- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
A Quick, No Frills Approach to Mouse Genotyping
Published: Vol 2, Iss 15, Aug 5, 2012 DOI: 10.21769/BioProtoc.244 Views: 27362

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
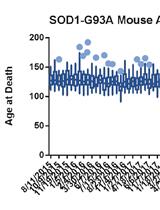
A High-throughput qPCR-based Method to Genotype the SOD1G93A Mouse Model for Relative Copy Number
Valerie R. Tassinari and Fernando G. Vieira
Jun 20, 2019 7319 Views
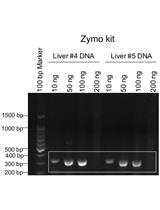
Genotyping of the OATP1B1 c. 521 T>C Polymorphism from the Formalin-Fixed Paraffin-Embedded (FFPE) Tissue Specimens: An Optimized Protocol
Alexandra R Crowe [...] Wei Yue
Aug 20, 2019 4334 Views
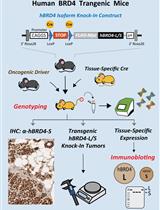
Conditional Human BRD4 Knock-In Transgenic Mouse Genotyping and Protein Isoform Detection
Michael Paul Lewis [...] Cheng-Ming Chiang
Apr 5, 2022 3632 Views
Abstract
Mice are extremely powerful mammalian genetic model organisms for basic and medical research, but managing a colony of transgenic mice is time consuming and expensive, many times requiring the help of dedicated technicians. Slow and laborious genotyping procedures add to the hassle. Outsourcing is costly and may not be as fast as desired, especially when setting up time sensitive experiments. Ultrafast genotyping protocols often require real-time PCR instruments and commercial reagents that may not be economical or practical. This protocol, adapted from methods suggested by The Jackson Laboratory, employs a minimalist approach that maximizes convenience by simplifying the tissue digestion/DNA extraction process and using a high-speed electrophoresis system for sample analysis. Genotype PCR results can be obtained in 3 h or less for as many samples as can fit in a PCR machine or can be efficiently handled by a user. Subsequent ethanol or chloroform purified DNA can be used in a standard PCR reaction to roughly identify a homozygous and a hemizygous mouse.
Materials and Reagents
- NaOH (NaOH pellets)
- Taq DNA Polymerase with ThermoPol buffer (New England Biolabs, catalog number: M0267X *, M0267L , or M0267S )
* Note: At 4,000U, ~800 μl of Taq serves several thousand PCR reactions. Buffer becomes a limiting reagent. ThermoPol Buffer recipe is available at NEB website. This buffer can be ordered separately from NEB (New England Biolabs, catalog number: B9004S ) - Primers, recommend to be 18-21 bp in length, have a melting temperature above 56 °C and around 58 °C, and produce amplicons of 150-600 bp.
- DNA loading buffer
Recommend Orange G (Sigma-Aldrich, catalog number: O3756 ) instead of bromophenol blue for loading dye - DNA ladder range for 100-800 bp range
Recommend 1 kb Plus DNA ladder (Life Technologies, catolog number: 10787-018 ) - Agarose
- EtBr (Sigma-Aldrich, catalog number: E8751 ) or SYBR Safe DNA Gel Stain (Life Technologies, catalog number: S33102 )
- NaOAc
- EtOH
- Phenol/Chloroform/Isoamyl alcohol (25: 24: 1) (Do not use acid phenol)
- 50 mM NaOH in dH2O (see Recipes)
- 10 mM dNTP Mix (see Recipes)
- 1x TAE buffer (see Recipes)
- dNTP set 100 mM each A, C, G, T (GE Life Sciences, catalog number: 28406552 ) (see Recipes)
- 0.3 M NaOAc in ddH2O (see Recipes)
Equipment
- PCR Thermal cycler (96 well capacity preferred)
- Centrifuges (mini for PCR tubes and microcentrifuge for 1.5 ml tubes)
- Liberty 1 buffer-less high speed gel system (Neuvitro, 6Mgel - SYS-LBT1)
- Liberty 1, 12-channel pipette compatible 13 teeth combs (Neuvitro, 6Mgel - CMS-1315)
- Multichannel Pipette 2-20 μl
- Repeat Pipettor 10-125 μl
- PCR tube with cap, 8 or 12 PCR strip tubes, or 96-well PCR plate
Procedure
- Part I. Digest tissue for genotyping
Note: This protocol is performed on 1-48 samples at the same time using 12-strip PCR tubes. The strip PCR tubes allow the use of multichannel pipettes to transfer solutions. 96 PCR plates can be used as well.- Tissue digestion:
- Place a roughly 2-3 mm ear clip, tail, or other tissue biopsy in each PCR tube as they are obtained.
- Add 75 μl of 50 mM NaOH to PCR tube. Make sure tissue sample is submerged.
Note: For adult mice ear clips add 75 μl, for adult tail clips 100 μl is preferred, and for neonatal mice tail or toe clips add 75 μl of 50 mM NaOH. Less than 75 μl makes it more difficult to use a repeat pipetor to squirt out a sufficient volume of 50 mM NaOH into each PCR tube. - Incubate in thermo cycler for 95 °C for 30 min to 1 h. 45 min is recommended.
Note: Samples have been heated for up to 2 h and as little as 15 min. 15-30 min is sufficient for neonatal tissue. - Immediately flick tubes.
Key feature: Tissue sample should partially break apart when PCR tube is flicked several times while still hot. Liquid should be cloudy. If tissue is not falling apart, incubate for longer. - Allow samples to cool to room temperature.
- Briefly spin to remove liquid from caps.
- Proceed to Part II or store samples at room temperature overnight for next day use.
Notes:- Samples can be kept at room temperature for up to 3 days. Samples left for over a week have been used successfully but not recommended.
- Freeze at -20 °C for long-term storage. Neutralization step for base pH created by NaOH is not required but may benefit long-term storage. 10x ThermolPol buffer or 1 volume of 0.3 M NaOAc in dH2O can be used to neutralize.
- Part II. PCR rection and gel electrophoresis
- PCR after tissue digestion:
- PCR master mix setup.
Note: Always make for more reactions than needed. For every 12 samples make for 2 samples extra, e.g. for 48 samples, make master mix for 56 reactions total.
Per reaction:
20 μl – dH2O
2.5 μl – 10x ThermoPol buffer
0.5 μl – 10 mM dNTPs
0.5 μl – 10 μM primer mix
0.1 μl – Taq DNA Polymerase
23.6 μl total - Aliquot 24 μl of master mix into new PCR tube(s) at room temperature.
- Transfer 2 μl of DNA from PCR tubes containing digested tissue (Part I) to PCR tubes containing aliquot of PCR master mix.
- Briefly spin.
- Samples can be kept at 4 °C until a PCR machine is available same day.
- PCR program
- 95 - 2 min
- 95 - 30 sec
- 56 - 30 sec
- 72 - 30 sec
- repeat steps ii, iii, iv 34x
- 72 - 1 min
- 10 - 5 min
END – completed reactions can be left at room temperature until ready for gel electrophoresis.
Tip: Do not modify the PCR reaction or program. Instead design all primers to be suitable under the same reaction condition. This way you can run multiple PCRs with different primers at the same time. This approach has been used to genotype mice containing multiple transgenes and for background backcrossing1,2.
- PCR master mix setup.
- 6 min gel electrophoresis.
- Cast a 2% agarose gel with EtBr or SYBR DNA gel stain in a liberty 1 apparatus (use 4 x 13 well combs to be able to load 48 samples and DNA ladder).
- Add 5 μl of DNA loading buffer to each PCR sample.
- Mix by pipetting up and down.
- Transfer 15 μl to 12 wells per lane in 2% agarose gel.
- Add DNA ladder with loading buffer to 13th well.
- Run gel electrophoresis at 200-220V for 6 min.
- Use appropriate gel viewer or imaging apparatus to identify bands.
- PCR after tissue digestion:
- Part III. Purifying genomic DNA and subsequent use
- You can purify DNA from digest in Part I in various ways. Two are listed:
- Precipitating DNA (crude purification suitable for a primer sequencing reaction or semi-quantitative PCR).
- Allow debri to settle after digestion procedure in Part I.
- Transfer top 50 μl of solution into 1.5 ml tube(s). Avoid picking up debri.
- Add 50 μl of 0.3 M NaOAc in dH2O, mix by flicking tubes.
- Add 300 μl of 100% EtOH, mix by flicking tubes.
- Incubate in -80 °C for 3 min.
- Centrifuge at 16,000 x g for 3 min.
- Dry pellet and resuspend in 50 μl dH2O or preferred buffer.
- DNA concentration can now be assessed by spectrophotometry.
- Phenol/Chloroform DNA purification (stringent purification)
- Allow debri to settle after digestion procedure in Part I.
- Transfer 50 μl of solution into 1.5 ml tube(s). Avoid picking up debri.
- Add 150 μl of 0.3 M NaOAc in dH2O, mix by flicking tubes.
- Add 200 μl of Phenol/Chloroform/Isoamyl alcohol (25: 24: 1), shake tubes.
- Centrifuge at 16,000 x g for 5 min.
- Transfer 150 μl of top phase to new 1.5 ml tube.
- Add 450 μl of 100% EtOH.
- Place tubes in -80 °C for 3 min.
- Centrifuge at 16,000 x g for 3 min.
- Remove supernatant without disturbing pellet.
- Add 1 ml of 75% EtOH and shake tube.
- Centrifuge at 16,000 x g for 3 min.
- Dry pellet and resuspend in 50 μl dH2O or preferred buffer.
- DNA concentration can now be assessed by spectrophotometry.
- Precipitating DNA (crude purification suitable for a primer sequencing reaction or semi-quantitative PCR).
- Purified DNA can be used to determine homozygote and heterozygote mice by semi-quantitative PCR
Note: This method will increase the chances of identifying a mouse homozygous for a gene if selective primers are not available. This method was used to select homozygote parent mice for either a transgene containing the tetracycline transactivator or yellow fluorescent protein1.- 2 μl of 0.05 μg μl-1 DNA can be used in PCR reaction outlined above with PCR program cycle number adjusted to 19-23x, 21x recommended.
Tip: PCR cycle number should be optimized to when a positive band for a given primer set can just be seen for a known heterozygous sample. - After gel run and imaging, differences in band intensity can be compared by eye or software, such as ImageJ.
- 2 μl of 0.05 μg μl-1 DNA can be used in PCR reaction outlined above with PCR program cycle number adjusted to 19-23x, 21x recommended.
- You can purify DNA from digest in Part I in various ways. Two are listed:
Recipes
- 50 mM NaOH in dH2O
Caution: NaOH (lye) solution is caustic! Wear gloves when handling. - 10 mM dNTP Mix
Recommend making mix from dNTP Set 100 mM each A,C,G,T - 10 μM Primer mix
- 1x TAE buffer
- 0.3 M NaOAc in ddH2O
Acknowledgments
This protocol was adapted from previous work (Lopez et al., 2011; Lopez et al., 2012).
References
- Lopez, M. E., Klein, A. D., Dimbil, U. J. and Scott, M. P. (2011). Anatomically defined neuron-based rescue of neurodegenerative Niemann-Pick type C disorder. J Neurosci 31(12): 4367-4378.
- Lopez, M. E., Klein, A. D., Hong, J., Dimbil, U. J. and Scott, M. P. (2012). Neuronal and epithelial cell rescue resolves chronic systemic inflammation in the lipid storage disorder Niemann-Pick C. Hum Mol Genet 21(13): 2946-2960.
Article Information
Copyright
© 2012 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Lopez, M. E. (2012). A Quick, No Frills Approach to Mouse Genotyping. Bio-protocol 2(15): e244. DOI: 10.21769/BioProtoc.244.
- Lopez, M. E., Klein, A. D., Dimbil, U. J. and Scott, M. P. (2011). Anatomically defined neuron-based rescue of neurodegenerative Niemann-Pick type C disorder. J Neurosci 31(12): 4367-4378.
Category
Molecular Biology > DNA > Electrophoresis
Molecular Biology > DNA > Genotyping
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link











