- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Labelling HaloTag Fusion Proteins with HaloTag Ligand in Living Cells
Published: Vol 7, Iss 17, Sep 5, 2017 DOI: 10.21769/BioProtoc.2526 Views: 12260
Reviewed by: Arsalan DaudiAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
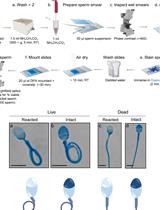
In vitro Induction and Detection of Acrosomal Exocytosis in Human Spermatozoa
Shenae L. Cafe [...] Brett Nixon
Jul 20, 2020 4804 Views
A New Method for Studying RNA-binding Proteins on Specific RNAs
Weiping Sun [...] Min Zhuang
May 20, 2021 10622 Views

Flow Cytometry Analysis of Microglial Phenotypes in the Murine Brain During Aging and Disease
Jillian E. J. Cox [...] Sarah R. Ocañas
Jun 20, 2024 3642 Views
Abstract
HaloTag has been widely used to label proteins in vitro and in vivo (Los et al., 2008). In this protocol, we describe labelling HaloTag-Cbx fusion proteins by HaloTag ligands for live-cell single-molecule imaging (Zhen et al., 2016).
Keywords: HaloTagBackground
Molecular processes of living organisms are intrinsically dynamics. Direct observation of the molecular processes in living cells is critical for quantitatively understanding of how biological systems function. Recent advances in fluorescence microscopy and fluorescent labelling enable to visualize trajectories of individually single molecules in living cells, providing insights about dynamic interactions and assemblies of biological molecules (Kusumi et al., 2014; Liu et al., 2015; Tatavosian et al., 2015; Cuvier and Fierz, 2017). Specific labelling of biomolecules with fluorophores is the key for fluorescence single-molecule imaging. HaloTag is self-labeling tag proteins that can be coupled to synthetic dyes in living cells (Los et al., 2008). The reaction occurs rapidly in living cells and the formed covalent bond is specific and irreversible. This technique has been utilized to study the genetic information flow in vivo, and to measure the kinetic of gene regulation in living mammalian cells (Liu et al., 2015; Zheng and Lavis, 2017). Janelia FluorTM dyes, such as Janelia FluorTM 549 (JF549), are bright and photostable fluorescent HaloTag ligands (Grimm et al., 2015). This protocol describes how to label HaloTag-Cbx proteins with JF549 for live-cell single-molecule imaging, which was developed in the recent publication (Zhen et al., 2016).
Materials and Reagents
- Pipette tips (BioExpress, catalog number: P-1236-200)
Manufacturer: Biotix, catalog number: P-1236-200CS . - 35 mm glass bottom dish made in the laboratory (see Video 1 for making glass-bottom dishes)Video 1. Making glass-bottom dishes. The video elaborates how to make glass-bottom dishes for live-cell single-molecule imaging.
- Cell lines used in protocol: mouse embryonic stem cells and HEK293T cells
- Janelia Fluor 549 dye (JF549) provided by Dr. Luke D. Lavis (Janelia Research Campus, Howard Hughes Medical Institute)
- Trypsin-EDTA (Thermo Fisher Scientific, GibcoTM, catalog number: 25300054 )
- Phosphate-buffered saline (PBS) (Sigma-Aldrich, catalog number: D8537-500ML )
- Gelatin (Sigma-Aldrich, catalog number: G1890-100G )
- DMEM (Sigma-Aldrich, catalog number: D5796 )
- Fetal bovine serum (FBS) (Sigma-Aldrich, catalog number: F0926 )
- Glutamine (Thermo Fisher Scientific, GibcoTM, catalog number: 25030081 )
- Penicillin-streptomycin (Thermo Fisher Scientific, GibcoTM, catalog number: 15140122 )
- β-Mercaptoethanol (Thermo Fisher Scientific, GibcoTM, catalog number: 21985023 )
- Non-essential amino acids (Thermo Fisher Scientific, catalog number: 1114050 )
- Leukemia inhibitor factor (LIF, made in the laboratory)
- FluoroBrite DMEM (Thermo Fisher Scientific, GibcoTM, catalog number: A1896701 )
- ES cell medium (see Recipes)
- Live-cell imaging medium (see Recipes)
Equipment
- Single channel pipette (BioExpress, Kaysville, USA)
- Heater controller (Warner Instruments, catalog number: TC-324 )
- Microscope (Manual Microscopy) (ZEISS, model: Axio Observer D1 )
- Alpha Plan-Apochromatic 100/1.46 NA Oil-immersion Objective (ZEISS, Germany)
- Evolve 512 x 512 EMCCD camera (Photometrics, Tucson, USA)
- Solid state laser (Intelligent Imaging Innovations, 3i LaserStack with Fiber)
Software
- Slidebook 6.0 software (Intelligent Imaging Innovations, Denver, Colorado)
- MATLAB R2015a (8.5.0.197613) (MathWorks, Natick, USA)
- U-track 2.0 (Danusar Lab, UT South Western Medical Center, Dallas, USA)
Procedure
- Trypsinize 70-90% confluent cells (mouse embryonic stem cells or HEK293 cells) of 100 mm plate stably expressing HaloTag-Cbx proteins (we recommend 0.6 ml of 0.05% trypsin-EDTA [1x] for 60 mm plate and 1.5 ml for 100 mm plate).
- Seed 20% of cells to 35 mm glass-bottom dish coated by Gelatin overnight (Note 1) (see Video 2 for gelatinization). Video 2. Gelatinization of glass-bottom dishes. The video describes how to gelatinize glass-bottom dishes before seeding cells.
- Following by overnight culture, the final confluency of the cells before was between 80-90%. Several concentrations (5 nM, 15 nM, and 30 nM) of JF549 are used to incubate with cells for 15 min at 37 °C in 5% CO2 (Notes 2 and 3) (see Video 3 for adding the JF549 dye). Video 3. Adding JF549 dyes to cells
- Gently wash cells with ES medium (see Recipes) once and incubate in ES medium for 30 min at 37 °C and 5% CO2 (see Video 4 for washing cells). Video 4. Washing cells with ES cell medium
- Replace ES medium with live-cell imaging medium (see Recipes) (see Video 5 for adding live-cell imaging medium).Video 5. Replacing ES cell medium with live-cell imaging medium
- Maintain 37 °C conditions during imaging by using a heater controller. Each plate should be imaged for the maximum of 1.5 h after placing on the microscope (see Video 6 for placing dishes on objective).Video 6. Placing dishes on objective
- The number of individual fluorescent spot per nucleus should be between 10-50 spots, controlled by adjusting the JF549 dye concentrations (Notes 4 and 5) (see Video 7 for single-molecule imaging).Video 7. Single-molecule imaging. The video describes how to image individual HaloTag-Cbx proteins within living cells.
- Movies are then uploaded to u-track 2.0. Each cell is cropped from the larger movie. Cropped movies are processed (Note 6).
Data analysis
- Our data were analyzed using MATLAB with u-track 2.0 plug-in, detail guide can be found at http://www.utsouthwestern.edu/labs/danuser/software/ (the software and pdf file guide are included in the download).
- Representative images and movies can be found in Zhen et al., 2016.
Notes
- Imaging dishes should be gelatinized overnight.
- We recommend using pipettes, not suction, when removing medium from imaging dishes.
- When add medium to the cells in imaging dishes, we recommend tilting plate at an angle, slowly add medium to the lower edge and let the medium reach the cells slowly.
- The TIRF angle used is unique for individual cells and should be adjusted to achieve the best possible movies.
- To obtain representative movies, the focus of the movie should be at the middle layer of the nucleus. This can be recognized by the deeper black area (this can be achieved by adjusting the angle). Bright field can be used to check the nucleus area.
- In u-track 2.0, multiple movies can be loaded and processed. After being processed, each movie should be checked to ensure that cells should be no drift and rotation.
Recipes
- ES cell medium
DMEM
15% FBS
2 mM glutamine
100 U/ml penicillin-streptomycin
0.1 mM β-mercaptoethanol
1,000 U/ml LIF
0.1 mM non-essential amino acids - Live-cell imaging medium
FluoroBrite DMEM supplemented with 15% FBS
2 mM glutamine
100 U/ml penicillin-streptomycin
0.1 mM β-mercaptoethanol
1,000 U/ml LIF
0.1 mM non-essential amino acids
Acknowledgments
This work was supported, in whole or in part, by the National Cancer Institute of the National Institutes of Health under Award Number R03CA191443 (to XR). This work was also supported by grants from the CU-Denver Office Research Service (to XR) and the American Cancer Society Grant IRG 57-001-53 subaward (to XR). This protocol was originally developed in Zhen et al., 2016.
References
- Cuvier, O. and Fierz, B. (2017). Dynamic chromatin technologies: from individual molecules to epigenomic regulation in cells. Nat Rev Genet.
- Grimm, J. B., English, B. P., Chen, J., Slaughter, J. P., Zhang, Z., Revyakin, A., Patel, R., Macklin, J. J., Normanno, D., Singer, R. H., Lionnet, T. and Lavis, L. D. (2015). A general method to improve fluorophores for live-cell and single-molecule microscopy. Nat Methods 12(3): 244-250, 243 p following 250.
- Kusumi, A., Tsunoyama, T. A., Hirosawa, K. M., Kasai, R. S. and Fujiwara, T. K. (2014). Tracking single molecules at work in living cells. Nat Chem Biol 10(7): 524-532.
- Liu, Z., Lavis, L. D. and Betzig, E. (2015). Imaging live-cell dynamics and structure at the single-molecule level. Mol Cell 58(4): 644-659.
- Los, G. V., Encell, L. P., McDougall, M. G., Hartzell, D. D., Karassina, N., Zimprich, C., Wood, M. G., Learish, R., Ohana, R. F., Urh, M., Simpson, D., Mendez, J., Zimmerman, K., Otto, P., Vidugiris, G., Zhu, J., Darzins, A., Klaubert, D. H., Bulleit, R. F. and Wood, K. V. (2008). HaloTag: a novel protein labeling technology for cell imaging and protein analysis. ACS Chem Biol 3(6): 373-382.
- Tatavosian, R., Zhen, C. Y. and Ren, X. J. (2015). Single-molecule fluorescence microscopy methods in chromatin biology. Acs Sym Ser 1215:129-136.
- Zhen, C. Y., Tatavosian, R., Huynh, T. N., Duc, H. N., Das, R., Kokotovic, M., Grimm, J. B., Lavis, L. D., Lee, J., Mejia, F. J., Li, Y., Yao, T. and Ren, X. (2016). Live-cell single-molecule tracking reveals co-recognition of H3K27me3 and DNA targets polycomb Cbx7-PRC1 to chromatin. Elife 5.
- Zheng, Q. and Lavis, L. D. (2017). Development of photostable fluorophores for molecular imaging. Curr Opin Chem Biol 39: 32-38.
Article Information
Copyright
Duc and Ren . This article is distributed under the terms of the Creative Commons Attribution License (CC BY 4.0).
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Duc, H. N. and Ren, X. (2017). Labelling HaloTag Fusion Proteins with HaloTag Ligand in Living Cells. Bio-protocol 7(17): e2526. DOI: 10.21769/BioProtoc.2526.
- Zhen, C. Y., Tatavosian, R., Huynh, T. N., Duc, H. N., Das, R., Kokotovic, M., Grimm, J. B., Lavis, L. D., Lee, J., Mejia, F. J., Li, Y., Yao, T. and Ren, X. (2016). Live-cell single-molecule tracking reveals co-recognition of H3K27me3 and DNA targets polycomb Cbx7-PRC1 to chromatin. Elife 5.
Category
Developmental Biology > Cell signaling
Biochemistry > Protein > Labeling
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










