kitabayashi, I. and Cleary, M. L. (2010). A higher-order complex containing AF4 and ENL family proteins with P-TEFb facilitates oncogenic and physiologic MLL-dependent transcription. Cancer Cell 17(2): 198-212.
Article Information
Copyright
© 2017 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Okuda, H. and Yokoyama, A. (2017). In vivo Leukemogenesis Model Using Retrovirus Transduction. Bio-protocol 7(23): e2627. DOI: 10.21769/BioProtoc.2627.
Category
Cancer Biology > General technique > Animal models
Cell Biology > Cell isolation and culture > Transformation
Cancer Biology > Oncogenesis > Leukemogenesis
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link








