- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
[2-3H]Mannose-labeling and Analysis of N-linked Oligosaccharides
Published: Vol 7, Iss 14, Jul 20, 2017 DOI: 10.21769/BioProtoc.2393 Views: 7854
Reviewed by: Ralph BottcherAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

A Robust, One-step FRET Assay for Human Heparanase
Jyothi C. Sistla and Umesh R. Desai
Sep 5, 2019 4671 Views
Analysis of Lipid-linked Oligosaccharides Synthesized in vivo in Schizosaccharomyces pombe
Ayelen Valko [...] Cecilia D’Alessio
Sep 20, 2022 2258 Views
A Radioisotope-free Oligosaccharyltransferase Assay Method
Takahiro Yamasaki and Daisuke Kohda
Mar 5, 2019 6359 Views
Abstract
Modifications of N-linked oligosaccharides of glycoproteins soon after their biosynthesis correlate to glycoprotein folding status. These alterations can be detected in a sensitive way by pulse-chase analysis of [2-3H]mannose-labeled glycoproteins, with enzymatic removal of labeled N-glycans, separation according to size by HPLC and radioactive detection in a scintillation counter.
Keywords: N-linked oligosaccharideBackground
Following entry of a nascent polypeptide into the ER, it is subjected to several post-translational modifications, which are crucial for folding, maturation and quality control processes. The addition of the core oligosaccharide, Glc3Man9GlcNAc2 to produce N-linked glycoproteins is a very common modification and the first to occur (Benyair et al., 2011). Processing of the precursor N-glycan directs the glycoproteins to maturation and quality control machineries by creating recognition tags while in early secretory compartments (Tannous et al., 2015). At later stages, throughout the secretory pathway the core of the oligosaccharide serves as a platform for expansion of the sugar chains into complex glycans, the structures of which relate to the trafficking and function of the glycoproteins (Kamiya et al., 2012). Because the early N-linked glycan modifications reflect glycoprotein biosynthesis and quality control, the oligosaccharide processing has been the subject of many studies (Avezov et al., 2010; Hosokawa et al., 2010; Ninagawa et al., 2014; Ogen-Shtern et al., 2016). Glycoproteomic methods have greatly improved N-glycan characterization, but they do not allow the study of the dynamics of glycan processing in the early secretory pathway.
Here we describe a simplified pulse-chase method for the isolation and analysis of metabolically labeled N-linked oligosaccharides. The method includes radioactive labeling by [2-3H]Man followed by enzymatic removal of oligosaccharides by endo-beta-N-acetylglucosaminidase H (Endo H). Then, N-linked oligosaccharide isolation by molecular filtration and separation by high-performance liquid chromatography (HPLC), which discriminates between high-mannose glycan structures depending on their number of monosaccharide residues. The protocol allows analysis of the dynamics of N-linked glycan modification under different conditions, e.g., after drug treatment or modification of protein levels by overexpression or knockdown. Trimming to shorter species, Man5-6GlcNAc2, is a requirement for glycoprotein targeting to endoplasmic reticulum-associated degradation (ERAD) (Frenkel et al., 2003). Man1A (α1,2 mannosidase 1A ) appears to be involved in endoplasmic reticulum (ER) quality control and required for this trimming, as we present in an example. This is a surprising finding considering that the enzyme was thought to be located in the Golgi complex (Igdoura et al., 1999; Herscovics, 2001); a recent reevaluation locates it in quality control vesicles (Ogen-Shtern et al., 2016).
To conclude, the protocol presented here enables the study of the dynamics of N-linked high-mannose glycan modifications, which has an important role in glycoprotein quality control and trafficking. The advantages of the method are its simplicity, high sensitivity of detection and unique information on the dynamics of N-glycan processing in the early secretory pathway.
Materials and Reagents
- 100 mm tissue culture dishes (Corning, catalog number: 430167 )
- 1.5 and 2 ml microcentrifuge tubes (Eppendorf)
- Microcon Amicon Ultra 0.5 ml 30K or Centricon ultracel YM-30 (EMD Millipore, catalog number: UFC503024 )
- Spherisorb NH2 column, 5 µm, 4.6 x 250 mm (WATERS, catalog number: PSS831115 )
- HEK-293 cells (ATCC, catalog number: CRL-1573 )
- Dulbecco’s modified Eagle’s medium (DMEM) (Thermo Fisher Scientific, GibcoTM, catalog number: 41965039 )
- Dulbecco’s modified Eagle’s medium-glucose free (Sigma-Aldrich, catalog number: D5030 )
- Fetal bovine serum (FBS) (Biological Industries, catalog number: 04-001-1A-US )
- Fetal bovine serum (FBS), dialyzed (Biological Industries, catalog number: 04-011-1A-US )
- Sodium pyruvate solution (100 mM) (Sigma-Aldrich, catalog number: S8636 )
- Mannose, D-[2-3H(N)]- (Specific Activity: 15-30 Ci/mmol) (PerkinElmer, catalog number: NET570A )
- Dulbecco’s phosphate buffered saline (PBS) (Sigma-Aldrich, catalog number: D1408 )
- Protein A-Agarose beads, IPA 300 (REPLIGEN, catalog number: 10-1003 )
- Rabbit polyclonal anti-H2 carboxy-terminal produced in our lab (Tolchinsky et al., 1996)
- Endo Hf Kit (1,000,000 U/ml) (New England Biolabs, catalog number: P0703S )
- Standard oligosaccharide mixture prepared by glycoprotein metabolic labeling with [14C] or [3H] and separation with endo H
- Acetonitrile LiChrosolv (gradient grade for liquid chromatography) (Merck, catalog number: 100030 )
- Phosphoric acid solution (49-51%, for HPLC) (Sigma-Aldrich, catalog number: 79607 )
- Opti-Fluor scintillation fluid (PerkinElmer, catalog number: 6013199 )
- Triton X-100 (BDH, catalog number: 306324N )
- Protease inhibitor cocktail (Sigma-Aldrich, catalog number: P2714 )
- Sodium deoxycholate (Sigma-Aldrich, catalog number: 30970 )
- Sodium dodecyl sulfate (SDS) (Bio-Rad Laboratories, catalog number: 1610301 )
- Sodium phosphate (Na3PO4) (Sigma-Aldrich, catalog number: 342483 )
- Buffer A (see Recipes)
- Buffer D (see Recipes)
- HPLC solvent (see Recipes)
Equipment
- Eppendorf centrifuge (Eppendorf, model: 5415 D )
- CO2 incubator
- -80 °C freezer
- LS 6500 Liquid Scintillation Counting systems (Beckman Coulter, model: LS 6500, catalog number: 510720 )
- 600E Multisolvent delivery System controller (HPLC) (WATERS, catalog number: WAT062710 )
Note: This product has been discontinued. - SpeedVac Concentrator 5301, incl. 48 x 1.5/2.0 ml fixed-angle rotor (Eppendorf, model: Concentrator 5301, catalog number: 5301 000.016 )
- Frac-100 fraction collector (Amersham Biosciences, model: FRAC-100, catalog number: 18-1000-77 )
- Vibra-Cell ultrasonic processors VCX 750 (Sonics and Materials, model: VCX 750 , catalog number: 690-003)
Procedure
- For metabolic labeling of N-linked glycans, include a pulse and chase samples (usually 1 to 3) for each condition analyzed, and a negative pulse control sample, to be immunoprecipitated with a preimmune antibody. Here HEK293 cells grown in a 90 mm dish were used for each sample. Transfections of HEK 293 cells (about 2 x 106) were carried out according to the calcium phosphate method, 24 h before the experiment. Other transfection methods can be used.
- Remove growth medium from 90 mm dish, scrape the cells with complete DMEM and transfer into a 2 ml Eppendorf tube. Spin the cells (6 sec at 16,000 x g) and rinse with 2 x 1 ml of glucose-free medium. Starve the cells from glucose by incubation in 1 ml of freshly prepared pre-warmed (37 °C) glucose-free medium supplied with 10% dialyzed FBS and 4 mM sodium pyruvate for 30 min in a CO2 incubator at 37 °C (Figure 1). Leave the lid of the tube open during this and subsequent incubations at 37 °C. Labeling in Eppendorf tubes allows the use of small volumes of medium and radioactive precursor in the following steps.
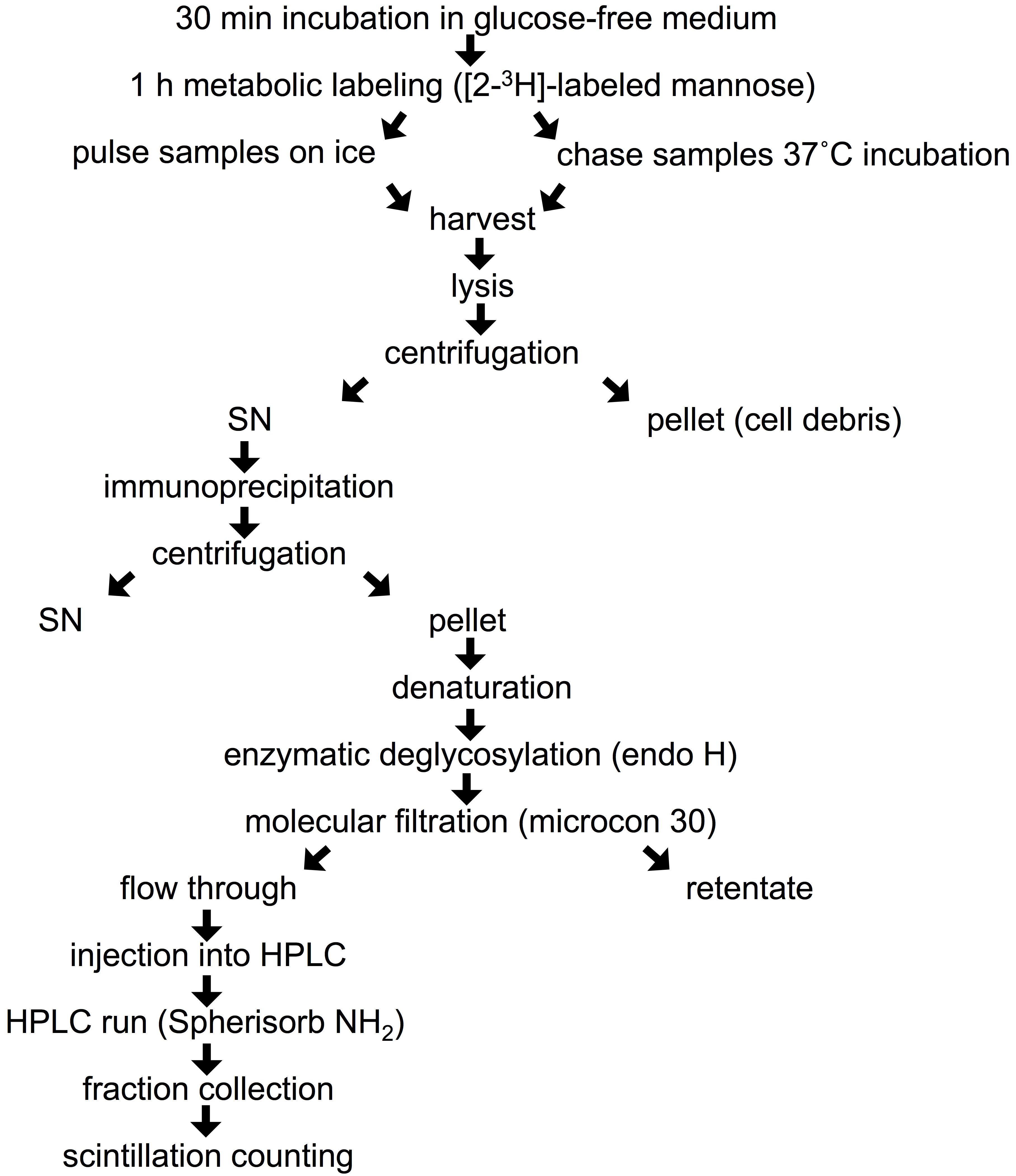
Figure 1. Flow chart of experimental procedure - Replace the starvation medium with 1 ml of pre-warmed (37 °C) glucose-free medium containing 10% dialyzed FBS, 4 mM sodium pyruvate and 400 μCi of [2-3H]-labeled mannose (stock concentration 1 μCi/1 μl), and incubate the cells in a CO2 incubator at 37 °C for 1 h (pulse).
- Remove the labeling medium from each sample, and add 2 ml of PBS at 4 °C to the samples corresponding to pulse and place them on ice, while the samples corresponding to the chase are rinsed 3 times with 1 ml of pre-warmed (37 °C) normal complete culture medium, and then placed in a CO2 incubator at 37 °C with 1 ml of pre-warmed normal complete culture medium for the desired chase periods (usually 30 min to 8 h, depending on the turnover of the studied protein).
- Rinse the pulse samples 3 times with 1 ml of ice-cold PBS.
- Short-spin the cells at 16, 000 x g (6-10 sec), discard the supernatant and proceed with step 7 or freeze the cell pellet at -80 °C (for up to several months).
- Perform steps 5 and 6 also for the chase samples at the end of the chase time.
- Remove the cells from the freezer and lyse them by addition of 400 μl of buffer A, vortexing briefly and incubating for 30 min on ice.
- Centrifuge the lysates at 16,000 x g for 30 min at 4 °C. Transfer the supernatant to a new 1.5 ml Eppendorf tube and discard the pellet.
- For immunoprecipitation of the H2a glycoprotein shown in Data analysis, add 20 μl/sample of Protein A-Agarose beads 1:1 suspension and 5 μl/sample of rabbit polyclonal anti H2a antibody to the supernatant from step 9.
- Incubate the samples for 4-16 h at 4 °C, constantly mixing by slow rotation.
- Spin down the beads at 16,000 x g for 30 sec, and carefully remove the supernatant.
- Wash the beads by adding 500 μl buffer D and vortexing. Spin as in step 12 and remove the supernatant. Repeat the rinse 3 times.
- Add 20 μl denaturing buffer (supplied with the NEB endo-H kit) to the bead pellet and boil the samples for 5 min.
- Spin down the beads (16,000 x g for 30 sec), and transfer the supernatant into a new Eppendorf tube. Then 0.5 μl of endo-H enzyme is added to each sample along with 2 μl of the reaction buffer (also supplied with the NEB endo-H kit), and the samples are incubated at 37 °C for 3 h.
- To separate the released glycans from the protein, the sample is diluted 5 times with double distilled water (DDW) and placed on top of a molecular filter (Microcon Ultracel YM30), with a 30 kDa cut-off, then centrifuged at 14, 000 x g for 3 min (the filter must be washed with 0.5 ml DDW and centrifuged for 3.5 min at 14,000 x g prior loading the sample). Apply 100 μl of DDW to Eppendorf tubes containing the endo-H reaction and transfer to the corresponding molecular filter. Repeat centrifugation of the samples. This washout is repeated 1 more time, keeping the flow through (total throughput is about 300 μl).
- Check 10% in a scintillation counter (30 μl). If you get at least 100 cpm, proceed to next step. Less than this amount will give insufficient cpm for further analysis.
- Place the tube containing the endo-H reaction flow through (steps 16 and 17) in a SpeedVac concentrator, and dry the samples completely (this can be done up to 45 °C to accelerate this process, usually taking 1-2 h).
- To prepare the samples for the HPLC, resuspend the dry pellets in 12 μl of the HPLC solvent (acetonitrile:water, 63:37, 2.5% phosphoric acid).
- Adjust the HPLC device (connected to the Spherisorb column) to constant flow of the solvent (1 ml/min) and pressure (a value between 1,000 and 2,000 psi), and place a fraction collector to change a tube every 1 min.
- Load the samples into the HPLC device and start the fraction collector simultaneously.
- Collect 65 fractions of 1 ml from each sample run and from a run of a standard oligosaccharide mixture. We use a standard mixture prepared by glycoprotein metabolic labeling with [14C] and separation with endo-H (kind gift of Armando Parodi).
- Transfer 500 μl from each fraction to a scintillation vial, and mix the contents with 3 ml of water-miscible scintillation fluid (Opti-fluor).
- Load the vials and read in a scintillation counter using settings for [3H].
- The cpm readout is plotted as a function of fraction number.
- The cpm values within a peak represent a specific glycan species, identified according to the parallel run of the standard oligosaccharide mixture.
- The sum of cpm values for the fractions within each peak reflects the absolute amount of a specific glycan species. In order to compensate for the fact that species containing more mannose residues acquire more label, the ‘relative molar amount’ of each glycan species is calculated as follows: the absolute amount in cpm of each glycan species is divided by the number of mannose residues that it contains, resulting in a ‘relative molar amount’. This value is then divided by the sum of the relative molar amounts of all glycan species obtained, to obtain the ‘molar percent of total’ for each specific glycan species (Figure 2). An example of the use of this procedure is shown in Figure 3.
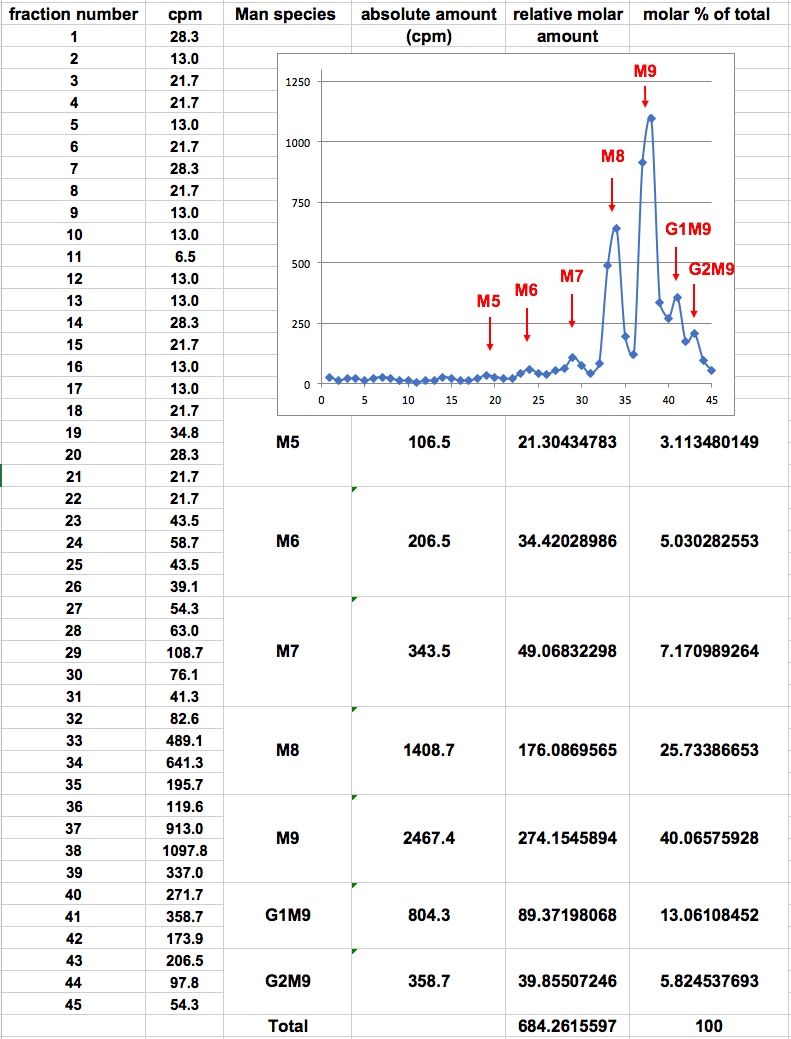
Figure 2. Example of calculation of relative molar amounts
Data analysis
The effect of knockdown of ManIA was tested in cells expressing the ERAD substrate H2a, labeled with [2-3H]Man. Cells were lysed, H2a was immunoprecipitated and treated with endo-H. The N-linked oligosaccharides were separated by HPLC, and fractions were counted in a beta counter. M9 to M5 stand for Man9GlcNAc2 to Man5GlcNAc2. Relative molar amounts of each oligosaccharide species were calculated based on mannose content. We converted the cpm values obtained for each glycan species, the percent of each species relative to the total sum of the relative molar amounts of all species present was then plotted. Only the values for 3 h chase are shown. Three independent biological replicate experiments were performed (Figure 3).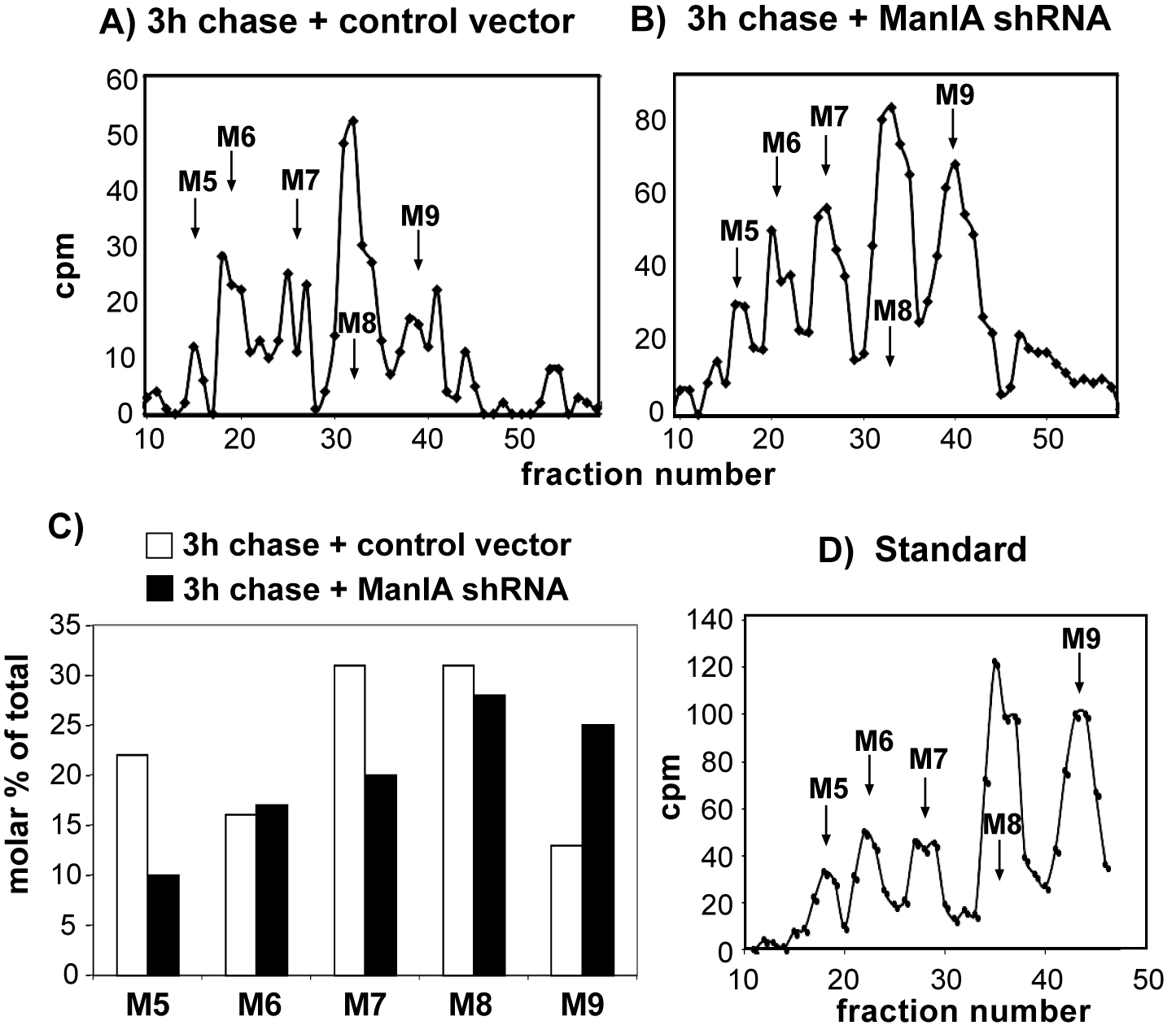
Figure 3. Trimming of high mannose residues on an ERAD substrate glycoprotein and the requirement of Man1A. HEK 293 cells cotransfected with an H2a-encoding vector together with pSUPER encoding control anti-lacZ shRNA (A) or anti-Man1A shRNA (B) were pulse-labeled for 1 h with [2-3H]Man in glucose-free medium and chased for 0 or 3 h in complete medium. C. The knockdown of Man1A significantly inhibited extensive mannose trimming, causing the accumulation of Man9GlcNAc2 (M9). One representative experiment of three independent biological replicate experiments is shown. D. Standard [14C] M9 to M5 oligosaccharide mixture.
Notes
Radioactive waste produced in the diverse steps should be collected and stored properly, according to the regulations at your institution.
Recipes
- Buffer A
1% Triton X-100
0.5% (w/v) sodium deoxycholate
Protease inhibitor cocktail 2% v/v in PBS - Buffer D
0.5% Triton X-100
0.25% (w/v) sodium deoxycholate
0.5% (w/v) SDS
Protease inhibitor cocktail 2% (v/v) in PBSNote: Stocks (5x) of buffers A and D can be prepared and stored in aliquots at -20 °C. Fresh protease inhibitor cocktail should be added just before use.
- HPLC solvent
63% acetonitrile
37% DDW
2.5% phosphoric acid
Acknowledgments
Research related to this work is supported by grants from the Israel Science Foundation (1593/16) and the Recanati Foundation. A related protocol can be found in Avezov et al., 2010.
References
- Avezov, E., Ron, E., Izenshtein, Y., Adan, Y. and Lederkremer, G. Z. (2010). Pulse-chase analysis of N-linked sugar chains from glycoproteins in mammalian cells. J Vis Exp (38): 1899.
- Benyair, R., Ron, E. and Lederkremer, G. Z. (2011). Protein quality control, retention, and degradation at the endoplasmic reticulum. Int Rev Cell Mol Biol 292: 197-280.
- Frenkel, Z., Gregory, W., Kornfeld, S. and Lederkremer, G. Z. (2003). Endoplasmic reticulum-associated degradation of mammalian glycoproteins involves sugar chain trimming to Man6-5GlcNAc2. J Biol Chem 278(36): 34119-34124.
- Herscovics, A. (2001). Structure and function of Class I α1,2-mannosidases involved in glycoprotein synthesis and endoplasmic reticulum quality control. Biochimie 83(8): 757-762.
- Hosokawa, N., Tremblay, L. O., Sleno, B., Kamiya, Y., Wada, I., Nagata, K., Kato, K. and Herscovics, A. (2010). EDEM1 accelerates the trimming of α1,2-linked mannose on the C branch of N-glycans. Glycobiology 20(5): 567-575.
- Kamiya, Y., Satoh, T. and Kato, K. (2012). Molecular and structural basis for N-glycan-dependent determination of glycoprotein fates in cells. Biochim Biophys Acta 1820(9): 1327-1337.
- Igdoura, S. A., Herscovics, A., Lal, A., Moremen, K. W., Morales, C. R. and Hermo, L. (1999). α-mannosidases involved in N-glycan processing show cell specificity and distinct subcompartmentalization within the Golgi apparatus of cells in the testis and epididymis. Eur J Cell Biol 78(7): 441-452.
- Ninagawa, S., Okada, T., Sumitomo, Y., Kamiya, Y., Kato, K., Horimoto, S., Ishikawa, T., Takeda, S., Sakuma, T., Yamamoto, T. and Mori, K. (2014). EDEM2 initiates mammalian glycoprotein ERAD by catalyzing the first mannose trimming step. J Cell Biol 206(3): 347-356.
- Ogen-Shtern, N., Avezov, E., Shenkman, M., Benyair, R. and Lederkremer, G. Z. (2016). Mannosidase IA is in quality control vesicles and participates in glycoprotein targeting to ERAD. J Mol Biol 428(16): 3194-205.
- Tannous, A., Pisoni, G. B., Hebert, D. N. and Molinari, M. (2015). N-linked sugar-regulated protein folding and quality control in the ER. Semin Cell Dev Biol 41: 79-89.
- Tolchinsky, S., Yuk, M. H., Ayalon, M., Lodish, H. F. and Lederkremer, G. Z. (1996). Membrane-bound versus secreted forms of human asialoglycoprotein receptor subunits. Role of a juxtamembrane pentapeptide. J Biol Chem 271(24): 14496-14503.
Article Information
Copyright
© 2017 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Shenkman, M., Ogen-Shtern, N. and Lederkremer, G. Z. (2017). [2-3H]Mannose-labeling and Analysis of N-linked Oligosaccharides. Bio-protocol 7(14): e2393. DOI: 10.21769/BioProtoc.2393.
Category
Biochemistry > Carbohydrate > Oligosaccharide > Labelling
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link











