- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Nematode Epicuticle Visualisation by PeakForce Tapping Atomic Force Microscopy
Published: Vol 7, Iss 21, Nov 5, 2017 DOI: 10.21769/BioProtoc.2596 Views: 8530
Reviewed by: Neelanjan BoseAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Preparation of Onion Epidermal Cell Walls for Imaging by Atomic Force Microscopy (AFM)
Tian Zhang and Daniel J. Cosgrove
Dec 20, 2017 15140 Views

Characterization of Amyloid Fibril Networks by Atomic Force Microscopy
Mirren Charnley [...] Nicholas P. Reynolds
Feb 20, 2018 9039 Views

High-speed Atomic Force Microscopy Observation of Internal Structure Movements in Living Mycoplasma
Kohei Kobayashi [...] Makoto Miyata
Mar 5, 2022 3035 Views
Abstract
The free-living soil nematode Caenorhabditis elegans has become an iconic experimental model animal in biology. This transparent animal can be easily imaged using optical microscopy to visualise its organs, tissues, single cells and subcellular events. The epicuticle of C. elegans nematodes has been studied at nanoscale using transmission and scanning (SEM) electron microscopies. As a result, imaging artefacts can appear due to embedding the worms into resins or coating the worms with a conductive gold layer. In addition, fixation and contrasting may also damage the cuticle. Conventional tapping mode atomic force microscopy (AFM) can be applied to image the cuticle of the dried nematodes in air, however this approach also suffers from imaging defects. Ideally, the nematodes should be imaged under conditions resembling their natural environment. Recently, we reported the use of PeakForce Tapping AFM mode for the successful visualisation and numerical characterisation of C. elegans nematode cuticle both in air and in liquid (Fakhrullina et al., 2017). We imaged the principal nematode surface structures and characterised mechanical properties of the cuticle. This protocol provides the detailed description of AFM imaging of liquid-immersed C. elegans nematodes using PeakForce Tapping atomic force microscopy.
Keywords: Atomic force microscopyBackground
Nematodes, both free-living and parasitic, have been extensively studied due to their remarkable biology and effects on agriculture and human well-being. Apparently, the most studied and famous species among nematodes is a free-living soil nematode Caenorhabditis elegans (Sterken et al., 2015). This round worm has been successfully employed as a versatile model organism in a number of investigations (Fire et al., 1998; Kenyon, 2010; Swierczek et al., 2011; O’Reilly et al., 2014; Stroustrup et al., 2016), securing eventually a Nobel prize for Sidney Brenner, John Sulston and Robert Horvitz in 2002. C. elegans nematode is a tiny (~1 mm long) transparent animal which can be easily visualised using an optical microscope. Its cuticle, though thin and transparent, serves as a principal protective barrier between the worm and its habitat. Cuticle is an important marker of disease, when pathogenic bacteria colonize it, leading to the death of the infected animal. In addition, the structure of the cuticle may indicate the ageing in the nematodes. From the human health care and industrial point of view, monitoring of the cuticle of nematodes might be helpful in elucidation of novel antinematode chemicals affecting the cuticle. As a result, nanoscale imaging of the nematode epicuticle in its natural liquid environments may open new avenues in biomedical research.
Previously, the cuticle of nematodes (mostly employing C. elegans as a model organism) has been studied using electron microscopy, both transmission and scanning. Unfortunately, electron microscopy does not allow imaging the worms in liquid, which is their native environment. Sample preparation for electron microscopy requires fixation, dehydration, contrast staining or surface sputtering, and also resin embedding for ultrathin slicing. As a result, nematode cuticle may exhibit secondary artifacts preventing from evaluation of native surface structure and mechanical characterisation. Atomic force microscopy has been established as a potent tool in imaging biological samples in situ, including imaging live cells in liquid media (Beaussart et al., 2015). Recently, the application of tapping mode AFM to visualise C. elegans nematodes in air has been reported (Allen et al., 2015). Imaging in air suffered from the same drawback as scanning electron microscopy, for example the images demonstrated the typical shrunk and collapsed cuticle surfaces apparently caused by dehydration and air imaging. We envisaged a different technique, based on PeakForce Tapping AFM mode (Alsteens et al., 2012) for successful visualisation of C. elegans nematode cuticle in water (Fakhrullina et al., 2017). Here we report a detailed protocol for this technique.
Materials and Reagents
Note: All chemicals were purchased from Sigma-Aldrich unless noted otherwise.
- Pipette tips
- Petri dishes
- Dust-free Nexterion glass slides (Schott)
- Wild type C. elegans (N2 Bristol) nematodes
- Escherichia coli OP50 bacteria
- Poly(allylamine hydrochloride) (PAH, molecular weight ~17,5 kDa) (Sigma-Aldrich, catalog number: 283215 )
- Poly(sodium 4-styrenesulfonate) (PSS, molecular weight ~70 kDa) (Sigma-Aldrich, catalog number: 243051 )
- Nematode growth media (NGM) (bioWORLD, catalog number: 30620040-1 )
- Nematode lysis solution (aqueous 2% NaOCl/0.45 M NaOH)
- Sodium hypochlorite (NaClO) (Sigma-Aldrich, catalog number: 425044 )
- Sodium hydroxide (NaOH) (Sigma-Aldrich, catalog number: S8045 )
- Levamisole hydrochloride (Sigma-Aldrich, catalog number: L9756 )
- Glutaraldehyde (25% aqueous) (Sigma-Aldrich, catalog number: G5882 )
- Ultrapure (type 1) water purified by Simplicity (Millipore) water purification system
- Potassium phosphate monobasic (KH2PO4) (Sigma-Aldrich, catalog number: NIST200B )
- Sodium phosphate dibasic (Na2HPO4) (Sigma-Aldrich, catalog number: 71649 )
- Sodium chloride (NaCl) (Sigma-Aldrich, catalog number: S7653 )
- Magnesium sulfate (MgSO4) (Sigma-Aldrich, catalog number: 793612 )
- Aqueous M9 buffer (see Recipes)
Equipment
- Pipettes (mechanical Eppendorf or Gilson pipettes)
- Eppendorf Scientific Excella E24 temperature-controlled benchtop shaker (Eppendorf, New BrunswickTM, model: Excella® E24 )
- Biosan V-1 plus vortex (Biosan, model: V-1 plus )
- Nikon SMZ 745 T stereomicroscope (Nikon, model: SMZ745T )
- Dimension FastScan Bio atomic force microscope (Bruker, model: Dimension FastScan ) equipped with a liquid submergible scanner
Note: Do not use air scanners for scanning in liquid, this may damage the instrument. - Biosan Microspin 12 centrifuge (Biosan, model: Microspin 12 )
- ScanAsyst-Fluid probes (nominal length 70 µm, tip radius 20 nm, spring constant of 0.7 N m-1) (Bruker, model: SCANASYST-FLUID )
Note: Other AFM probes from Bruker or other producers approved for using in PeakForce Tapping mode having characteristics similar to ScanAsyst-Fluid probes can also be used.
Software
- NanoScope AFM operating software (Bruker Corporation)
- NanoScope Analysis v.1.7. software (Bruker Corporation)
Procedure
- Fabrication of layer-by-layer (LbL) polyelectrolyte coated glass substrates
- Prepare 5 mg ml-1 aqueous solutions of PAH and PSS polymers (5 ml each).
- Drop 50 µl of PAH solution onto the central part of a dust-free glass slide, incubate for 20 min in a humidified chamber (any tightly closed glass or plastic container with moist cotton pad might be used to prevent the premature evaporation), wash several times with water.
- Repeat the same using PSS solution.
- Repeat steps A2 and A3 until the resulting coating architecture is (PAH/PSS)12PAH (12 consecutive PAH/PSS bilayers followed by a final PAH layer). The outmost polyelectrolyte should be PAH to ensure the positive charge of the deposited film. More information on fabrication of LbL films can be found elsewhere (Lvov et al., 1995).
- Prepare 5 mg ml-1 aqueous solutions of PAH and PSS polymers (5 ml each).
- Caenorhabditis elegans nematodes cultivation
- Cultivate wild type C. elegans (N2 Bristol) nematodes on NGM-supplemented agar plates at 20 °C using a thermostated shaker according to a standard protocol (Brenner, 1974). Use Escherichia coli OP50 bacteria to feed worms. Observe the animals using a stereomicroscope after 48 h to detect gravid nematodes.
- Wash off gravid adult hermaphrodites with M9 buffer (see Recipes) into a tube and harvest by centrifugation for 2 min (1,100 x g, applicable to eggs, larvae and adult hermaphrodites). Dissolve the animals using aqueous 2% NaOCl/0.45 M NaOH solution (4.5 ml) to obtain nematode eggs. Wash the eggs with M9 buffer several times, then collect by centrifugation as described above and place onto the sterile NGM agar plates.
- Cultivate the nematodes at 20 °C to obtain synchronised adult hermaphrodites. Alternatively, cultivate the worms for shorter time to obtain larvae animals (Brenner, 1974).
- Collect the nematodes as described in step B2. Euthanise the collected worms with 45 mM aqueous levamisole hydrochloride. Wash consequently with water.
- Fixate the animals using 2 ml of buffered glutaraldehyde (in M9 buffer) for 2 h at 25 °C, then wash with water.
- Cultivate wild type C. elegans (N2 Bristol) nematodes on NGM-supplemented agar plates at 20 °C using a thermostated shaker according to a standard protocol (Brenner, 1974). Use Escherichia coli OP50 bacteria to feed worms. Observe the animals using a stereomicroscope after 48 h to detect gravid nematodes.
- Atomic force microscope setup
- It is assumed that the AFM operator is familiar with the AFM working principles and has previous experience with Dimension FastScan Bio AFM. For introduction into PeakForce Tapping AFM mode refer to microscope manual or to the following publication (Durkovic et al., 2014)
- AFM should be initiated before the sample is ready for imaging (see below). Ideally, these steps should be performed at the same time.
- Switch on the microscope and the operating software.
- Install and calibrate the mechanical properties of the probe (ScanAsyst-Fluid) according to the manufacturer’s manual. Start calibrating in air, then repeat in water.
- It is assumed that the AFM operator is familiar with the AFM working principles and has previous experience with Dimension FastScan Bio AFM. For introduction into PeakForce Tapping AFM mode refer to microscope manual or to the following publication (Durkovic et al., 2014)
- Atomic force microscopy imaging
- Place a droplet (100 µl) of water-suspended fixated nematodes onto the LbL-coated glass slides.
- Wait 30 min for settling and attaching the animals to the LbL polyelectrolyte film.
- Remove the excess water from the glass surface by pipetting, then wash the slide with water 3 times to remove any loosely immobilised worms.
- Add 200 µl of water on the top of the LbL-deposited film retaining the worms. The adhesion of the animals to the glass slide can be checked using a stereomicroscope prior to AFM imaging.
- Secure the glass slide on the microscope stage using vacuum suction. This should be done with due care to avoid removing the liquid from the glass slide.
- Locate a worm using the internal AFM optical camera. The worms will appear on the computer screen connected to the AFM internal camera as they would be seen under a stereomicroscope (Figure 1).
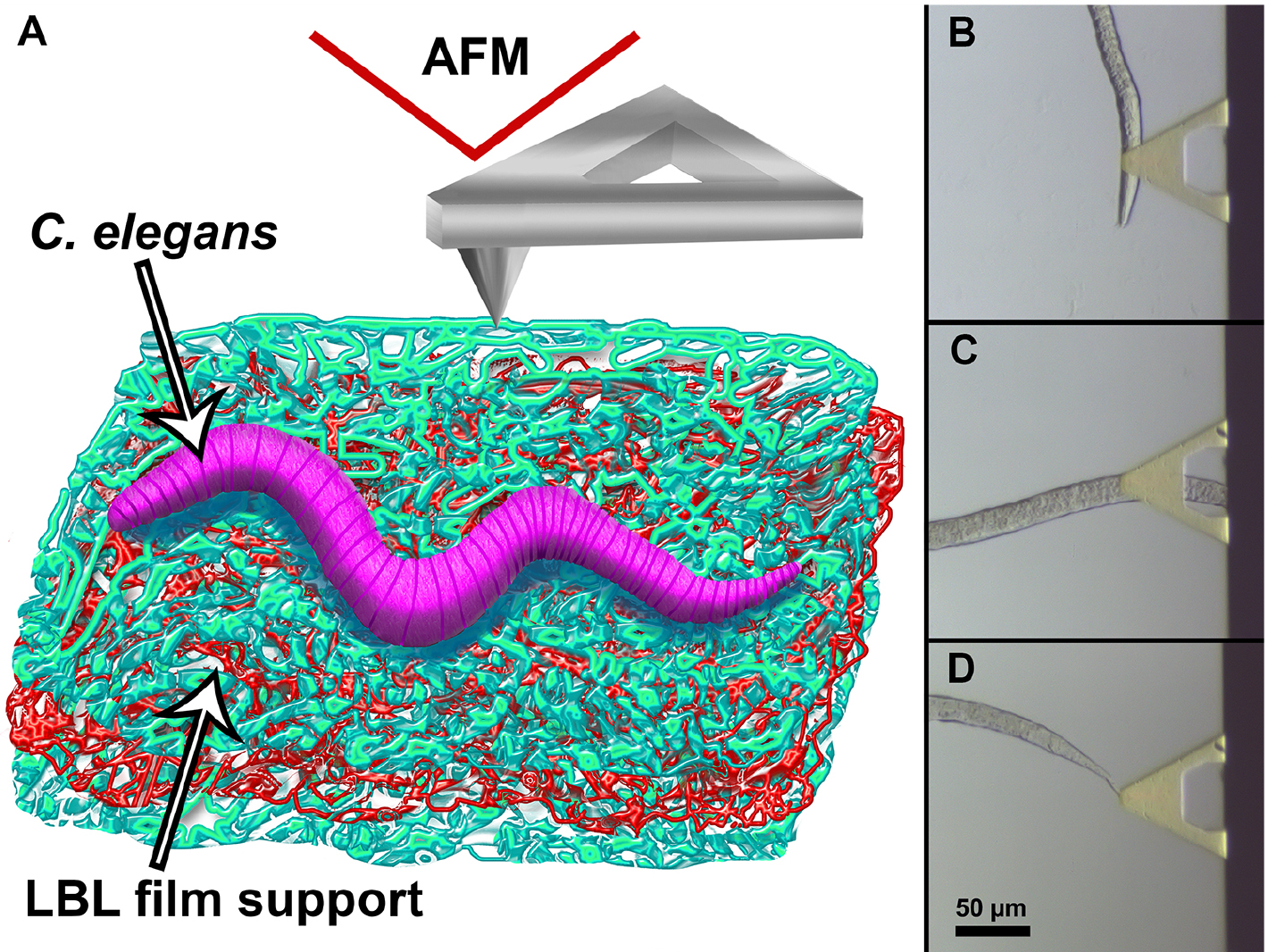
Figure 1. Immobilisation of nematodes on the support film. A sketch demonstrating immobilisation of an adult C. elegans nematode on (PAH/PSS)12PAH LbL-deposited film and subsequent AFM imaging in liquid (A). Optical microscopy camera snapshot demonstrating the approach of the AFM probe to head (B), body (C) and tail (D) region of an immobilised C. elegans adult hermaphrodite. - Position the probe above a certain area of the worm (i.e., head, tail or body regions).
- Approach the tip to the surface. Use relatively low scanning resolution (i.e., 64 lines) for locating the cuticle surface.
- After locating the region of interest (ROI), reduce the scanning frequency (0.8-0.9 Hz) and peak force setpoint (1 nN); increase the scanning resolution (up to 512 lines). The typical AFM images of an adult C. elegans nematode are shown in Figure 2.
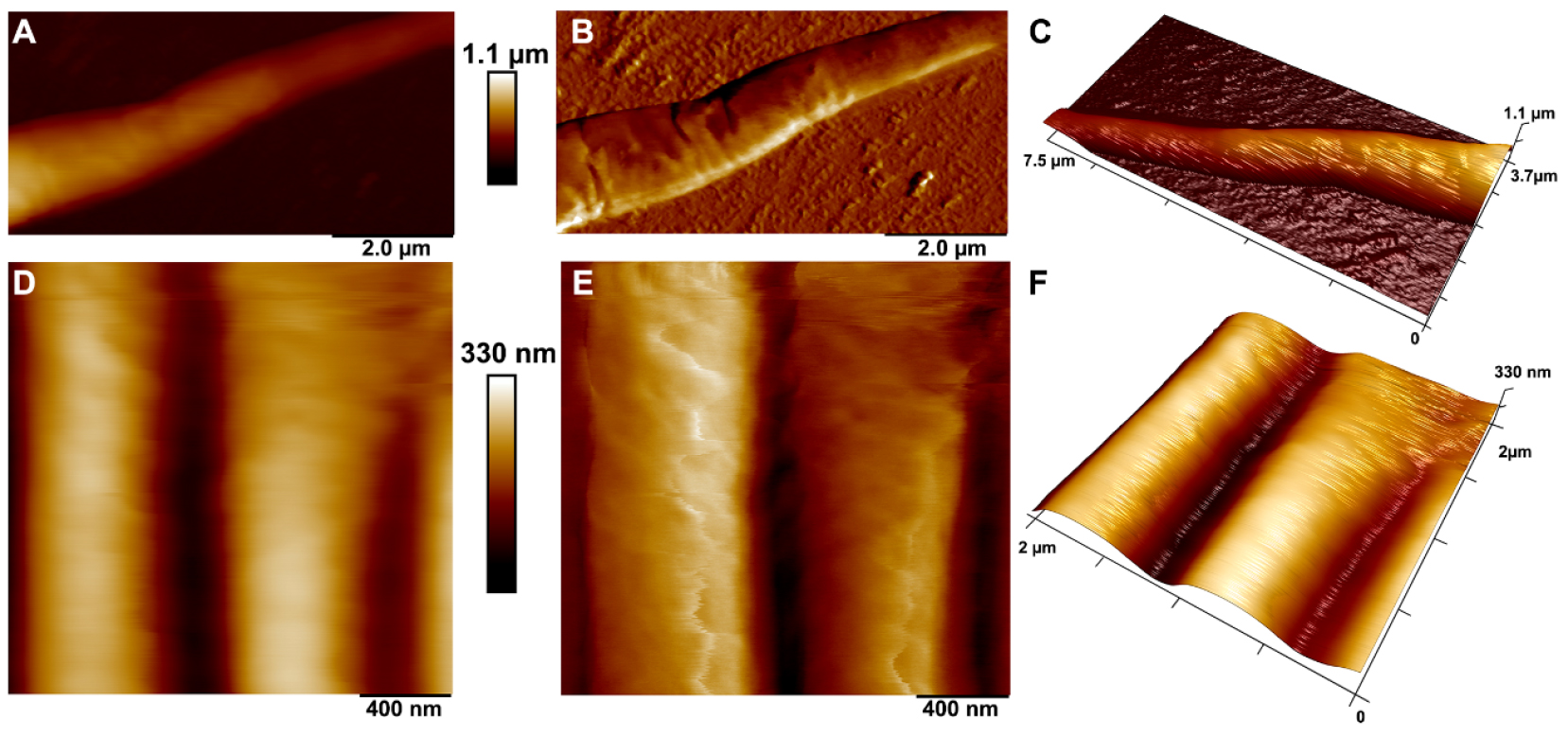
Figure 2. PeakForce Tapping AFM images of an adult C. elegans nematode inmmobilised on (PAH/PSS)12PAH LbL-deposited film: tail region (upper row) and body region (lower row). A and D. Height sensor topography images; B and E. Peak force error images; C and F. 3-D rendering of height sensor data with peak force error data overlayed. - Collect images in the following channels: height sensor, peak force error, adhesion and Sneddon modulus mode. Figure 3 demonstrates the images obtained in all four channels.

Figure 3. PeakForce Tapping AFM images of L1 larvae (A) and adult C. elegans (B) nematode imaged in water in topography (height sensor), peak force error, adhesion and modulus (Sneddon model) channels. Scale bars = 1 µm. - Scans can be obtained using higher aspect ratios (i.e., 5-6), which allows increasing the longitude of the images (compare aspect ratios in Figures 2 and 3).
- For statistically-significant quantitative evaluation examine 5-10 individual randomly selected nematodes.
- Place a droplet (100 µl) of water-suspended fixated nematodes onto the LbL-coated glass slides.
- Atomic force microscopy raw data processing
- Export raw data files to NanoScope Analysis software.
- Open a file, then flatten the image using 3d flattening order.
- Adjust colour scale using ‘Color Bar Scale Relative to Minimum Data Cursor’ function.
- Export high-resolution tiff files using Journal Quality Export function.
- For advanced image processing and nanomechanical characterisation refer to the NanoScope Analysis software operation manual.
- Export raw data files to NanoScope Analysis software.
Recipes
- M9 buffer
22 mM KH2PO4
42 mM Na2HPO4
85.5 mM NaCl
1 mM MgSO4
Acknowledgments
This study was performed according to the Russian Government Program of Competitive Growth of Kazan Federal University. The authors acknowledge the RFBR 17-04-02182А grant. This protocol was adapted from our previous publication (Fakhrullina et al., 2017). We thank Ms. E. Dubkova for technical help. The authors have no conflicts of interest.
References
- Allen, M. J., Kanteti, R., Riehm, J. J., El-Hashani, E. and Salgia, R. (2015). Whole-animal mounts of Caenorhabditis elegans for 3D imaging using atomic force microscopy. Nanomedicine 11(8): 1971-1974.
- Alsteens, D., Dupres, V., Yunus, S., Latge, J. P., Heinisch, J. J. and Dufrene, Y. F. (2012). High-resolution imaging of chemical and biological sites on living cells using peak force tapping atomic force microscopy. Langmuir 28(49): 16738-16744.
- Beaussart, A., El-Kirat-Chatel, S., Fontaine, T., Latge, J. P. and Dufrene, Y. F. (2015). Nanoscale biophysical properties of the cell surface galactosaminogalactan from the fungal pathogen Aspergillus fumigatus. Nanoscale 7(36): 14996-15004.
- Brenner, S. (1974). The genetics of Caenorhabditis elegans. Genetics 77(1): 71-94.
- Durkovic, J., Kardosova, M. and Lagana, R. (2014). Imaging and measurement of nanomechanical properties within primary xyleme cell walls of broadleaves. Bio Protoc 4 (24) e1360.
- Fakhrullina, G., Akhatova, F., Kibardina, M., Fokin, D. and Fakhrullin, R. (2017). Nanoscale imaging and characterisation of Caenorhabditis elegans epicuticle using atomic force microscopy. Nanomedicine 13(2): 483-491.
- Fire, A., Xu, S., Montgomery, M. K., Kostas, S. A., Driver, S. E. and Mello, C. C. (1998). Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391(6669): 806-811.
- Kenyon, C. J. (2010). The genetics of ageing. Nature 464(7288): 504-512.
- Lvov, Y., Ariga, K., Ichinose, I. and Kunitake, T. (1995). Assembly of multicomponent protein films by means of electrostatic layer-by-layer adsorption. J Am Chem Soc 117(22): 6117-6123.
- O’Reilly, L. P., Luke, C. J., Perlmutter, D. H., Silverman, G. A. and Pak, S. C. (2014). C. elegans in high-throughput drug discovery. Adv Drug Deliv Rev 69-70: 247-253.
- Sterken, M. G., Snoek, L. B., Kammenga, J. E. and Andersen, E. C. (2015). The laboratory domestication of Caenorhabditis elegans. Trends Genet 31(5): 224-231.
- Stroustrup, N., Anthony, W. E., Nash, Z. M., Gowda, V., Gomez, A., Lopez-Moyado, I. F., Apfeld, J. and Fontana, W. (2016). The temporal scaling of Caenorhabditis elegans ageing. Nature 530(7588): 103-107.
- Swierczek, N. A., Giles, A. C., Rankin, C. H. and Kerr, R. A. (2011). High-throughput behavioral analysis in C. elegans. Nat Methods 8(7): 592-598.
Article Information
Copyright
© 2017 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Akhatova, F., Fakhrullina, G., Gayazova, E. and Fakhrullin, R. (2017). Nematode Epicuticle Visualisation by PeakForce Tapping Atomic Force Microscopy. Bio-protocol 7(21): e2596. DOI: 10.21769/BioProtoc.2596.
Category
Cell Biology > Cell imaging > Atomic force microscopy
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link











