- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Immunogold Localization of Molecular Constituents Associated with Basal Bodies, Flagella, and Extracellular Matrices in Male Gametes of Land Plants
($Retired) Published: Vol 7, Iss 21, Nov 5, 2017 DOI: 10.21769/BioProtoc.2599 Views: 7867
Reviewed by: Scott A M McAdamAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
An In-depth Guide to the Ultrastructural Expansion Microscopy (U-ExM) of Chlamydomonas reinhardtii
Nikolai Klena [...] Virginie Hamel
Sep 5, 2023 4852 Views
Fast and High-Resolution Imaging of Pollinated Stigmatic Cells by Tabletop Scanning Electron Microscopy
Lucie Riglet and Isabelle Fobis-Loisy
Nov 20, 2024 1823 Views
Cryo-SEM Investigation of Chlorella Using Filter Paper as Substrate
Peng Wan [...] Jinghan Wang
Dec 20, 2024 1485 Views
Abstract
Male gametes (spermatozoids) are the only motile cells produced during the life cycle of land plants. While absent from flowering and most cone-bearing plants, motile cells are found in less derived taxa, including bryophytes (mosses, liverworts and hornworts), pteridophytes (lycophytes and ferns) and some seed plants (Ginkgo and cycads). During development, these cells undergo profound changes that involve the production of a locomotory apparatus, unique microtubule (MT) arrays, and a series of special cell walls that are produced in sequence and are synchronized with cellular differentiation. Immunogold labeling in the transmission electron microscope (TEM) provides information on the exact location and potential function of macromolecules involved with this developmental process. Specifically, it is possible to localize epitopes to proteins that are associated with the cellular inclusions involved in MT production and function. Spermatogenesis in these plants is also ideal for examining the differential expression of carbohydrates and glycoproteins that comprise the extracellular matrixes associated with the dramatic architectural changes in gamete shape and locomotory apparatus development. Here we provide methodologies using monoclonal antibodies (MAbs) and immunogold labeling in the TEM to localize macromolecules that are integral to spermatozoid development.
Keywords: Arabinogalactan proteinsBackground
Motile gametes of land plants are strikingly diverse with numbers of flagella ranging from two to greater than 40,000 (Renzaglia and Garbary, 2001). Following a series of synchronized mitotic divisions within antheridia, nascent sperm cells (spermatids) undergo a sequence of developmental changes within the confines of a dynamic and growing cell wall. A complex locomotory apparatus is produced and flagella elongate around the cell as the organelles are repositioned and shaped. Synchronized development yields hundreds of cells in a single stage of maturation and in different planes of section within a single antheridium.
This profound cellular differentiation involves the development of unique MT arrays, the spline and flagella, that emanate from discrete microtubule organizing centers (MTOCs), the only centriole-containing centrosomes in land plants. Because of the exclusive occurrence of basal bodies, flagella and associated complexes in developing male gametes, studies of spermatogenesis have revealed important information on the structure, composition, and developmental changes in MT arrays as they relate to the cell cycle, MTOCs and cellular differentiation in plants (Joshi et al., 1992; Lui et al., 1993; Vaughn and Renzaglia, 1993; Hoffman et al., 1994; Hoffman and Vaughn, 1995; Vaughn and Harper, 1998; Klink and Wolniak, 2003; Vaughn and Renzaglia, 2006; Vaughn and Bowling, 2008; Vaughn, 2013). A dynamic and flexible extracellular matrix is necessary for spermatogenesis to take place (Garbary and Renzaglia, 2001; Lopez and Renzaglia, 2014); thus spermatogenesis in plants provides an opportunity to examine cell wall changes during development.
The purpose of this review is to describe the methodologies used in localizing proteins, carbohydrates and glycoproteins during the development of motile gametes in land plants. One of the most powerful tools in these studies involves antibodies that recognize epitopes to macromolecules using immunogold labeling techniques at the TEM level (Vaughn, 2013). Here we provide images and a brief discussion of results using immunogold labeling to examine the molecular constituents involved in sperm cell development in plants. Two procedures for these investigations are provided that use the same Materials and Reagents, and Equipment: Procedure A describes the protocol for microtubule-related proteins and procedure B for localizing cell wall constituents.
Procedure A: Immunogold labeling has led to important advances in understanding the role of the proteins centrin and tubulin in plants (Figures 1A-1E). Centrin is a ~20 kDa Ca-binding protein first discovered in motile green algae (Satisbury, 1995), where it is localized to the stellate pattern in the transition zone of the flagella and a dense band of fibers (distal fibers) that connect the nucleus to the basal bodies. In spermatogenous cells, centrin localizes to specific, seemingly diverse structures (Figures 1A-1D). The plates of the multilayered structure (MLS) of the locomotory apparatus, but not the microtubules (MTs), are strongly labeled with antibodies that recognize centrin (Vaughn and Renzaglia, 1993; Vaughn and Harper, 1998) (Figures 1A and 1B). The transition zone that occurs in the flagella of most plants with motile cells also strongly labels with antibodies to centrin, indicating homology with a similar zone in the basal body apparatus in green algae (Vaughn and Renzaglia, 1993; Hoffman et al., 1994; Vaughn and Harper, 1998; Klink and Wolniak, 2003; Vaughn and Renzaglia, 2006) (Figures 1C and 1D). In tracheophytes, an electron opaque pericentriolar type material, called the amorphous zone (AZ), runs along the top of the spline (MT band), connects the basal bodies of adjacent flagella and labels with centrin antibodies (Figure 1B) (Hoffman et al., 1994; Hoffman and Vaughn, 1995). The AZ thus serves as that same sort of basal body connector as found in the green algae.
These localizations are indicative of two possible functions for centrin, as an MTOC protein, and as a contractile protein. In the MLS, centrin appears to be involved in MT nucleation and organization of the spline MT array. In the AZ and transition zone, it is more likely that the centrin is involved in contractile functions. The AZ might also be involved in nucleation/organization of MTs as it lies at the base of the basal body and is close to the spline MT array.
Gamma tubulin was the last of the tubulin proteins to be discovered (Oakley et al., 1990) and occurs in a much lower quantity than alpha and beta tubulin. In mammalian cells, gamma tubulin is restricted to the ends of MTs, where it forms a template for the MTs to form (Joshi et al., 1992). In contrast, gamma tubulin in plants occurs along MTs and not just at their termini (Liu et al., 1993; Liu et al., 1994; Hoffman et al., 1994; Vaughn and Harper, 1998). These sites are in fact new nucleating sites as plant MTs form more of a ‘fir tree’ or highly branched pattern than that is noted in mammalian cells (Murata et al., 2005; Murata and Hasebe, 2007).
The blepharoplast occurs in the last two spermatogenous cell divisions in pteridophytes and serves as the spindle pole body in these divisions as well as the template for basal body production (Hepler, 1976; Hoffman and Vaughn, 1995). In an attempt to determine its ability to nucleate MTs, Vaughn and Bowling (2008) treated Ceratopteris antheridia with the potent microtubule-disrupter oryzalin (ChemService Inc., West Chester, PA) leading to the loss of all microtubules except those in stabilized MT arrays such as in flagella. In these oryzalin-treated cells, the blepharoplast was clearly recognizable but free of MTs and covered with pits that were the size and structure of the MT templates (tubulin ring complexes) recognized in mammalian cells. 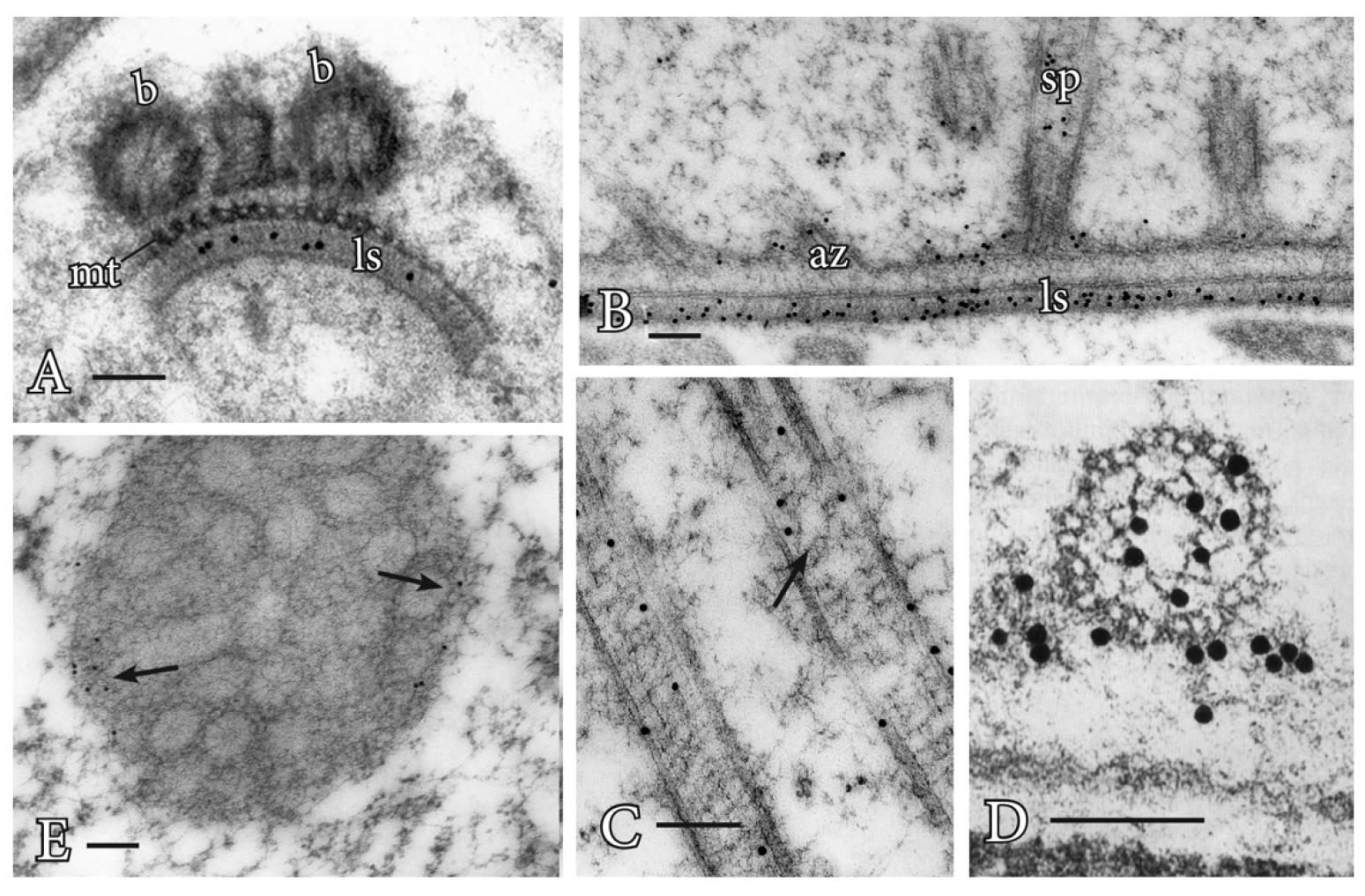
Figure 1. Immunogold labeling of basal bodies and flagella of land plants. A. Centrin localization in the lamella strip (ls) that subtends the band of microtubules (mt) and basal bodies (b) of Phaeoceros carolinianus, a hornwort. B. In the seed plant, Ginkgo biloba, centrin epitopes are localized in the lamellar strip (ls), amorphous zone (az) where the basal bodies insert and stellate pattern (sp) of the transition zone. C. Longitudinal section of the long stellate pattern in Ceratopteris basal bodies that label with anti-centrin. A faint outline of the stellate pattern is visible at the arrow. D. Cross section at the basal body of Ceratopteris showing centrin localizations in the stellate pattern and amorphous zone around the basal body. E. In Ceratopteris spermatogenous cells, gamma tubulin (arrows) localizes around the periphery of the blepharoplast following oryzalin treatment. Bars = 0.1 µm.
Gamma tubulin antibodies label the periphery of the blepharoplast in oryzalin treated cells (Figure 1E). When the oryzalin is washed from the antheridia, MTs are quickly reformed along this pitted surface, further indicating the ability of the blepharoplast to serve as an MTOC.
In mammalian cells, the centrioles are surrounded by an electron opaque material where spindle MTs emanate. To identify the components of this pericentriolar material, monoclonal antibodies (MAbs) were raised to mitotic cells and MAbs that recognize the centriolar material could be used not only for mammalian cells but also for other materials, including spermatogenous cells. For example, MPM-2 recognizes a phosphorylated-protein epitope (Davis et al., 1983; Vandre et al., 1984) in spermatogenous cells. In cells without blepharoplasts, this MAb recognizes the surface of the nuclear envelope immediately before mitosis (Hoffman et al., 1994; Klink and Wolniak, 2003). These are the sites where MTs appear to be produced prior to mitosis in all plant cells. In cells with a blepharoplast, these antibodies strongly label the interior of this structure, not the edges (Hoffman et al., 1994; Vaughn and Bowling, 2008). Interestingly, as the blepharoplast begins to reorganize, the reactivity of the antibody is lost and centrin labeling increases in the pericentriolar material. Thus, as different MT arrays are formed, changes occur in proteins of the MTOC.
Procedure B: Immunogold localizations of the sequential matrices that are made during spermatogenesis has revealed differential labeling of carbohydrate-specific MAbs during development and across phylogeny. Callose is a prominent wall constituent in spermatogenesis of ferns, especially in the thickened wall of rounded spermatids in the early stages of ontogeny. In this stage, the locomotory apparatus originates and consists of a multilayered structure (MLS) and basal bodies (Figure 2A). Pectin is absent in this thickened wall in ferns (Lopez and Renzaglia, 2017). In contrast, mosses have a comparable wall that is deposited as spermatids become round, but it is devoid of callose and contains scattered aggregates of esterified pectin as localized with the JIM7 MAb (Figure 2B). By far the most abundant polysaccharide in this thickened wall layer in mosses is hemicellulose that localizes with both LM15 and LM25 MAbs (Figures 2C and 2D) (Lopez-Swalls, 2016).
In addition to carbohydrates, the walls involved in plant spermatogenesis contain abundant but diverse arabinogalactan proteins (AGPs) (Figures 2E-2G). AGPs recognized by the LM2 MAb replace the hemicelluloses around moss spermatids (Figure 2E). As the spermatid matures and begins to develop flagella and assume a coiled configuration, a flexible extraprotoplasmic matrix forms between the plasmalemma and thick callosic wall in ferns and between the plasmalemma and hemicellulosic-pectinaceous wall of mosses (Figures 2F and 2G). The matrix does not label with monoclonal antibodies raised against standard cell wall polysaccharide epitopes such as pectins, cellulose, and hemicelluloses. Rather, MAbs that recognize sugar residues of AGPs abundantly label the matrix as well as the plasmalemma of elongating flagella in fern and moss spermatids (Figures 1F and 1G) (Lopez and Renzaglia, 2014). These results coupled with light and fluorescence microscopy and inhibitor experiments with Yariv (a reagent that binds and precipitates AGPs) suggest that AGPs are involved in growth and positioning of flagella. The implication of AGPs as calcium modulators through binding and release of Ca2+ (Lamport and Várnai, 2013) is a potential mechanism for the regulation of cellular development in plants, and spermatogenesis is an ideal system in which to further pursue this hypothesis.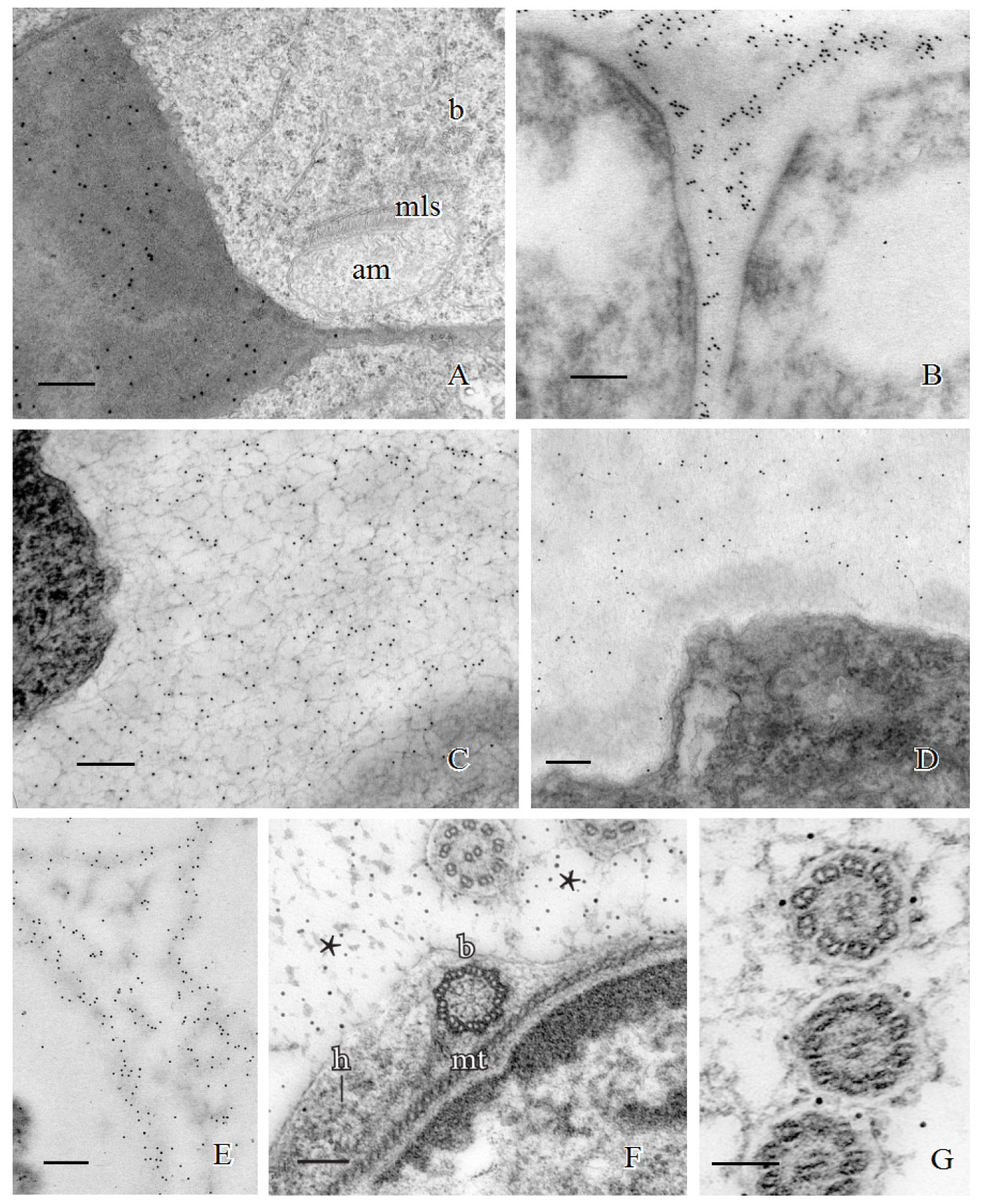
Figure 2. Immunogold labeling of spermatogenous cell walls in land plants. A. Callose localization in the unevenly-thickened wall layer that surrounds spermatids of Ceratopteris richardii during the formation of the locomotory apparatus which includes an anterior mitochondrion (am), a multilayered structure (mls) and basal bodies (b). B. A thickened wall layer comparable to that in Ceratopteris spermatids, is deposited by young spermatids in the moss, Physcomitrella patens. This wall labels intensely with the JIM7 MAb that binds to esterified pectin epitopes. C-D. Immunogold labeling of hemicelluloses in the thickened wall layer of young spermatids in the moss Aulacomnium palustre. C. This wall layer contains abundant xyloglucan epitopes recognized by the LM15 MAb. D. Similarly, galactoxyloglucan epitopes (LM25 MAb) are a rich component of these thickened walls. (Note: Spermatids in C. were post-fixed in osmium tetroxide (OsO4) that reveals the loose fibrillar consistency of the wall compared to the spermatids in D. that were not post-fixed in OsO4.) E-G. Immunogold labeling of arabinogalactan proteins (AGPs) in spermatid walls. E. AGP epitopes recognized by the LM2 MAb replace the hemicelluloses in the wall around spermatids in P. patens. F. JIM13, a monoclonal antibody to the epitope structure (β)-D-GlcpA1-(1,3)-α-D-GalpA-(1,2)-L-Rha of AGPs, is expressed in the extracellular matrix (*) around flagella during development in C. richardii. The microtubule band (mt), basal body (b) and a hub extension (h) are visible inside the developing spermatid. G. Cross sections of Ceratopteris flagella labelled with JIM8, a monoclonal antibody that identifies an unknown AGP epitope, showing specific localization on the plasmalemma. Bars = 0.5 µm for A-E; 0.1 µm for F-G.
Materials and Reagents
- Immunogold labeling of microtubules and microtubule organizing center proteins
- Scintillation vials with aluminum covered caps (Fisher Scientific, catalog number: 03-340-4B)
Manufacturer: DWK Life Sciences, Kimble, catalog number: 7450320 .
- Pasteur pipette (Fisher Scientific, catalog number: 13-678-20C )
- Gelatin capsules (Electron Microscopy Sciences, catalog number: 70100 )
- 200 mesh nickel grids (Electron Microscopy Sciences, catalog number: EMS200-Ni )
- 300 mesh gold grids (Electron Microscopy Sciences, catalog number: EMS300-Au )
- Glass Petri dishes (Corning, catalog number: 3160-101 )
- 90 mm filter paper (GE Healthcare, catalog number: 1004-090 )
- Sterile filters (0.2 µm) (Corning, catalog number: 431212 )
- Parafilm (Sigma-Aldrich, Parafilm, catalog number: P7793 )
- Dental wax plate (Electron Microscopy Sciences, catalog number: 72670 )
- Glass slides (Fisher Scientific, catalog number: 12-544-1 )
- Microcentrifuge tubes, 1.5 ml natural (USA Scientific, catalog number: 1615-5500 )
- Ethylene dichloride (Electron Microscopy Sciences, catalog number: 13250 )
- LR white resin (Electron Microscopy Sciences, catalog number: 14383 )
- Primary antibodies: (see Table 1), centrin (Sigma-Aldrich, catalog number: ABE480 ); MPM-2 (EMD Millipore, catalog number: 05-368 )
Table 1. Primary antibodies used to immunogold label microtubules, microtubule organizing center proteins, and carbohydrates and arabinogalactan proteins in extracellular matrices during spermatogenesis

aPhaeoceros carolinianus (hornwort); bPhyscomitrella patens (moss); cAulacomnium palustre (moss); dCeratopteris richardii (fern), and eGinkgo biloba (seed plant).
- Secondary antibody: goat anti-mouse conjugated with gold (Millipore Sigma, catalog number: G7652 )
- Sorenson’s phosphate buffer, 0.2 M, pH 7.2 (Electron Microscopy Sciences , catalog number: 11600-10 )
- Glutaraldehyde (Electron Microscopy Sciences, catalog number: 16120 )
- Osmium tetroxide (Electron Microscopy Sciences, catalog number: 19150 )
- PIPES buffer, 0.2 M, pH 7.2 (Sigma-Aldrich, catalog number: P6757 )
- Bovine serum albumin (BSA) (Sigma-Aldrich, catalog number: B4287 )
- Uranyl acetate (Polyscience, catalog number: 21447 )
- Lead nitrate (Electron Microscopy Sciences, catalog number: 17900 )
- Sodium citrate (Electron Microscopy Sciences, catalog number: 21140 )
- 1 N NaOH (Electron Microscopy Sciences, catalog number: 21170-01 )
- 0.01 M phosphate buffer (pH 7.2) (see Recipes)
- 0.05 M phosphate buffer (pH 7.2) (see Recipes)
- 2.5% glutaraldehyde (see Recipes)
- 2% aqueous osmium tetroxide (see Recipes)
- 0.02 M phosphate buffer (pH 7.2) (see Recipes)
- Antibodies (see Recipes)
- 2% PBS/BSA (see Recipes)
- 0.05 M PIPES buffer (pH 7.2) (see Recipes)
- 2% uranyl acetate (see Recipes)
- 1 N NaOH (see Recipes)
- Reynold’s lead citrate (see Recipes)
- Scintillation vials with aluminum covered caps (Fisher Scientific, catalog number: 03-340-4B)
- Immunogold labeling of spermatozoid matrix and cell wall constituents
- Scintillation vials with aluminum covered caps (Fisher Scientific, catalog number: 03-340-4B)
Manufacturer: DWK Life Sciences, Kimble, catalog number: 7450320 .
- Pasteur pipette (Fisher Scientific, catalog number: 13-678-20C )
- 200 mesh nickel grids (Electron Microscopy Sciences, catalog number: EMS200-Ni )
- 300 mesh gold grids (Electron Microscopy Sciences, catalog number: EMS300-Au )
- 90 mm filter paper (GE Healthcare, catalog number: 1004-090 )
- Dental wax plate (Electron Microscopy Sciences, catalog number: 72670 )
- Glass Petri dish (Corning, catalog number: 3160-101 )
- Glass Slides (Fisher Scientific, catalog number: 12-544-1 )
- Gelatin capsules (Electron Microscopy Sciences, catalog number: 70100 )
- Microcentrifuge tubes, 1.5 ml natural (USA Scientific, catalog number: 1615-5500 )
- Parafilm (Sigma-Aldrich, Parafilm, catalog number: P7793 )
- Sterile filters (0.2 µm) (Corning, catalog number: 431212 )
- Ethylene dichloride (Electron Microscopy Sciences, catalog number: 13250 )
- LR white resin (Electron Microscopy Sciences, catalog number: 14383 )
- Primary antibodies (PlantProbes) (see Table 1)
- Secondary antibody: Goat-Anti-Rat IgG-gold (Sigma-Aldrich, catalog number: G7035 )
- Secondary antibody: goat anti-mouse conjugated with gold (Sigma-Aldrich, catalog number: G7652 )
- Sorensen’s phosphate buffer, 0.2 M, pH 7.2 (Electron Microscopy Sciences , catalog number: 11600-10 )
- Glutaraldehyde (Electron Microscopy Sciences, catalog number: 16120 )
- Osmium tetroxide (Electron Microscopy Sciences, catalog number: 19150 )
- PIPES buffer, 0.2 M, pH 7.2 (Sigma-Aldrich, catalog number: P6757 )
- Bovine serum albumin (BSA) (Sigma-Aldrich, catalog number: B4287 )
- Uranyl acetate (Polyscience, Inc., catalog number: 21447 )
- Lead nitrate (Electron Microscopy Sciences, catalog number: 17900 )
- Sodium citrate (Electron Microscopy Sciences, catalog number: 21140 )
- 1 N NaOH (Electron Microscopy Sciences, catalog number: 21170-01 )
- 0.01 M phosphate buffer (pH 7.2) (see Recipes)
- 0.05 M phosphate buffer (pH 7.2) (see Recipes)
- 2.5% glutaraldehyde (see Recipes)
- 2% aqueous osmium tetroxide (see Recipes)
- 0.02 M phosphate buffer (pH 7.2) (see Recipes)
- Antibodies (see Recipes)
- 2% PBS/BSA (see Recipes)
- 0.05 M PIPES buffer (pH 7.2) (see Recipes) (see Recipes)
- 2% uranyl acetate (see Recipes)
- 1 N NaOH (see Recipes)
- Reynold’s lead citrate (see Recipes)
- Scintillation vials with aluminum covered caps (Fisher Scientific, catalog number: 03-340-4B)
Equipment
- Immunogold labeling of microtubules and microtubule organizing center proteins
- Bunsen burner (Fisher Scientific, catalog number: S12809 )
- -20 °C freezer
- M2100 Benchmark Tube Rocker (Benchmark Scientific Inc.)
- Oven (General Signal, model: Gravity convection )
- Water bath (Sheldon Manufacturing, model: SWB23 )
- Micropipette (Bioexpress, GeneMate, catalog number: P-4963-20 )
- Centrifuge (Thermo Fisher Scientific, Thermo ScientificTM, catalog number: 75004061 )
- Diamond knife (Diatome, specs: Ultra, 45°, 4 mm, Wet)
- Transmission electron microscope (Hitachi, model: HF7100 )
- Bunsen burner (Fisher Scientific, catalog number: S12809 )
- Immunogold labeling of spermatozoid matrix and cell wall constituents
- Bunsen burner (Fisher Scientific, catalog number: S12809 )
- M2100 Benchmark Tube Rocker (Benchmark Scientific Inc.)
- Oven (General Signal, model: Gravity convection )
- Micropipette (Bioexpress, catalog number: P-4963-20 )
- Centrifuge (Thermo Fisher Scientific, Thermo ScientificTM, catalog number: 75004061 )
- Diamond knife (Diatome, specs: Ultra, 45°, 4 mm, Wet)
- Humid chamber (see Notes)
- Transmission electron microscope (Hitachi, model: HF7100 )
- Bunsen burner (Fisher Scientific, catalog number: S12809 )
Procedure
- Immunogold labeling of microtubules and microtubule organizing center proteins (Vaughn and Bowling, 2008; Vaughn, 2013)
Note: For each solution change described below, completely saturate samples by adding solution until it covers over the material in the vial/tube.- Fix small pieces of several antheridia or whole gametophyte plants (if small, like those of ferns) in 2.5% glutaraldehyde in 0.05 M Sorenson’s phosphate buffer (pH 7.2) at room temperature. Scintillation vials with aluminum-covered caps are ideal for this process (alternatively, you can also use 1.5 ml microcentrifuge tubes). If parts of plants are used, do all cutting of samples in a drop of the glutaraldehyde solution.
- Wash in the same 0.05 M Sorenson’s buffer, 2-3 exchanges, 15 min each at 4 °C. To facilitate exchanging solutions without losing the small samples, the tip of the Pasteur pipette is heated over a Bunsen burner so that the tip aperture is down to about 1/3 of its original size. In this way, clean exchanges may be made so that the tiny samples are not sucked up into the pipette and lost during the exchanges.
- Post-fix tissue for 10 min in 2% (v/v) osmium.
- Serial dehydration: Remove the PIPES buffer and add 25% ethanol (in 0.05 M PIPES buffer) to the vial, increasing the percentage of ethanol by 25% increments until 75% ethanol is reached.
- Exchange the solution with 100% ethanol several times. These steps should be at 4 °C, and the vials transferred to a -20 °C freezer after the second 100% ethanol transfer.
- Add LR White resin (London Resin Company) to reach 25% plastic (in ethanol). Shake vial to mix plastic into the solvent and maintain at -20 °C for one day. Repeat this procedure by adding plastic to 50% for one day and then add plastic to 75% for another day. Exchange the solutions, putting 100% resin in the vial. Shake the vial and make sure the samples have sunk in the resin. Allow them to spend 24 h in 100% resin.
- Allow the vials to warm to room temperature and shake on a rocking platform for 24 h. This will allow for infiltration of the resin throughout the sample.
- Transfer the samples to gelatin capsules and fill the capsule with 100% resin forming a meniscus. Rotate the cap as it is inserted to create an oxygen-excluding shield. Place in a 50 °C oven and remove in ~2 d.
- Trim the samples and cut 100 nm sections (pale gold reflectance colors) with a diamond knife and fume the sections with ethylene dichloride to stretch the sections. Pick up the sections by touching the dull side of 300 mesh gold grids or 200 mesh nickel grids.
- Place mounted grids in a closed CLEAN Petri dish and dry for 1-2 h at room temperature or place the Petri dish on a slide warmer (45-50 °C) for 30 min.
- Float the grids, section-side down on 10 µl drops of the following solutions (all of which are filtered through 250 nm filters) and times:
- 2% BSA in 0.02 M sodium phosphate buffer (PBS/BSA; pH 7.2), 30 min (BLOCKING).
- On drops of primary antibodies or monoclonal antibodies diluted from 1:20 in 2% PBS/BSA for 3-4 h at room temperature.
- 2% BSA in 0.02 M sodium phosphate buffer (PBS/BSA; pH 7.2), 30 min (BLOCKING).
- Rinse grids by placing them on drops of 2% BSA/PBS 4 x 3 min.
- Place grids on 5-15 µl drops of secondary gold-labeled antibody or Protein A/gold diluted 1/20-in 2% BSA/PBS in ‘humid chamber’ 1-3 h.
- Move grids to 5-10 µl drops of 0.02 M PBS for rinsing 4 x 3-5 min (FILTER and CENTRIFUGE).
- Rinse each grid with a jet of double distilled autoclaved water that has been filtered (0.2 µm) and let it dry.
- Post-stain the grids for 2 min in 2% uranyl acetate, wash in double distilled autoclaved water and post-stain in Reynold’s lead citrate. Wash in double distilled autoclaved water, dry and examine under transmission electron microscope.
- Fix small pieces of several antheridia or whole gametophyte plants (if small, like those of ferns) in 2.5% glutaraldehyde in 0.05 M Sorenson’s phosphate buffer (pH 7.2) at room temperature. Scintillation vials with aluminum-covered caps are ideal for this process (alternatively, you can also use 1.5 ml microcentrifuge tubes). If parts of plants are used, do all cutting of samples in a drop of the glutaraldehyde solution.
- Immunogold labeling of spermatozoid matrix and cell wall constituents (Lopez and Renzaglia, 2014)
- Fix small pieces of antheridial tissue or whole gametophyte plants (if small, like those of ferns) in 2.5% glutaraldehyde in 0.05 M Sorenson’s phosphate buffer (pH 7.2) at room temperature. Scintillation vials with aluminum-covered caps are ideal for this process (alternatively, you can also use 1.5 ml microcentrifuge tubes). If parts of plants are used, do all cutting of samples in a drop of the glutaraldehyde solution.
- Wash in the same 0.05 M Sorenson’s buffer, 2-3 exchanges, 15 min each at 4 °C. To facilitate exchanging solutions without losing the small samples, the tip of the Pasteur pipette is heated over a Bunsen burner so that the tip aperture is down to about 1/3 of its original size. In this way, clean exchanges may be made so that the tiny samples are not sucked up into the pipette during the exchanges.
- Post-fix tissue for 10 min in 1% (v/v) osmium tetroxide (OsO4).
- Rinse tissue 3 x in distilled water (10 min each).
- Serial dehydration: Remove the water and add 25% ethanol (in water) to the vial, increasing the percentage of ethanol by 25% increments until 75% ethanol is reached.
- Exchange the solution 3 x with 100% ethanol.
- Add LR White resin (London Resin Company) to reach 25% plastic (in ethanol). Shake vial to mix plastic into the solvent and maintain at -20 °C for one day. Repeat this procedure by adding plastic to 50% for one day and then add plastic to 75% for another day. Exchange the solutions, putting 100% resin in the vial. Shake the vial and make sure the samples have sunk in the resin. Allow them to spend 24 h in 100% resin.
- Allow the vials to warm to room temperature and shake on a rocking platform for 24 h. This will allow for infiltration of the resin throughout the sample.
- Transfer the samples to gelatin capsules and fill the capsule with 100% resin forming a meniscus. Rotate the cap as it is inserted to create an oxygen-excluding shield. Place in a 50 °C oven and remove in ~2 d.
- Trim the samples and cut 90-100 nm sections (pale gold reflectance colors) with a diamond knife and fume the sections with ethylene dichloride to stretch the sections. Pick up the sections by touching the dull side of 300 mesh gold grids or 200 mesh nickel grids.
- Place mounted grids on a clean piece of filter paper in a closed CLEAN Petri dish and dry for 1-2 h at room temperature or place the Petri dish on a slide warmer (45-50 °C) for 30 min.
- Using a humid chamber (see Note 2) place a piece of dental wax on completely wet filter paper, float the grids, section-side down on 10 µl drops in the following solutions (all of which are centrifuged and filtered through 250 nm filters) and times:
- 2% BSA in 0.05 M sodium phosphate buffer (BSA/PBS; pH 7.2), overnight (BLOCKING).
- On drops of primary antibodies diluted from 1:20 in 2% BSA/PBS for 3-4 h at room temperature. Alternatively, grids can be left in primary antibody overnight at 4 °C.
- 2% BSA in 0.05 M sodium phosphate buffer (BSA/PBS; pH 7.2), overnight (BLOCKING).
- Rinse grids by placing them on drops of 2% BSA/PBS 4 x 3 min.
- Place grids on 10 µl drops of secondary gold-labeled antibody diluted 1/20 in 2% BSA/PBS in ‘humid chamber’ 1-3 h.
- Move grids to 10 µl drops of 0.02 M PBS (FILTER and CENTRIFUGE) for rinsing 4 x 3-5 min.
- Rinse each grid with a jet of double distilled autoclaved water that has been filtered (0.2 µm) and let it dry.
- Post-stain the grids for 2 min in 2% uranyl acetate, wash in double distilled autoclaved water and post-stain in Reynold’s lead citrate for 30 sec. Wash in double distilled autoclaved water, dry and examine under transmission electron microscope.
- Fix small pieces of antheridial tissue or whole gametophyte plants (if small, like those of ferns) in 2.5% glutaraldehyde in 0.05 M Sorenson’s phosphate buffer (pH 7.2) at room temperature. Scintillation vials with aluminum-covered caps are ideal for this process (alternatively, you can also use 1.5 ml microcentrifuge tubes). If parts of plants are used, do all cutting of samples in a drop of the glutaraldehyde solution.
Notes
- Osmium tetroxide (OsO4) is a heavy fixative that stains lipids in membranous structures and vesicles. Additionally, the reduced heavy metal adds density and contrast to biological tissue.
- Humid chamber is a Petri dish, whose bottom is covered with a fresh, clean wet filter paper and cover is sealed with Parafilm if sitting overnight. Aliquots of BSA/PBS can be kept frozen until used. Centrifuge prior to use and store refrigerated (4 °C). 1-2% BSA/PBS can be divided in small aliquots (100-200 µl) for each time of labeling and kept at -20 °C, up to 6 months.
- The primary antibody in BSA/PBS should not be frozen and thawed repeatedly; it decreases the strength of enzyme.
- Use completely clean tools and Petri dishes during the labeling process.
- Millipore filter or centrifuge every single agent and only draw from supernatant, use sterile filtered (0.2 µm) water for rinses after PBS, lead citrate and uranyl acetate stains.
- Aliquots (40-50 µl) of primary antibody must be kept at -20 °C.
- Secondary antibody with gold particles must be kept at 4 °C, never place in freezer.
- After labeling agents are thawed (aliquots) do not freeze them again. They are good for maximum 3-4 days at 4 °C.
- Drop size for BSA/PBS as well as other liquids could be between 5 to 10 µl, if you are stopping for a step overnight use 10 µl.
- If you stop at any of the above steps (prior to putting grids in the secondary antibody) it should not be more than overnight and humid chamber should be sealed and kept in the refrigerator.
Note: Grids should not be left in the secondary antibody overnight because it will increase the amount of background ‘noise’ (i.e., over labels).
- It is recommended to wick grids on a small piece of filter paper prior to moving them to a new solution and between different steps of the same rinses.
- Use a new piece of Parafilm (or rinse and wipe dental wax) each time you move grids to a new solution.
- Glass Petri dishes are preferred. Plastic creates static electricity and causes the grids to cling to the top of the dish.
Recipes
- 0.01 M phosphate buffer (pH 7.2)
Add 5 parts 0.20 M Sorenson’s phosphate buffer (pH 7.2) to 95 parts double distilled autoclaved water
- 0.05 M phosphate buffer (pH 7.2)
Add 25 parts 0.20 M Sorenson’s phosphate buffer (pH 7.2) to 75 parts double distilled autoclaved water
- 2.5% glutaraldehyde
- Add 1 part 10% glutaraldehyde to 1 part double distilled autoclaved water
- Add 1 part 0.20 M Sorensen’s phosphate buffer (pH 7.2) to 1 part double distilled autoclaved water
- Add 1 part 5% glutaraldehyde to 1 part 0.10 M Sorensen’s phosphate buffer (pH 7.2)
- Add 1 part 10% glutaraldehyde to 1 part double distilled autoclaved water
- 2% aqueous osmium tetroxide
Add 1 part 4% aqueous osmium tetroxide to 1 part double distilled autoclaved water
- 0.02 M phosphate buffer (pH 7.2)
Add 1 part 0.20 M Sorenson’s phosphate buffer (pH 7.2) to 10 parts double distilled autoclaved water
- Antibodies
Add 1 part antibody to 20 parts 2% PBS/BSA
- 2% PBS/BSA
Add 0.20 g of bovine serum albumin to 10 ml 0.02 M phosphate buffer (pH 7.2)
- 0.05 M PIPES buffer (pH 7.2)
Add 1 part 0.30 M PIPES buffer (pH 7.2) to 6 parts double distilled autoclaved water
- 2% uranyl acetate
Add 1 g uranyl acetate in 50 ml double distilled autoclaved water
- 1 N NaOH
Dissolve 40.00 g sodium hydroxide in water to make volume 1 L
- Reynold’s lead citrate
- Add 1.33 g lead nitrate and 1.76 g sodium citrate to 30 ml boiled double distilled autoclaved water that has cooled to room temperature and shake for 30 min
- Add 8 ml 1 N NaOH
- Add 12 ml double distilled autoclaved water
- Add 1.33 g lead nitrate and 1.76 g sodium citrate to 30 ml boiled double distilled autoclaved water that has cooled to room temperature and shake for 30 min
Acknowledgments
This research was supported by research grants (DEB-0322664, DEB-0423625, DEB0521177, and DEB-0228679) from the National Science Foundation as part of the Research Experience for Undergraduates and Assembling the Tree of Life Programs.
The Authors declare we have no conflicts of interest or competing interest.
References
- Davis, F. M., Tsao, T. Y., Fowler, S. K. and Rao, P. N. (1983). Monoclonal antibodies to mitotic cells. Proc Natl Acad Sci U S A 80(10): 2926-2930.
- Hepler, P. K. (1976). The blepharoplast of Marsilea: its de novo formation and spindle association. J Cell Sci 21(2): 361-390.
- Hoffman, J. C. and Vaughn, K. C. (1995). Using the developing spermatogenous cells of Ceratopteris to unlock the mysteries of the plant cytoskeleton. Int J Plant Sci 156(3): 346-358.
- Hoffman, J. C., Vaughn, K. C. and Joshi, H. C. (1994). Structural and immunocytochemical characterization of microtubule organizing centers in pteridophyte spermatogenous cells. Protoplasma 179(1): 46-60.
- Joshi, H. C., Palacios, M. J., McNamara, L. and Cleveland, D. W. (1992). γ-Tubulin is a centrosomal protein required for cell cycle-dependent microtubule nucleation. Nature 356(6364): 80-83.
- Klink, V. P. and Wolniak, S. M. (2003). Changes in the abundance and distribution of conserved centrosomal, cytoskeletal and ciliary proteins during spermiogenesis in Marsilea vestita. Cell Motil Cytoskeleton 56 (1): 57-73.
- Knox, J. P., Linstead, P. J., King, J., Cooper, C., Roberts, K. (1990). Pectin esterification is spatially regulated both within cell walls and between developing tissues of root apices. Planta 181(4): 512-521.
- Lamport, D. T. and Várnai, P. (2013). Periplasmic arabinogalactan glycoproteins act as a calcium capacitor that regulates plant growth and development. New Phytol 197(1): 58-64.
- Liu, B., Joshi, H. C., Wilson, T. J., Silflow, C. D., Palevitz, B. A. and Snustad, D. P. (1994). gamma-Tubulin in Arabidopsis: gene sequence, immunoblot, and immunofluorescence studies. Plant Cell 6(2): 303-314.
- Liu, B., Marc, J., Joshi, H. C. and Palevitz, B. A. (1993). A gamma-tubulin-related protein associated with the microtubule arrays of higher plants in a cell cycle-dependent manner. J Cell Sci 104(4): 1217-1228.
- Lopez, R. A. and Renzaglia, K. S. (2014). Multiflagellated sperm cells of Ceratopteris richardii are bathed in arabinogalactan proteins throughout development. Am J Bot 101(12): 2052-2061.
- Lopez, R. A. and Renzaglia, K. S. (2017). The Ceratopteris (fern) developing motile gamete walls contain diverse polysaccharides but not pectin. Planta.
- Lopez-Swalls and Renee A. (2016). The special walls around gametes in Ceratopteris richardii and Aulacomnium palustre: using immunocytochemistry to expose structure, function, and development. Southern Illinois University Carbondale.
- Manandhar, G., Simerly, C., Salisbury, J. L. and Schatten, G., (1999). Centriole and centrin degeneration during mouse spermiogenesis. Cytoskeleton 43(2): 137-144.
- Marcus, S. E., Verhertbruggen, Y., Hervé, C., Ordaz-Ortiz, J. J., Farkas, V., Pedersen, H. L., Willats, W. G., Knox, J. P. (2008). Pectic homogalacturonan masks abundant sets of xyloglucan epitopes in plant cell walls. BMC Plant Biol 8(1): 60.
- Meikle, P. J., Bonig, I., Hoogenraad, N. J., Clarke, A. E. and Stone, B. A. (1991). The location of (1→3)-β-glucans in the walls of pollen tubes of Nicotiana alata using a (1→3)-β-glucan-specific monoclonal antibody. Planta 185(1): 1-8.
- Murata, T. and Hasebe, M. (2007). Microtubule-dependent microtubule nucleation in plant cells. J Plant Res 120 (1): 73-78.
- Murata, T., Sonobe, S., Baskin, T. I., Hyodo, S., Hasezawa, S., Nagata, T., Horio, T. and Hasebe, M. (2005). Microtubule-dependent microtubule nucleation based on recruitment of gamma-tubulin in higher plants. Nat Cell Biol 7(10): 961-968.
- Oakley, B. R., Oakley, C. E., Yoon, Y. and Jung, M. K. (1990). γ-Tubulin is a component of the spindle pole body that is essential for microtubule function in Aspergillus nidulans. Cell 61(7): 1289-1301.
- Pedersen, H. L., Fangel, J. U., McCleary, B., Ruzanski, C., Rydahl, M. G., Ralet, M. C., Farkas, V., von Schantz, L., Marcus, S. E., Andersen, M. C. and Field, R. (2012). Versatile high resolution oligosaccharide microarrays for plant glycobiology and cell wall research. J Biol Chem 287(47): 39429-39438.
- Pennell, R. I., Janniche, L., Kjellbom, P., Scofield, G. N., Peart, J. M. and Roberts, K. (1991). Developmental regulation of a plasma membrane arabinogalactan protein epitope in oilseed rape flowers. Plant Cell 3(12): 1317-1326
- Renzaglia, K. S. and Garbary, D. J. (2001). Motile male gametes of land plants: Diversity, development, and evolution. Crit Rev Sci 20(2):107-213.
- Satisbury, J. L. (1995). Centrin, centrosomes, and mitotic spindle poles. Curr Opin Cell Biol 7(1): 39-45.
- Showalter, A. M. (2001). Arabinogalactan-proteins: structure, expression and function. Cell Mol Life Sci 58(10): 1399-1417.
- Vandre, D. D., Davis, F. M., Rao, P. N. and Borisy, G. G. (1984). Phosphoproteins are components of mitotic microtubule organizing centers. Proc Natl Acad Sci U S A 81(14): 4439-4443.
- Vaughn, K. (2013). Immunocytochemical techniques. In: Immunocytochemistry of Plant Cells. Springer pp: 1-41.
- Vaughn, K. C. and Bowling, A. J. (2008). Recovery of microtubules on the blepharoplast of Ceratopteris spermatogenous cells after oryzalin treatment. Protoplasma 233 (3-4): 231-240.
- Vaughn, K. C. and Harper, J. D. (1998). Microtubule-organizing centers and nucleating sites in land plants. Int Rev Cytol 181: 75-149.
- Vaughn, K. C. and Renzaglia, K. S. (1993). Centrin in spermatogenesis of archegoniates. Am J Bot (Suppl.) 80(1): 58-66.
- Vaughn, K. C. and Renzaglia, K. S. (2006). Structural and immunocytochemical characterization of the Ginkgo biloba L. sperm motility apparatus. Protoplasma 227(2-4):165-173.
- Verhertbruggen, Y., Marcus, S. E., Haeger, A., Ordaz-Ortiz, J. J. and Knox, J. P. (2009). An extended set of monoclonal antibodies to pectic homogalacturonan. Carbohyd Res 344(14): 1858−186.
- Yates, E. A., Valdor, J-F., Haslam, S. M., Morris, H. R., Dell, A., Mackie, W. and Knox, J. P. (1996). Characterization of carbohydrate structural features recognized by anti-arabinogalactan-protein monoclonal antibodies. Glycobiology 6(2): 131-139.
Article Information
Copyright
© 2017 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Lopez, R. A., Mansouri, K., Henry, J. S., Flowers, N. D., Vaughn, K. C. and Renzaglia, K. S. (2017). Immunogold Localization of Molecular Constituents Associated with Basal Bodies, Flagella, and Extracellular Matrices in Male Gametes of Land Plants. Bio-protocol 7(21): e2599. DOI: 10.21769/BioProtoc.2599.
- Renzaglia, K. S., Villarreal, J. C., Piatkowski, B. T., Lucas, J. R. and Merced, A. (2017). Hornwort Stomata: Architecture and Fate Shared with 400-Million-Year-Old Fossil Plants without Leaves. Plant Physiol 174(2): 788-797.
Category
Cell Biology > Cell imaging > Electron microscopy
Cell Biology > Cell imaging > Fixed-cell imaging
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link